- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Determination of Hydrodynamic Radius of Proteins by Size Exclusion Chromatography
Published: Vol 7, Iss 8, Apr 20, 2017 DOI: 10.21769/BioProtoc.2230 Views: 34368
Reviewed by: Yanjie LiWenrong HeMurugappan Sathappa

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Heterologous Expression and Purification of Catalytic Domain of CESA1 from Arabidopsis thaliana
Venu Gopal Vandavasi and Hugh O’ Neill
Oct 20, 2016 7547 Views
Expression and Purification of the GRAS Domain of Os-SCL7 from Rice for Structural Studies
Shengping Li [...] Yunkun Wu
Feb 5, 2017 8563 Views
X-ray Crystallography: Seeding Technique with Cytochrome P450 Reductase
Bixia Zhang [...] ChulHee Kang
Nov 5, 2022 2328 Views
Abstract
Size exclusion chromatography (SEC) or gel filtration is a hydrodynamic technique that separates molecules in solution as a function of their size and shape. In the case of proteins, the hydrodynamic value that can be experimentally derived is the Stokes radius (Rs), which is the radius of a sphere with the same hydrodynamic properties (i.e., frictional coefficient) as the biomolecule. Determination of Rs by SEC has been widely used to monitor conformational changes induced by the binding of calcium (Ca2+) to many Ca2+-sensor proteins. For this class of proteins, SEC separation is based not just on the variation in protein size following Ca2+ binding, but likely arises from changes in the hydration shell structure.
This protocol aims to describe a gel filtration experiment on a prepacked column using a Fast Protein Liquid Chromatography (FPLC) system to determine the Rs of proteins with some indications that are specific for Ca2+ sensor proteins.
Background
Gel filtration separates molecules of different sizes and shapes based on their relative abilities to penetrate a bed of porous beads with well-defined pore sizes, which identifies the fractionation range. Molecules larger than the fractionation range, which are completely excluded from entering the pores, flow quickly through the column and elute first at the void volume (V0), which is the interstitial volume outside the support particles. Molecules smaller than the fractionation range, which are able to diffuse into the pores of the beads, have access to the total volume available to the mobile phase, therefore they move through the bed more slowly and elute last. Molecules with intermediate dimensions will be eluted with an elution volume (Ve) comprised between the void volume and the total volume available to the mobile phase (the smaller the molecule, the greater its access to the pores of the matrix, and thus the greater is its Ve).
The molecular weight of a protein can be determined by comparison of its elution volume parameter Kd, which represents the distribution of a given solute between the stationary and mobile phases (see Data analysis below), with those of different known calibration standards.
If the protein of interest has the same shape (generally globular) as the standard calibration proteins, the gel filtration experiment provides a good estimate of its molecular weight. However, because the shape of proteins can vary significantly and may be not known for an unknown protein, care must be taken in the determination of molecular size from elution volume. For example, a protein with an elongated shape could elute at a position that does not correspond to its dimension and which is significantly different from the position of a spherical protein having the same molecular weight. This is the case for some Ca2+ sensor proteins, e.g., calmodulins from different organisms (Sorensen and Shea, 1996; Sorensen et al., 2001; Astegno et al., 2014; Astegno et al., 2016; Vallone et al., 2016), which have anomalous migrations in gel filtration, resulting in a defined overestimation of the molecular weight due to their highly extended conformation. Thus, it is clear that in a gel filtration experiment the elution profile of proteins is closer to their Stokes radius (Rs) rather than to their molecular weight. Rs is a hydrodynamic value indicative of the apparent size of the dynamic solvated/hydrated protein.
For this reason, a SEC-based approach has been employed to resolve Ca2+-induced changes in the hydrated shape of Ca2+ sensor proteins by determination of their Rs in apo- and Ca2+-bound conditions. Ca2+ binding usually causes a decrease in the Rs (Sorensen and Shea, 1996; Sorensen et al., 2001; Astegno et al., 2016). The same SEC-based approach may be applicable to the detection of other protein-small molecule (e.g., other metals) interactions that cause changes in the structure of the protein to a less or more extended conformation (Asante-Appiah and Skalka, 1997; Bagai et al., 2007; De Angelis et al., 2010).
