- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Determination of Adeno-associated Virus Rep DNA Binding Using Fluorescence Anisotropy
Published: Vol 7, Iss 6, Mar 20, 2017 DOI: 10.21769/BioProtoc.2194 Views: 7870
Reviewed by: Yannick DebingAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
General Maintenance and Reactivation of iSLK Cell Lines
Ariana C. Calderón-Zavala [...] Ekaterina E. Heldwein
Jun 5, 2025 1933 Views
Inducible HIV-1 Reservoir Reduction Assay (HIVRRA), a Fast and Sensitive Assay to Test Cytotoxicity and Potency of Cure Strategies to Reduce the Replication-Competent HIV-1 Reservoir in Ex Vivo PBMCs
Jade Jansen [...] Neeltje A. Kootstra
Jul 20, 2025 2489 Views
Assembly and Mutagenesis of Human Coronavirus OC43 Genomes in Yeast via Transformation-Associated Recombination
Brett A. Duguay and Craig McCormick
Aug 20, 2025 3071 Views
Abstract
Quantitative measurement of proteins binding to DNA is a requisite to fully characterize the structural determinants of complex formation necessary to understand the DNA transactions that regulate cellular processes. Here we describe a detailed protocol to measure binding affinity of the adeno-associated virus (AAV) Rep68 protein for the integration site AAVS1 using fluorescent anisotropy. This protocol can be used to measure the binding constants of any DNA binding protein provided the substrate DNA is fluorescently labeled.
Keywords: Adeno-associated virusBackground
Fluorescence polarization anisotropy has become one of the most popular methods to measure the interaction of proteins with a large variety of ligands including small molecules, nucleic acids, peptides and other proteins. The method is quick, inexpensive and can be modified to be used in plate readers equipped with fluorescence detectors. The technique is based on the principle that when a fluorescent molecule is excited with plane polarized light, the emitted light remains polarized in the same plane if the molecule is stationary or if it rotates slowly. In contrast, if the molecule rotates rapidly (due to small size), the light is emitted in a different plane. These changes can be quantified by the normalized differences in parallel and perpendicular intensities. Polarization is defined as P = (I= - I⊥)/(I= + I⊥), where I= is the parallel intensity and I⊥ is the perpendicular intensity. An alternative way is to define the anisotropy, A = (I= - I⊥)/(I= + 2I⊥). Both parameters can be used interchangeably to describe the changes in polarization. Thus, when a small fluorescent DNA molecule binds a protein, the larger complex will rotate more slowly than the DNA molecule, changing the plane of the polarized light and increasing the anisotropy value. We have used this technique to measure the binding affinity of AAV Rep68 for different DNA substrates (Yoon-Robarts et al., 2004; Musayev et al., 2015; Bardelli et al., 2016). The non-structural AAV Rep proteins carry out most of the DNA transactions that are required to complete the virus life cycle. These include DNA replication, transcriptional regulation, site-specific integration and packaging of DNA into preformed capsids (Im and Muzyczka, 1990; Weitzman et al., 1994; Wonderling et al., 1995). The large AAV Rep proteins (Rep78/Rep68) contain an N-terminal origin binding domain (OBD) that specifically binds the Rep binding sites (RBS) and displays nuclease activity (Hickman et al., 2004; Musayev et al., 2015). The RBS sites consist of two or more 5’-GCTC-3’ repeats and are found at the viral origin of replication, in several promoters and at the AAVS1 integration site (Weitzman et al., 1994; McCarty et al., 1994). In addition, a C-terminus SF3 helicase domain is required for high affinity binding and DNA unwinding (James et al., 2003; Mansilla-Soto et al., 2009).The protocol described here can be modified to fit any protein-DNA system or any other instrument such as plate readers.
