- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Two-electrode Voltage-clamp Recordings in Xenopus laevis Oocytes:Reconstitution of Abscisic Acid Activation of SLAC1 Anion Channel via PYL9 ABA Receptor
Published: Vol 7, Iss 2, Jan 20, 2017 DOI: 10.21769/BioProtoc.2114 Views: 12459
Reviewed by: Marisa RosaKathrin SutterAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
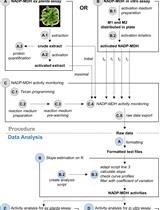
A Semi-throughput Procedure for Assaying Plant NADP-malate Dehydrogenase Activity Using a Plate Reader
Kevin Baudry and Emmanuelle Issakidis-Bourguet
Aug 20, 2023 1496 Views
An in vitro Assay to Probe the Formation of Biomolecular Condensates
Yu Zhang and Shen Lisha
Sep 5, 2023 3237 Views
Immunofluorescence for Detection of TOR Kinase Activity In Situ in Photosynthetic Organisms
Ana P. Lando [...] Giselle M. A. Martínez-Noël
Dec 20, 2024 1846 Views
Abstract
Two-Electrode Voltage-Clamp (TEVC) recording in Xenopus laevis oocytes provides a powerful method to investigate the functions and regulation of ion channel proteins. This approach provides a well-known tool to characterize ion channels or transporters expressed in Xenopus laevis oocytes. The plasma membrane of the oocyte is impaled by two microelectrodes, one for voltage sensing and the other one for current injection. Here we list a protocol that allows robust reconstitution of multi-component signaling pathways. This protocol has been used to study plant ion channels, including the SLAC1 channel (SLOW ANION CHANNEL-ASSOCIATED 1), in particular SLAC1 activation by either the protein kinase OST1 (OPEN STOMATA 1), Ca2+-dependent protein kinases (CPKs) or the GHR1 (GUARD CELL HYDROGEN PEROXIDE-RESISTANT 1) transmembrane receptor-like protein. Data are presented showing reconstitution of abscisic acid activation of the SLAC1 anion channel by the ‘monomeric’ ABA (abscisic acid) receptor RCAR1/PYL9 (PYRABACT INRESISTANCE1 [PYR1]/PYR1-LIKE [PYL]/REGULATORYCOMPONENTS OF ABA RECEPTORS [RCAR]) by co-expressing four components of the abscisic acid signaling core. This protocol is also suitable for studying other ion channel functions and regulation mechanisms, as well as transporter proteins.
Keywords: Ion channelBackground
Ion channels expressed in Xenopus laevis oocytes can be studied using two-electrode voltage-clamping. This protocol provides a method to measure ion channel or transporter currents expressed in oocytes, including plant ion channels. In this protocol, we not only summarize how to prepare cRNA, isolate oocytes, inject cRNA and record currents, but also provide information on how to succeed in completing experiments upon co-expressing a signal transduction cascade from receptor to ion channel.
Materials and Reagents
- Borosilicate glass capillaries (World Precision Instruments, catalog number: 1B100F-4 )
- Parafilm (Sigma-Aldrich, catalog number: P7793-1EA )
- Xenopus laevis oocytes (Ecocyte Bioscience, catalog number: 0-100-2 )
- Vector: pNB1 oocyte expression vector harboring the cDNA of interest using the USER method (Nour-Eldin et al., 2006), or other oocytes expression vector like
- mMESSAGE mMACHINE® T7 Kit (Thermo Fisher Scientific, AmbionTM, catalog number: AM1344 )
- Collagenase D (Roche Diagnostics, catalog number: 11088882001 )
- Mineral oil (Sigma-Aldrich, catalog number: M5904 )
- MES hydrate (Sigma-Aldrich, catalog number: M2933 )
- Tris-base (Thermo Fisher Scientific, Fisher Scientific, catalog number: BP152-5 )
- Calcium chloride (CaCl2) (Sigma-Aldrich, catalog number: C5670 )
- Magnesium chloride (MgCl2) (Sigma-Aldrich, catalog number: M8266 )
- Sodium chloride (NaCl) (Thermo Fisher Scientific, Fisher Scientific, catalog number: S271-10 )
- Potassium chloride (KCl) (Sigma-Aldrich, catalog number: P9333 )
- Na-gluconate (Sigma-Aldrich, catalog number: S2054 )
- D-sorbitol (Sigma-Aldrich, catalog number: S1876 )
- Gentamicin solution (Sigma-Aldrich, catalog number: G1272 )
- ND96 buffer (see Recipes)
- Recording buffer (see Recipes)
Equipment
- Two-electrode voltage clamp amplifier (e.g., Warner Instrument, model: Oocyte Clamp OC-725C )
- Digidata 1440A low-noise data acquisition system (Molecular Devices, model: Digidata 1440A)
- P-87 flaming/brown microelectrode micropipette puller (Sutter Instrument, model: P-87)
- Osmometer (e.g., Wescor, model: Vapor Pressure Osmometer 5500 )
- Microdispenser (Drummond Scientific, catalog number: 3-000-510 )
- Custom glass tubing (Drummond Scientific, catalog number: 3-000-210-G8 )
Software
- pCLAMP 10 Electrophysiology Data Acquisition and Analysis Software (Molecular Devices).