The values of Rs have been reported for a large number of proteins; in particular, some proteins are especially convenient for calibration of gel filtration columns (le Maire et al., 1986; Uversky, 1993) (Table 1). A gel filtration column can determine the hydrodynamic size, Rs, of a sample protein by comparison with the Rs of these water-soluble calibration proteins.
Table 1. Standards for calibrating analytical gel filtration
Materials and Reagents
- 0.22 µm syringe filters with low protein retention (Thermo Fischer Scientific, Thermo ScientificTM, catalog number: 42204-PV )
- 0.22 µm vacuum filtration unit (Sartorius, catalog number: 180C7-E )
- Superdex 200 10/300 GL prepacked column (GE Healthcare, catalog number: 17517501 )
- Superose 12 10/300 GL prepacked column (GE Healthcare, catalog number: 17-517-301 )
- 20% EtOH–minimum 1 L (Sigma-Aldrich, catalog number: 51976 )
- MilliQ water–minimum 1 L
- Protein markers:
Thyroglobulin (Sigma-Aldrich, catalog number: T9145 )
Apo-Ferritin (Sigma-Aldrich, catalog number: A3660 )
β-amylase (Sigma-Aldrich, catalog number: A8781 )
Catalase (Sigma-Aldrich, catalog number: C9322 )
Aldolase (Sigma-Aldrich, catalog number: A2714 )
Alcohol dehydrogenase (Sigma-Aldrich, catalog number: A8656 )
Albumin (Sigma-Aldrich, catalog number: A8531 )
Ovalbumin (Sigma-Aldrich, catalog number: A5503 )
Carbonic anhydrase (Sigma-Aldrich, catalog number: C7025 )
Myoglobin (Sigma-Aldrich, catalog number: M0630 )
Cytochrome c (Sigma-Aldrich, catalog number: C7150 ) - Gel filtration Calibration Kits with an optimized range of proteins are also commercially available:
- Gel Filtration Markers Kit for Protein Molecular Weights 29,000-700,000 Da (Sigma-Aldrich, catalog number: MWGF1000 )
- Gel Filtration Markers Kit for Protein Molecular Weights 12,000-200,000 Da (Sigma-Aldrich, catalog number: MWGF200 )
- Gel Filtration Markers Kit for Protein Molecular Weights 6,500-66,000 Da (Sigma-Aldrich, catalog number: MWGF70 )
- Gel Filtration Calibration Kits
High molecular weight (GE Healthcare, catalog number: 28-4038-42 )
Low molecular weight (GE Healthcare, catalog number: 28-4038-41 ) - Calcium chloride dihydrate (CaCl2·2H2O) (Sigma-Aldrich, catalog number: 21097 )
- Ethylene glycol-bis(2-aminoethylether)-N,N,N’,N’-tetraacetic acid (EGTA) (Sigma-Aldrich, catalog number: E4378 )
- Sodium hydroxide (NaOH) (Sigma-Aldrich, catalog number: 1.06462 )
- Hydrochloric acid (HCl) (Sigma-Aldrich, catalog number: H1758 )
- DL-dithiothreitol, anhydrous (DTT) (Sigma-Aldrich, catalog number: D9779 )
- Trizma® base (Sigma-Aldrich, catalog number: T1503 )
- Potassium chloride (KCl) (Sigma-Aldrich, catalog number: 31248 or P9333 )
Note: The product “ 31248 ” has been discontinued. - HEPES (Sigma-Aldrich, catalog number: H3375 )
- Sodium chloride (NaCl) (Sigma-Aldrich, catalog number: S7653 )
- Acetone (Sigma-Aldrich, catalog number: 439126 )