Materials and Reagents
- Pipette tips
- 15 ml conical tubes (USA scientific, catalog number: 1475-1611 )
- Black 1.5 ml Eppendorf tubes (Argos Technologies, catalog number: T7456-001 )
- 16 gauge needle (BD, catalog number: 305197 )
- Fluorescein labeled AAVS1 sense DNA strand with the following sequence:
5’-TGGCGGCGGTTGGGGCTCGGCGCTCGCTCGCTCGCTGGGCG-3’ - AAVS1 anti-sense strand with the sequence: 5’-CGCCCAGCGAGCGAGCGA GCGCCGAGCCCCAACCGCCGCCA-3’
Note: DNA can be synthesized using any synthesis facility such as IDT services at a 100 nanomole scale. The sense strand can be labeled at the 5’ end with fluorescein (6-FAM). - Purified recombinant AAV Rep68 protein was expressed in E.coli and purified using Ni-NTA affinity column followed by a gel filtration column as described previously (Musayev et al., 2015)
- Sodium hydroxide (NaOH)
- Sodium chloride (NaCl) (Fisher Scientific, catalog number: BP358 )
- 2-Amino-2-(hydroxymethyl)propane-1,3-diol (Tris) (Sigma-Aldrich, catalog number: T1503 )
- Ethylendiaminetetraacetic acid (EDTA) (Gold Bio, catalog number: E-210-1 )
- 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES) (Gold Bio, catalog number: H-401-500 )
- Tris(2-carboxyethyl)phosphine (TCEP) (Gold Bio, catalog number: TECP )
- Double distilled water
- Q1 buffer (see Recipes)
- Q2 buffer (see Recipes)
- TES buffer (see Recipes)
- Binding buffer (see Recipes)
Equipment
- Pipettes
- MonoQ anion-exchange column (GE Healthcare, catalog number: 17-5166-01 )
- HiTrap 5 ml desalting column (GE Healthcare, catalog number: 11-0003-29 )
- GE Healthcare AKTA purifier
- Labconco Freezone 2/5 Benchtop lyophilizer
- Thermo Scientific NanoDrop ND-2000c spectrophotometer (Thermo Fisher Scientific, model: NanoDropTM 2000/2000c )
- Denville IncuBlock heating block
- ISS PC1 fluorimeter (ISS, model: PC1TM )
Software
- Microsoft Excel
- GraphPad Prism 7TM
- Vinci Instrument control and data acquisition software from ISS
Procedure
- Preparation of fluorescein labeled AAVS1 DNA
- Fluorescent labeled DNA can be purified by HPLC or using preparative acrylamide gel electrophoresis (PAGE) directly by the synthesis facility or can be purified by anion-exchange chromatography as described next. DNA is dissolved in 500 μl of Q1 buffer and injected into a MonoQ anion-exchange column. DNA was purified using a linear gradient from buffer Q1 (100 mM NaCl) to 100% Q2 buffer (1 M NaCl).
- Fractions are desalted using a 5 ml GE HiTrap desalting column pre-equilibrated with water. DNA elutes in the void volume.
- Collected DNA fractions are placed in a 15 ml conical tube, frozen and placed in a Labcono Freezone 2.5 lyophilizer overnight. To allow drying of the sample, the lid of the 15 ml conical tube should be pierced with a needle 4-5 times. The DNA should be fibrous and dry by the next day.
- After lyophilization, resuspend the DNA in 100 μl of TES buffer. Measure the DNA concentrations using a Thermo Scientific NanoDrop ND-2000c spectrophotometer using the calculated extinction coefficients of the DNA oligonucleotides at a wavelength of 260 nm.
- To prepare the double-stranded DNA, combine the two DNA strands in a ratio where the non-labeled strand is in 1.1 molar excess with respect to the fluorescein-labeled strand. Place the DNA strands in a black 1.5 ml microcentrifuge tube. Using a Denville IncuBlock heating block, heat the DNA at 99 °C for 3 min. Then, turn off the heating block and leave the DNA cooling to room temperature.
- The final DNA concentration is calculated using the number of moles of the labeled-DNA strand and the final volume after mixing the two strands.
- Fluorescent labeled DNA can be purified by HPLC or using preparative acrylamide gel electrophoresis (PAGE) directly by the synthesis facility or can be purified by anion-exchange chromatography as described next. DNA is dissolved in 500 μl of Q1 buffer and injected into a MonoQ anion-exchange column. DNA was purified using a linear gradient from buffer Q1 (100 mM NaCl) to 100% Q2 buffer (1 M NaCl).
- DNA binding studies with Rep68
Rep68 concentration is determined by absorbance at 280 nm using the extinction coefficient calculated from the Rep68 amino acid sequence using EXPASY Protparam site (http://web.expasy.org/protparam/). Prepare different concentrations of Rep68 in binding buffer and mix with DNA allowing the binding reaction to reach equilibrium at 20 °C for 20-30 min. For other proteins, a preliminary experiment needs to be performed to obtain a rough estimate of the concentrations to be used. Titrations are carried out using a final 5 nM DNA concentration in binding buffer. A typical binding experiment is performed as follows:- It is recommended that the PC-1 fluorimeter is turned on the night before the experiment to stabilize the instrument (Figure 1). Start the Vinci software on the computer. Access instrument control and set the excitation and emission filters to 492 nm and 528 nm, respectively.
- Turn-on the lamp of the PC1 fluorimeter and let it warm up for at least 1 h.
- Calculate the amount of Rep68 stock solution needed to achieve the following final concentrations in the binding reactions: 0, 10, 25, 50, 75, 100, 150, 200, 300, 400 and 500 nM.