Procedure
- Prepare cRNA
- All constructs were cloned into the pNB1 oocyte expression vector using the USER method (Nour-Eldin et al., 2006).
- cRNAs were synthesized from 0.5-1 μg of linearized plasmid DNA template using the mMESSAGE mMACHINE® T7 Kit from Thermo Fisher Scientific.
- Isolation of oocytes
- Individual Xenopus laevis oocytes can be ordered from Ecocyte Bioscience and arrives the next day on dry ice. Alternatively, ovary lobes can be surgically extracted as described in previous reports (Miledi, 1982; Gundersen et al., 1983; Stühmer and Parekh, 1995).
- To isolate oocytes from ovary lobes, wash ovary lobes with ND96 buffer 3 times, incubate it in ND96 buffer at 16 °C overnight.
- Place 2-3 ovary lobes in ND96 buffer containing 30 mg/ml collagenase D, shake 2-3 h to remove follicle cell layer at room temperature (~23 °C) (Figure 1A).
- Wash oocytes with ND96 buffer 5 times, sort stage IV and V oocytes (approximate diameter 1 mm, Wasserman et al., 1984) in ND96 buffer. Incubate sorted oocytes in ND96 buffer at 16 °C overnight before injecting cRNA (Figure 1B).
- Injection of cRNA
- Use stage IV and V oocytes, for which the follicle cell layer has been removed (Wasserman et al., 1984). Inject more than 30 oocytes with each mRNA combination that is to be investigated.
- Pull injection glass needles on P-87 flaming/brown microelectrode micropipette puller using borosilicate glass capillaries to produce injection needles (Figure 1C).
- Break off the needle tip with forceps so that it is easier to inject cRNA into oocytes (Figure 1D).
- Fill pipette tip with mineral oil to about one-third to two-thirds.
- Mount pipette to the microdispenser.
- Deposit cRNA sample onto Parafilm, to a total volume of cRNA > 1 μl.
- Fill the pipette with RNA solution with microdispenser by applying negative pressure to the pipette.
- Inject cRNA into selected oocytes. The volume of cRNA injected into oocytes is about 50 nl (oocyte volume ~500 nl). The concentration of injected cRNA is better more than 2 ng/μl (Figure 1E).
- Incubate oocytes in ND96 buffer at 16 °C for 2-3 days before recording currents.

Figure 1. Oocytes and glass needle at different steps. A. Oocyte ovary lobes; B. Oocytes after isolation; C. Injection glass needle before the tip is ‘broken’; D. ‘Broken’ needle; E. An oocyte being injected with cRNA. Note that injection pipette is out of focus. F. An oocyte impaled with two electrodes for voltage clamping.
- Recording
- Pull recording glass electrodes on P-87 flaming/brown microelectrode micropipette puller using custom glass tubing.
- In some case, slightly break the needle tip so that it is easier to insert the micropipette into oocytes since needle tips can be too soft for multiple sequential injections into oocytes. Alternatively, press the electrode against the oocyte and tap the recording table. Vibrations aid in impaling the oocyte.