- Dextran blue (Sigma-Aldrich, catalog number: D4772 )
- 2 M CaCl2 (20 ml) (see Recipes)
- 0.1 M EGTA (100 ml, pH 8) (see Recipes)
- 0.1 M DTT (see Recipes)
- Mobile phase (see Recipes)
- 5 mM Tris, 150 mM KCl, pH 7.5 (for calibration of Superose 12 column 10/300GL)
- 5 mM Tris, 150 mM KCl, 5 mM EGTA, 1 mM DTT pH 7.5
- 5 mM Tris, 150 mM KCl, 5 mM CaCl2, 1 mM DTT pH 7.5
- 50 mM HEPES, 150 mM NaCl, 0.1 mM DTT pH 7.5 (for calibration of Superdex 200 10/300 GL)
- Total volume available to the mobile phase marker (see Recipes)
- Void volume marker (see Recipes)
- Protein calibration standards (see Recipes)
- Protein sample (see Recipes)
Equipment
- ÄKTA FPLC system (or similar liquid chromatography system, e.g., ÄKTA pure , ÄKTA prime plus , ÄKTA start ) including injector, one pump, UV-detector, Fraction collector (GE Healthcare, models: ÄKTA pure , ÄKTA prime plus , ÄKTA start )
Note: ÄKTA FPLC has been discontinued and replaced with ÄKTA pure . - Sample loop kit (GE Healthcare, catalog number: 18-0404-01 )
- Hamilton syringe (500 µl) (Hamilton, catalog number: 81217 or Sigma-Aldrich, catalog number: S9266 )
- Vacuum pump
- Tabletop centrifuge (Eppendorf, model: 5424 R )
Software
- UNICORNTM 4.0 control software for ÄKTA FPLC chromatography system (GE Healthcare)
- Origin 8.0 (OriginLab Corporation, Northampton, MA)
Procedure
The following protocol describes the determination of Rs by using a prepacked column in a typical FPLC system (Figure 1). Buffers and other solutions are delivered via a system pump, and the sample can be load in different ways (e.g., by a syringe and a sample loop or by a sample pump). A detection system (e.g., UV/Vis absorbance, conductivity) is located after the column to control the separation process and the proteins are collected in the fraction collector. It is possible to operate the system manually, although the system is usually controlled by software. In particular, in the ÄKTA FPLC all data and parameters of the separation run are displayed in the System Control module of UNICORN software, which permits control of chromatography systems (e.g., system settings, run data, curves) and on-line monitoring of separation processes. Refer to the UNICORN User Reference Manual for software functionality.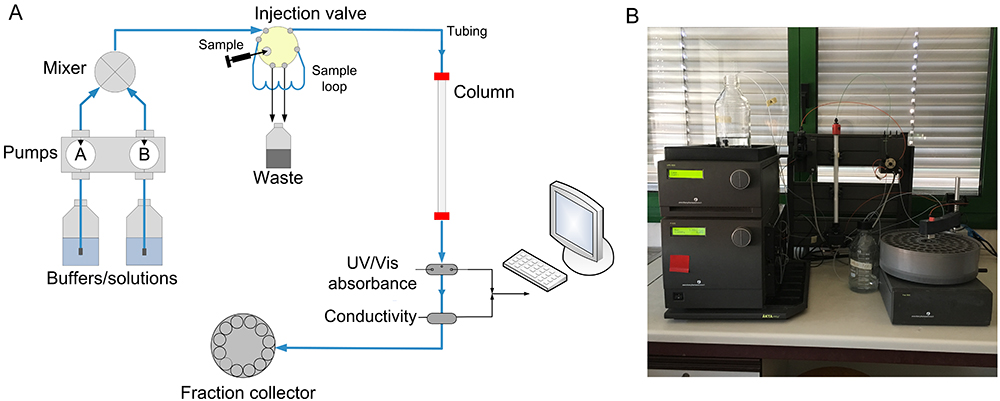
Figure 1. Typical FPLC system. A. Scheme of basic components and typical flow path for a chromatography system; B. Picture of GE Healthcare ÄKTA FPLC apparatus.
- Filter and de-gas all solutions (MilliQ water, eluent, 20% ethanol) through a 0.22 µm filter.
- Move the input lines of FPLC system from 20% EtOH and place them into MilliQ water.
Note: Make sure that the back-pressure over the column does not exceed the maximum pressure value mandated by the manufacturer. - Connect the column (injector outlet–column inlet and column outlet–detector) (Figure 2).