Generally, three replicates are done for each concentration. - Mix the different Rep68 concentrations with 5 nM DNA in a final volume of 300 μl. Incubate samples for 20-30 min away from light.
- Measure the anisotropy value of each concentration point using the ‘Single point Polarization’ function in the Vinci software and record 10 measurements per concentration.

Figure 1. PC1 fluorimeter from ISS
- It is recommended that the PC-1 fluorimeter is turned on the night before the experiment to stabilize the instrument (Figure 1). Start the Vinci software on the computer. Access instrument control and set the excitation and emission filters to 492 nm and 528 nm, respectively.
Data analysis
- Open all the data in Microsoft Excel and for each concentration calculate the average anisotropy value from the 10 measurements.
- Make a new Graphpad project using the Rep68 concentration as the x values and the average anisotropy as the y value.
- Subtract the DNA anisotropy value from all concentrations.
- Using Prism 7TM (GraphPad), perform a nonlinear fitting using a single specific binding model. Prism uses the following equation:
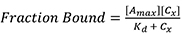
Where,
Amax is the maximum anisotropy value at saturation,
Cx is the concentration of Rep68,
Kd is binding constant. - To calculate the fraction of DNA bound, the maximum value obtained from the fitting is used as the anisotropy value at saturation when all DNA is bound (Amax). The fraction of DNA bound at each concentration is the anisotropy at each concentration (Ax) divided by Amax. This is shown in Figure 2A. Anisotropy values (Ax) at each concentration (Group A) were converted to fraction bound (Group B).
- Below is an example of a typical experiment.
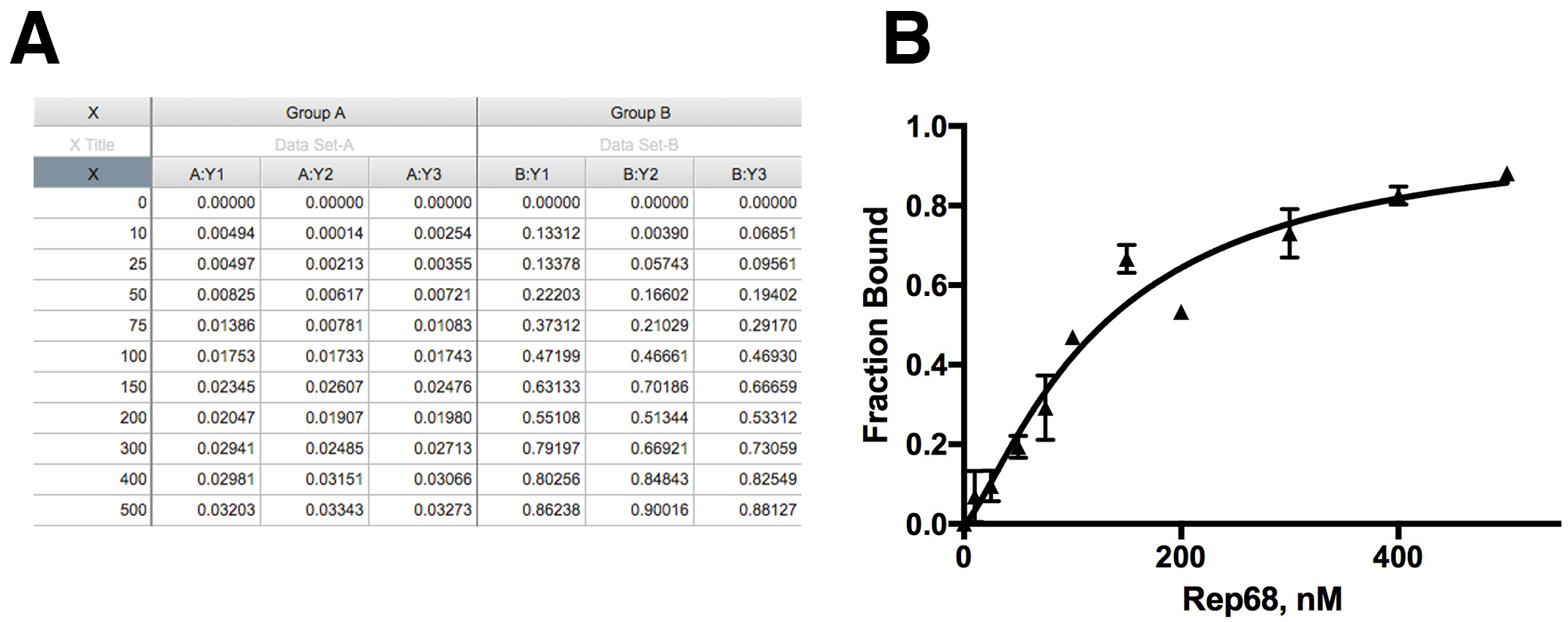
Figure 2. Data analysis. A. Anisotropy data and correction for fraction bound; B. Nonlinear Fitting of the data to specific binding model.