- Fill the micropipette with 3 M KCl. Place one electrode into each of the two holders, making sure that Ag/AgCl electrode wire contacts the KCl solution in the micropipette. The resistances of the filled electrodes were 0.5-1.5 M (Figure 1F).
- For anion channel recordings, steady state currents are recorded starting from a holding potential of 0 mV and ranging from +40 to -160 mV in -20 mV decrements, followed by a -120 mV voltage ‘tail’ pulse (Figure 2).
Note: The time-dependent properties of SLAC1 channel currents in Xenopus laevis oocytes vary among individual oocytes. This may depend on posttranslational modification of the channel protein. This property is also known from guard cell recordings of the SLAC1-encoding S-type anion channels (Schmidt and Schroeder, 1994).
Figure 2. OST1 activates SLAC1 anion channel currents in Xenopus laevis oocytes. A. The voltage protocol used for recording SLAC1 anion channel currents. B and C. Example of whole-cell current traces recorded from oocytes injected with cRNA of (B) SLAC1yc only and (C) SLAC1yc+OST1yn (in SLAC1yc, SLAC1 is tagged with the C-terminal ‘half’ of YFP and in OST1yn, OST1 is tagged with N-terminal ‘half’ of YFP. Constructs for similar experiments see Geiger et al., 2009; Lee et al., 2009; Hua et al., 2012; Brandt et al., 2015; Wang et al., 2016). - To assess whether ‘monomeric’ abscisic acid ABA receptor PYL9 (Dupeux et al., 2011; Hao et al., 2011) activation of SLAC1 can be reconstituted, we co-injected cRNAs of the SLAC1yc channel, ABI1 (ABA-INSENSITIVE1) protein phosphatase, the PYL9 receptor, and the OST1yn protein kinase into oocytes (Geiger et al., 2009; Brandt et al., 2012). Without ABA, PYL9 did not activate the anion channel currents in oocytes expressing the ABI1 with SLAC1yc and OST1yn. When ABA was injected into oocytes 30 min before measuring of oocyte currents, SLAC1 anion channels activity was dramatically enhanced in oocytes expressing SLAC1yc, OST1yn, PYL9 and ABI1 (Figure 3). Thus, the ‘monomeric’ ABA receptor PYL9 enables ABA signaling reconstitution in oocytes.