Note: Make sure that there is no air in the valves and tubing prior to connecting the column to a chromatography system. Ensure that the inlet of the column is filled with liquid; connect it to the system in a drop-to-drop fashion (unscrew the inlet of the column, let the buffer flow through the system and drop to the inlet of the column; then, unscrew the outlet of the column and connect to the detector).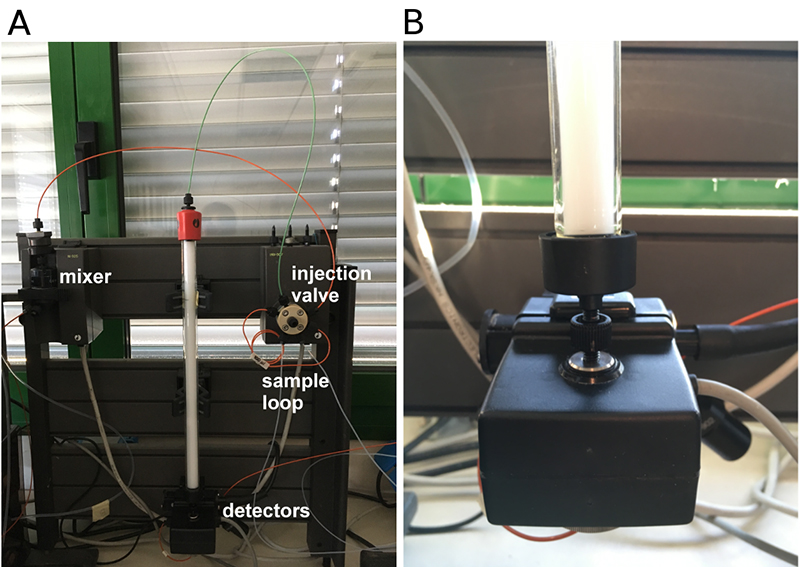
Figure 2. Chromatography column connection. A. Detailed view of Superose 12 10/300 GL prepacked column connected to ÄKTA FPLC system and of some important components of the instrument, i.e., mixer valve, injection valve, sample loop, and detectors. B. Detail of the column outlet and its connection to the detectors. - When using the column after prolonged storage or for the first time equilibrate the column as follows:
- At least 2 times the volume of the column (2x column volumes) of MilliQ water at a flow rate of 0.5 ml/min. Columns are usually stored in 20% ethanol, which produce high back pressures, so the initial flow rates should not exceed half the maximum recommended flow rate. Since ethanol might cause precipitation of the salts used in the mobile phase, it is advisable to flow MilliQ water until complete ethanol removal before proceeding with the equilibration.
- 2x column volumes of mobile phase (eluent) at a flow rate of 0.5-1 ml/min.
Note: The eluent should be chosen to ensure full sample solubility. In order to avoid any unwanted ionic interactions between the matrix and the solute molecules, the use of a buffer with an ionic strength equivalent or greater to 0.15 M NaCl or KCl is recommended. The manufacturer usually provides a list of some useful eluent compositions for a given column. - Connect the loop (100 μl loop size) to injection valve. Wash the loop by injection with MilliQ water (at least 5 times the volume of the loop). Repeat the wash with the protein’s eluent (according to the column used).
- Prepare the samples and remove particulate material from protein solution by either centrifuging (10,000 x g for 15 min) or filtering samples through a 0.22 µm filter.
- The sample is injected through the injection valve which has two positions, load and inject. When the valve is set on load, the loop can be filled with the sample (Figure 3A, blue line) and the eluent is being bypassed to the column (Figure 3A, green line). When the valve is set on inject, the eluent flows through the loop and moves the sample to the column (Figure 3B, green line).