Notes
- It is recommended that during the incubation step, samples be kept away from light and incubated at the temperature of the instrument by means of a water bath or heating/cooling block.
- Protein stocks are stored at high concentration (~0.1-0.5 mM) at -80 °C and are diluted in binding buffer to the required concentrations. Generally, a series of dilutions from 100 μM to 100 nM are prepared to cover the concentration range of the titration curve.
- If incubation time is not known, this can be determined by incubating sample and reading anisotropy every five minutes. Equilibrium is estimated when the measured anisotropy remains constant.
Recipes
- Q1 buffer
10 mM NaOH
100 mM NaCl (pH 12.0) - Q2 buffer
10 mM NaOH
1 M NaCl (pH 12.0) - TES buffer
10 mM Tris
100 mM NaCl
1 mM EDTA (pH 8.0) - Binding buffer
25 mM HEPES
200 mM NaCl
1 mM TCEP (pH 7.0)
Acknowledgments
This work was supported by NIH grant R01-GM092854. The authors declare no conflict of interests.
References
- Bardelli, M., Zarate-Perez, F., Agundez, L., Linden, R. M., Escalante, C. R. and Henckaerts, E. (2016). Identification of a functionally relevant AAV Rep68 oligomeric interface. J Virol 90(15): 6612-24.
- Hickman, A. B., Ronning, D. R., Perez, Z. N., Kotin, R. M. and Dyda, F. (2004). The nuclease domain of adeno-associated virus rep coordinates replication initiation using two distinct DNA recognition interfaces. Mol Cell 13(3): 403-414.
- Im, D. S. and Muzyczka, N. (1990). The AAV origin binding protein Rep68 is an ATP-dependent site-specific endonuclease with DNA helicase activity. Cell 61(3): 447-457.
- James, J. A., Escalante, C. R., Yoon-Robarts, M., Edwards, T. A., Linden, R. M. and Aggarwal, A. K. (2003). Crystal structure of the SF3 helicase from adeno-associated virus type 2. Structure 11(8): 1025-1035.
- Mansilla-Soto, J., Yoon-Robarts, M., Rice, W. J., Arya, S., Escalante, C. R. and Linden, R. M. (2009). DNA structure modulates the oligomerization properties of the AAV initiator protein Rep68. PLoS Pathog 5(7): e1000513.
- McCarty, D. M., Pereira, D. J., Zolotukhin, I., Zhou, X., Ryan, J. H. and Muzyczka, N. (1994). Identification of linear DNA sequences that specifically bind the adeno-associated virus Rep protein. J Virol 68(8): 4988-4997.
- Musayev, F. N., Zarate-Perez, F., Bishop, C., Burgner, J. W., 2nd and Escalante, C. R. (2015). Structural insights into the assembly of the adeno-associated virus type 2 Rep68 protein on the integration site AAVS1. J Biol Chem 290(46): 27487-27499.
- Weitzman, M. D., Kyostio, S. R., Kotin, R. M. and Owens, R. A. (1994). Adeno-associated virus (AAV) Rep proteins mediate complex formation between AAV DNA and its integration site in human DNA. Proc Natl Acad Sci U S A 91(13): 5808-5812.
- Wonderling, R. S., Kyostio, S. R. and Owens, R. A. (1995). A maltose-binding protein/adeno-associated virus Rep68 fusion protein has DNA-RNA helicase and ATPase activities. J Virol 69(6): 3542-3548.
- Yoon-Robarts, M., Blouin, A. G, Bleker, S., Kleinmschmidt, J. A., Aggarwal, A. K., Escalante, C. R. and Linden, R. M. (2004). Residues within the B’ motif are critical for DNA binding by the superfamily 3 helicase Rep40 of adeno-associated virus type 2. J Biol Chem 279(48): 50472-50481.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Zarate-Perez, F., Santosh, V., Bardelli, M., Agundez, L., Linden, R. M., Henckaerts, E. and Escalante, C. R. (2017). Determination of Adeno-associated Virus Rep DNA Binding Using Fluorescence Anisotropy. Bio-protocol 7(6): e2194. DOI: 10.21769/BioProtoc.2194.
Category
Microbiology > Microbe-host interactions > Virus
Biochemistry > Protein > Activity
Molecular Biology > Protein > Protein-DNA binding
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