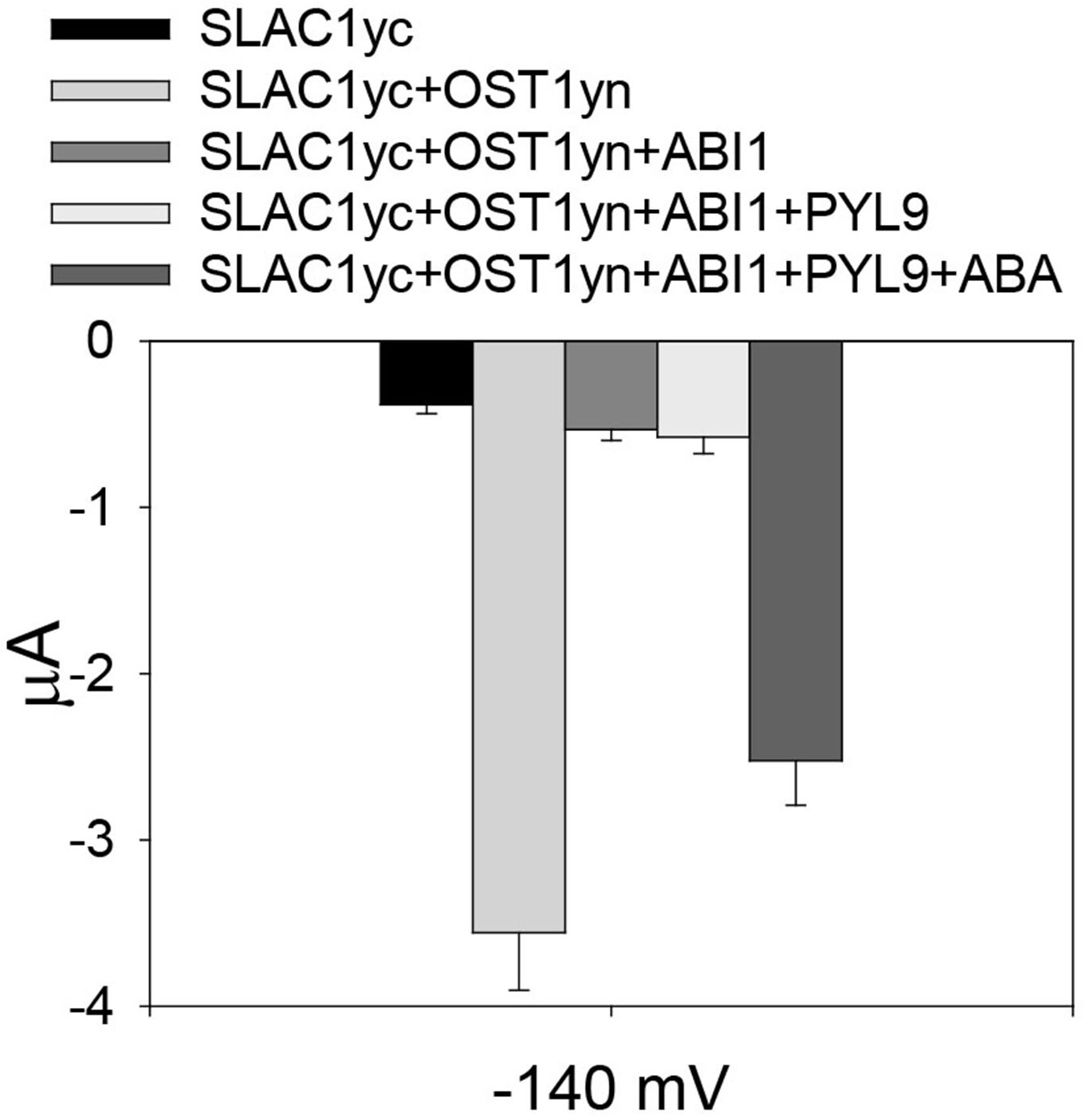
Figure 3. Reconstitution of ABA activation of SLAC1 anion channels by PYL9 ABA receptor. Average current of SLAC1 anion channels recorded at -140 mV. In the absence of ABA, the ABA receptor PYL9 is unable to enhance SLAC1 anion channels currents. However, in the presence of injected ABA, SLAC1 anion channels currents are greatly increased. Data are mean ± SEM (SLACyc, n = 9; SLAC1yc+OST1yn, n = 13; SLAC1yc+OST1yn+ABI1, n = 10; SLAC1yc+OST1yn+ABI1+PYL9, n = 12; SLAC1yc+OST1yn+ABI1+PYL9+ABA, n = 15).
Recipes
- ND96 buffer
10 mM MES/Tris (pH 7.5)
1 mM CaCl2
1 mM MgCl2
96 mM NaCl
Osmolality is adjusted to 220 mM using D-sorbitol (Geiger et al., 2009; Wang et al., 2016) - Recording buffer
10 mM MES/Tris (pH 7.5)
1 mM MgCl2
1 mM CaCl2
2 mM KCl
24 mM NaCl
70 mM Na-gluconate
Osmolality is adjusted to 220 mM using D-sorbitol (Geiger et al., 2009; Wang et al., 2016)
Acknowledgments
This research was surpported by grants from the National Institutes of Health (GM060396) and the National Science Foundation (MCB1616236) to J.I.S.
References
- Brandt, B., Brodsky, D. E., Xue, S., Negi, J., Iba, K., Kangasjarvi, J., Ghassemian, M., Stephan, A. B., Hu, H. and Schroeder, J. I. (2012). Reconstitution of abscisic acid activation of SLAC1 anion channel by CPK6 and OST1 kinases and branched ABI1 PP2C phosphatase action. Proc Natl Acad Sci USA 109(26): 10593-10598.
- Brandt, B., Munemasa, S., Wang, C., Nguyen, D., Yong, T., Yang, P. G., Poretsky, E., Belknap, T. F., Waadt, R., Aleman, F. and Schroeder, J. I. (2015). Calcium specificity signaling mechanisms in abscisic acid signal transduction in Arabidopsis guard cells. Elife 4: e03599.