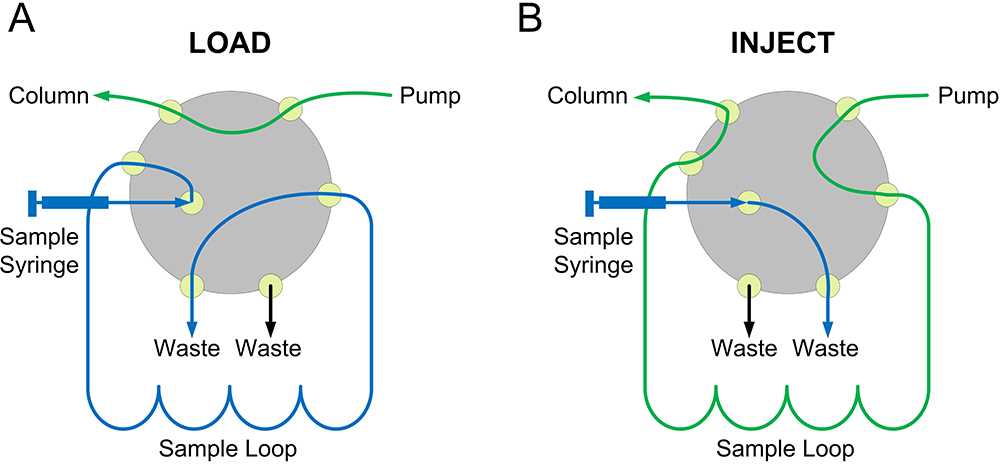
Figure 3. Schematic representation of the injection valve in the load (A) and inject (B) positions - Determine the void volume by loading blue dextran (200-300 μl at 1-2 mg/ml dissolved in the mobile phase, see Recipes) to the column via loop. This concentration of blue dextran will give an absorbance at 280 nm (A280) of 0.5-1 in the peak fraction. Dextran blue can also be used to control column packing.
Note: An excess of sample is used to ensure that the sample loop is filled completely (complete filling method). To achieve 95% of maximum loop volume about 2 to 3 loop volumes of sample are necessary. Before injection, make sure the flow rate is correctly set and that the injection valve is set to ‘load’. Load the syringe with the sample (2-3 times the loop volume) and expel the air bubbles. It may be necessary to flick the syringe to push the bubbles up. Gently inject the syringe contents into the loop. Leave the syringe in position and set the injection valve to ‘Inject’. The ideal volume loads correspond to sample volumes between 1-2% of the total column volume to obtain sharp peaks. Additionally, solutions containing high protein concentrations (> 2 mg/ml) can lead to increased viscosity and hence decreased diffusion coefficients of solutes (Ricker and Sandoval, 1996).The typical flow rate is 0.5-1 ml/min, although by decreasing the flow rate in SEC experiments, greater resolution can be obtained on a given column (refer to the manufacturer’s recommendation for flow rate). - Inject an aliquot of 200-300 μl of acetone at 5 mg/ml (0.63% v/v dissolved in the mobile phase, see Recipes) into the column to determine the total volume available to the mobile phase. Detection of acetone is possible at 280 nm.
- Prepare a mixture of selected standard proteins by dissolving them in the eluent buffer at 0.5-1 mg/ml (or at concentrations indicated by the manufacturer, see Recipes). Check for protein complete dissolution before applying to the column and then determine Ve for each protein standard.
Note: Typically, five protein standards are sufficient to obtain a good calibration plot. It is possible to use more than one standard in the same run (only if the peaks of the different standards are completely resolved). The standards must be chosen accurately, taking into account the fractionation range of the selected column. - Apply the protein sample (200-300 μl at 1 mg/ml) with the unknown Rs and determine the corresponding Ve.
Note: For determination of Ca2+-dependent change in Rs of Ca2+ sensor proteins, the sample protein must be loaded first in the absence and then in the presence of Ca2+. Apo conditions include 5 mM Tris, 150 mM KCl, 1 mM DTT, and 5 mM EGTA at pH 7.5, while Ca2+-saturated conditions include 5 mM Tris, 150 mM KCl, 1 mM DTT, and 5 mM CaCl2 at pH 7.5. - If the column has to be stored, wash it with 2x column volumes of MilliQ water and then at least 2x column volumes of 20% ethanol. Store at 4 °C.
Note: For column data, cleaning-in-place conditions, column performance controls, buffers, and solvent resistance always refer to the manufacturer’s instructions.
Data analysis
- Results in gel filtration are typically visualized as a plot of the volume of the mobile phase eluted through the column versus the detector signal (for proteins usually signal at 280 nm) which is called chromatogram. The elution volume of a given molecule is determined by calculating the volume of eluent from the point of injection to the center of the elution peak.
Note: Peak integration in the chromatography software is used to identify and measure curve characteristics, such as elution volumes, peak areas, and peak widths. In particular, to perform a basic peak integration in the Evaluation module of Unicorn software 4.0:
Select the Integrate:Peak Integrate menu command→Select a source curve→Select Calculate baseline in the Baseline list. The peaks in the integrated chromatogram are automatically labelled with their elution volumes. Several other peak characteristics are also calculated automatically. To display other characteristics, select the options that you want to display from the Select peak table columns list. - The distribution coefficient Kd, which represents the fraction of the stationary phase available for diffusion of a given solute species, can be expressed as follows:
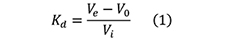
where (see also Figure 4),
Ve is the elution volume for a particular protein,
V0 is the void volume for the column, i.e., the elution volume of molecules totally excluded from the pores of the beads,
Vi is the volume of the solvent inside the matrix and is equal to the volume of the stationary phase.
In the method described in this protocol, Vi is determined by the measurement of the elution volume of acetone (i.e., the maximum retention volume experienced by a small molecule that has access to the total volume available to the mobile phase, since it will distribute freely between the mobile and stationary solvent phases) minus the void volume V0.
However, for the molecular weight and/or Rs estimations in SEC, it is widely accepted to substitute the term Vt - Vo for Vi and use the partition coefficient (Kav):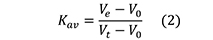
where,
Vt is the total volume of the packed bed column (Figure 4) and is measured by geometrically calculating the volume of a cylinder, (πr2h, where r and h are the radius and the length of the column, respectively, both expressed in cm).
Vt represents the sum of the total volume available to the mobile phase and the solid support volume, which is not accessible to solvent. Since the estimated volume of the stationary phase (Vt - Vo) will include the volume of the gel-forming substance, Kav is not a true distribution coefficient. However, Kav is directly proportional to Kd and, like Kd, defines solute behavior independently of the bed dimensions and packing (Figure 4) (Irvine, 2001 and GE Healthcare Handbook).
Figure 4. Gel filtration column parameters. A. Schematic representation of V0, Vt, and Vi, which are depicted as grey areas. B. Theoretical chromatogram for a gel filtration experiment with the summary of the various gel filtration parameters. - Calculate Kd for the standards and obtain a calibration curve by plotting Kd versus the logarithm of Rs (for Rs values see Table 1). An example of column calibration (Superdex 200 HR 10/300 GL prepacked column) obtained using a commercially available gel filtration calibration kit is shown in Figure 5. The calibration curve is prepared by plotting the Kd value for each standard versus its corresponding log10 Rs value. An essentially linear relationship should be obtained. Once the Kd value is calculated for the unknown sample, its Rs value can be determined from the calibration graph. Data were analyzed using Origin 8.0 (OriginLab Corporation, Northampton, MA).
Note: Plotting elution volumes vs. Rs for the standard proteins is usually adequate, and usually produces an approximately linear plot.
Figure 5. Superdex 200 HR 10/300 GL prepacked column (GE Healthcare) for size exclusion chromatography. Chromatographic separation and calibration curve for standard proteins. For this gel filtration analysis the mobile phase is 50 mM HEPES, 150 mM NaCl, 0.1 mM DTT pH 7.5. A. Overlap of chromatograms obtained from two runs (solid and dotted lines) of standard proteins. 1. Thyroglobulin; 2. Ferritin; 3 β-amylase; 4. Alcohol dehydrogenase; 5. Albumin bovine serum; 6. Carbonic anhydrase; 7. Cytochrome c. B. Calibration curve prepared by plotting the Kd value for each standard in (A) versus its corresponding log10 Rs value used to determine Rs of the protein sample. Calibration data are from (Astegno et al., 2015).