- Dupeux, F., Santiago, J., Betz, K., Twycross, J., Park, S. Y., Rodriguez, L., Gonzalez-Guzman, M., Jensen, M. R., Krasnogor, N., Blackledge, M., Holdsworth, M., Cutler, S. R., Rodriguez, P. L. and Marquez, J. A. (2011). A thermodynamic switch modulates abscisic acid receptor sensitivity. EMBO J 30(20): 4171-4184.
- Geiger, D., Scherzer, S., Mumm, P., Stange, A., Marten, I., Bauer, H., Ache, P., Matschi, S., Liese, A., Al-Rasheid, K. A., Romeis, T. and Hedrich, R. (2009). Activity of guard cell anion channel SLAC1 is controlled by drought-stress signaling kinase-phosphatase pair. Proc Natl Acad Sci USA 106(50): 21425-21430.
- Gundersen, C. B., Miledi, R. and Parker, I. (1983). Voltage-operated channels induced by foreign messenger RNA in Xenopus oocytes. Proc R Soc Lond B Biol Sci 220(1218): 131-140.
- Hao, Q., Yin, P., Li, W., Wang, L., Yan, C., Lin, Z., Wu, J. Z., Wang, J., Yan, S. F. and Yan, N. (2011). The molecular basis of ABA-independent inhibition of PP2Cs by a subclass of PYL proteins. Mol Cell 42(5): 662-672.
- Hua, D., Wang, C., He, J., Liao, H., Duan, Y., Zhu, Z., Guo, Y., Chen, Z. and Gong, Z. (2012). A plasma membrane receptor kinase, GHR1, mediates abscisic acid- and hydrogen peroxide-regulated stomatal movement in Arabidopsis. Plant Cell 24(6): 2546-2561
- Lee, S. C., Lan, W., Buchanan, B. B. and Luan, S. (2009). A protein kinase-phosphatase pair interacts with an ion channel to regulate ABA signaling in plant guard cells. Proc Natl Acad Sci U S A 106(50): 21419-21424.
- Miledi, R. (1982). A calcium-dependent transient outward current in Xenopus laevis oocytes. Proc R Soc Lond B Biol Sci 215(1201): 491-497.
- Nour-Eldin, H. H., Hansen, B. G., Norholm, M. H., Jensen, J. K. and Halkier, B. A. (2006). Advancing uracil-excision based cloning towards an ideal technique for cloning PCR fragments. Nucleic Acids Res 34(18): e122.
- Schmidt, C. and Schroeder, J. I. (1994). Anion selectivity of slow anion channels in the plasma membrane of guard cells (large nitrate permeability). Plant Physiol 106(1): 383-391.
- Stühmer, W. and Parekh, A. B. (1995). Electrophysiological recordings from Xenopus oocytes. In: Bert S. and Erwin N. (Eds.). Single-Channel Recording. Springer pp: 341-356.
- Wang, C., Hu, H., Qin, X., Zeise, B., Xu, D., Rappel, W. J., Boron, W. F. and Schroeder, J. I. (2016). Reconstitution of CO2 regulation of SLAC1 anion channel and function of CO2-permeable PIP2;1aquaporin as CARBONIC ANHYDRASE4 interactor. Plant Cell 28(2): 568-582.
- Wasserman, W. J., Houle, J. G. and Samuel, D. (1984). The maturation response of stage IV, V, and VI Xenopus oocytes to progesterone stimulation in vitro. Dev Biol 105(2): 315-324.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Wang, C., Zhang, J. and Schroeder, J. I. (2017). Two-electrode Voltage-clamp Recordings in Xenopus laevis Oocytes:Reconstitution of Abscisic Acid Activation of SLAC1 Anion Channel via PYL9 ABA Receptor. Bio-protocol 7(2): e2114. DOI: 10.21769/BioProtoc.2114.
Category
Plant Science > Plant biochemistry > Protein > Activity
Cell Biology > Cell-based analysis > Ion analysis
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link