Practical example: determination of Ca2+-dependent changes in the Rs of the Ca2+-sensor protein CaM1 from Arabidopsis thaliana
SEC was used to determine Ca2+-induced variations in the hydrated shape of CaM1 from Arabidopsis thaliana (AtCaM1) (Astegno et al., 2016). A Superose 12 10/300 GL prepacked column was calibrated with standard proteins using 5 mM Tris, 150 mM KCl, pH 7.5 as the mobile phase (Figure 6A). The hydrodynamic behavior of AtCaM1 was then analyzed in the absence and presence of Ca2+ by adding 5 mM EGTA or 5 mM CaCl2, respectively, to the mobile phase and to the sample. Superposition of the chromatograms of AtCaM1 in the presence and the absence of Ca2+ (Figure 6B) indicates that the Ve of AtCaM1 is increased when the protein is in the Ca2+-bound form. The calibration curve was then used to evaluate the Rs of Ca2+-free and Ca2+-bound states of AtCaM1 and shows that the addition of Ca2+ to AtCaM1 is accompanied by a decrease in its Rs value (Figure 6C) as a consequence of the large conformational change that AtCaM1 undergoes upon Ca2+ binding. AtCaM1 is a monomer in both the apo and Ca2+-bound forms at the concentrations used in this study. The Rs values were derived from at least 3 replicates for each protein sample and mean ± SD were calculated.
Figure 6. Determination of Rs for apo- and Ca2+-bound CaM1 from Arabidopsis thaliana. The Rs difference between the Ca2+-free and Ca2+-bound states of AtCaM1 was determined using a Superose 12 column 10/300GL (GE Healthcare). A. Overlap of chromatograms obtained from two runs (solid and dotted lines) of standard proteins. 1. Albumin bovine serum; 2. Ovalbumin; 3. Carbonic anhydrase; 4. Myoglobin; 5. Cytochrome c. B. Elution profiles of AtCaM1 in the absence (apo-AtCaM1) and presence (Ca2+-AtCaM1) of Ca2+; C. Calibration curve prepared by plotting the Kd value for each standard in (A) versus its corresponding log10 Rs value. The graph was used to determine the Rs of AtCAM1 in apo- and Ca2+-bound conditions. The average Rs of apo- and Ca2+-AtCaM1 at pH 7.5 was 2.77 ± 0.03 nm and 2.53 ± 0.04 nm, respectively, and for both states results in molecular weight overestimation. Moreover, Ca2+ binding to AtCaM1 results in a decrease of Rs (~0.24 nm). Experimental data are from (Astegno et al., 2016).
Notes
- Instability of the column temperature negatively affects reproducibility of separation and column efficiency. We recommend that you keep the column at controlled temperature (usually 25 °C).
- Avoid extreme changes in pH or other conditions that can cause protein inactivation or even precipitation. If the sample precipitates in the SEC column, the column will be blocked, possibly irreversibly, and the sample might be lost.
- SEC columns must not run dry. Ensure that there is sufficient buffer for long runs. Columns that run dry must be repacked.
- Bubble formation on mixing of solvents can lead to many problems in FPLC analysis (e.g., appearance of spurious peaks, decrease in pumping reliability) which can be prevented by thoroughly degassing all solutions.
Recipes
- 2 M CaCl2 (20 ml)
Dissolve 5.88 g of CaCl2 in 20 ml of MilliQ water - 0.1 M EGTA (100 ml, pH 8)
Dissolve 3.8 g of EGTA in about 20 ml MilliQ water, bring to pH 11 with NaOH and then to pH 8.0 with HCl. Finally, add MilliQ water to a final volume of 100 ml - 0.1 M DTT
Dissolve 0.46 g of DTT (DL-dithiothreitol, anhydrous) in 25 ml of deionized or distilled H2O, adjust volume to 30 ml, dispense into aliquots, and store at -20 °C - Mobile phase
- Mobile phase marker
Dissolve acetone in the mobile phase at 5 mg/ml (0.63 % v/v). Centrifuge at 10,000 x g for 15 min immediately before column loading - Void volume marker
Prepare a solution of blue dextran (average MW 2,000,000 Da) by dissolving it at 1-2 mg/ml in the mobile phase. Centrifuge at 10,000 x g for 15 min immediately before column loading - Protein calibration standards
Solubilize each protein standard in the mobile phase at a concentration 1-2 mg/ml or as specified in the manufacturer’s instructions without heating or mixing vigorously. Centrifuge at 10,000 x g for 15 min immediately before column loading - Protein sample
Dissolve the protein sample in the mobile phase at 1 mg/ml (for determination of Ca2+-dependent change in Rs of Ca2+ sensor proteins add 5 mM EGTA or 5 mM CaCl2 to the sample)
Acknowledgments
This protocol was adapted from the previously published studies by Uversky, 1993; Sorensen and Shea, 1996; Astegno et al., 2016; Vallone et al., 2016.
References
- Asante-Appiah, E. and Skalka, A. M. (1997). A metal-induced conformational change and activation of HIV-1 integrase. J Biol Chem 272(26): 16196-16205.
- Astegno, A., Capitani, G. and Dominici, P. (2015). Functional roles of the hexamer organization of plant glutamate decarboxylase. Biochim Biophys Acta 1854(9): 1229-1237.
- Astegno, A., La Verde, V., Marino, V., Dell'Orco, D. and Dominici, P. (2016). Biochemical and biophysical characterization of a plant calmodulin: Role of the N- and C-lobes in calcium binding, conformational change, and target interaction. Biochim Biophys Acta 1864(3): 297-307.
- Astegno, A., Maresi, E., Marino, V., Dominici, P., Pedroni, M., Piccinelli, F. and Dell'Orco, D. (2014). Structural plasticity of calmodulin on the surface of CaF2 nanoparticles preserves its biological function. Nanoscale 6(24): 15037-15047.
- Bagai, I., Liu, W., Rensing, C., Blackburn, N. J. and McEvoy, M. M. (2007). Substrate-linked conformational change in the periplasmic component of a Cu(I)/Ag(I) efflux system. J Biol Chem 282(49): 35695-35702.
- De Angelis, F., Lee, J. K., O'Connell, J. D., 3rd, Miercke, L. J., Verschueren, K. H., Srinivasan, V., Bauvois, C., Govaerts, C., Robbins, R. A., Ruysschaert, J. M., Stroud, R. M. and Vandenbussche, G. (2010). Metal-induced conformational changes in ZneB suggest an active role of membrane fusion proteins in efflux resistance systems. Proc Natl Acad Sci U S A 107(24): 11038-11043.
- Ge Healthcare Life Science Handbook: Size Exclusion Chhromatography. Principles and Methods.
- Irvine, G. B. (2001). Determination of molecular size by size-exclusion chromatography (gel filtration). Curr Protoc Cell Biol Chapter 5: Unit 5 5.
- le Maire, M., Aggerbeck, L. P., Monteilhet, C., Andersen, J. P. and Møller, J. V. (1986). The use of high-performance liquid chromatography for the determination of size and molecular weight of proteins: a caution and a list of membrane proteins suitable as standards. Anal Biochem 154(2): 525-535.
- Ricker, R. D. and Sandoval, L. A. (1996). Fast, reproducible size-exclusion chromatography of biological macromolecules. J Chromatogr A 743(1): 43-50.
- Sorensen, B. R., Eppel, J. T. and Shea, M. A. (2001). Paramecium calmodulin mutants defective in ion channel regulation associate with melittin in the absence of calcium but require it for tertiary collapse. Biochemistry 40(4): 896-903.
- Sorensen, B. R. and Shea, M. A. (1996). Calcium binding decreases the stokes radius of calmodulin and mutants R74A, R90A, and R90G. Biophys J 71(6): 3407-3420.
- Vallone, R., La Verde, V., D'Onofrio, M., Giorgetti, A., Dominici, P. and Astegno, A. (2016). Metal binding affinity and structural properties of calmodulin-like protein 14 from Arabidopsis thaliana. Protein Sci 25(8): 1461-1471.
- Uversky, V. N. (1993). Use of fast protein size-exclusion liquid chromatography to study the unfolding of proteins which denature through the molten globule. Biochemistry 32(48):13288-98.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
La Verde, V., Dominici, P. and Astegno, A. (2017). Determination of Hydrodynamic Radius of Proteins by Size Exclusion Chromatography. Bio-protocol 7(8): e2230. DOI: 10.21769/BioProtoc.2230.
Category
Biochemistry > Protein > Structure
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










