- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Assessment of Wheat Resistance to Fusarium graminearum by Automated Image Analysis of Detached Leaves Assay
Published: Vol 6, Iss 24, Dec 20, 2016 DOI: 10.21769/BioProtoc.2065 Views: 12784
Reviewed by: Arsalan DaudiMalou FraitureBaohua Li

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Quantification of the Composition Dynamics of a Maize Root-associated Simplified Bacterial Community and Evaluation of Its Biological Control Effect
Ben Niu and Roberto Kolter
Jun 20, 2018 10538 Views
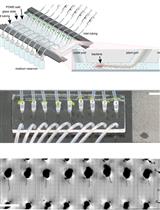

Tracking Root Interactions System (TRIS) Experiment and Quality Control
Hassan Massalha [...] Asaph Aharoni
Apr 20, 2019 7373 Views

A Quick Method for Screening Biocontrol Efficacy of Bacterial Isolates against Bacterial Wilt Pathogen Ralstonia solanacearum in Tomato
Heena Agarwal [...] Niraj Agarwala
Nov 20, 2020 5592 Views
Abstract
Fusarium head blight (FHB) caused by Fusarium pathogens is a globally important cereal disease. To study Fusarium pathogenicity and host disease resistance, robust methods for disease assessment and quantification are needed. Here we describe the procedure of a detached leaves assay emphasizing the image analysis. The protocol provides the different steps of a rapid, automatic and quantitative image analysis to evaluate leaf area infected by Fusarium graminearum.
Keywords: Detached leaf assayBackground
Evaluation of wheat FHB resistance at the whole plant level is estimated by visual scoring at flowering stage which is laborious, time consuming and requires space. Therefore in vitro methods that expedites disease assessment for FHB resistance at early plant stage have been developed, such as seed germination assay (Browne, 2009), coleoptiles assay (Shin et al., 2014), detached leaf assay (Browne and Cooke, 2004) and seedling assay (Li et al., 2010). Detached leaves assay is commonly used to assess host responses to Fusarium and was successful in identifying components of FHB resistance (Browne and Cooke, 2004). In such assays pathogen establishment is visually assessed which is time-consuming and limits accurate measurement. The method described recently by Perochon et al. (2015) and detailed here resolved these two limitations by using an automatic method that quantifies leaf area infected by image analysis based on particle size.
Materials and Reagents
- Petri dish, triple vent 94 x 15mm (Greiner Bio One, catalog number: 633 185 )
- Filter paper (Whatman, catalog number: 1001-090 )
- Parafilm (Parafilm, catalog number: PM992 )
- Plant container pots, 3 L (National Agrochemical Distributors, catalog number: POTS34 )
- John Innes compost No. 2 (Westland Horticulture)
- Square Petri dish 100 x 100 x 20 mm (SARSTEDT, catalog number: 82.9923.422 )
- Glass Pasteur pipette (VWR, catalog number: 14673-010 )
- Fusarium graminearum strain GZ3639 (Proctor et al., 1995)
- Plant agar (Duchefa, catalog number: P1001.1000 )
- Benzimidazole (Stock solution at 500 mM in ethanol stored in aliquots at -20 °C) (Sigma-Aldrich, catalog number: 194123 )
- Ethanol (Sigma-Aldrich, catalog number: E7023 )
- Tween-20 (Sigma-Aldrich, catalog number: P2287 )
Equipment
- Incubator (20 °C)
- Glasshouse (Cambridge HOK production glasshouse) (20-22 °C with a 16 h light/8 h dark photoperiod at 300 μmol m-2 s-1 and 70% relative humidity)
- Plant growth room (20 °C with a 16 h light/8 h dark photoperiod at 200 μmol m-2 s-1 and 70% relative humidity)
- Digital camera (Nikon, model: COOLPIX P500 with a NIKKOR 36X wide optical ZOOM ED VR, 4.0 - 144 mm, 1:3.4 - 5.7)
- Copy stand (RPS Studio RS-CS920 copy stand)
- Lights (cold-light fluorescent lighting system)
Software
- Fiji (http://fiji.sc/)
Note: Fiji is an open source software using the same functionality as ImageJ with many bundled plugins. For that reason the image analysis presented here could be executed with ImageJ.
Procedure
- Place wheat seeds in a 94 mm diameter Petri dish plate containing 2 filter papers and 6 ml of sterile water.
- Seal the plates with a piece of Parafilm and germinate in the dark for 3 days at 20 °C in an incubator.
- Transplant seedlings to 3 L pots containing moistened John Innes compost No 2 with maximum 6 seedlings/pot.
- Grow the plants in the glasshouse under controlled environment conditions at 20-22 °C with a 16 h light/8 h dark photoperiod at 300 μmol m-2 s-1 and 70% relative humidity.
- After approximately 20 days at the 3-leaf-stage (growth stage 13; [Zadoks et al., 1974]), cut an 8 cm section from the second leaf at a fixed distance from the leaf base (Figure 1A).
Note: 8 leaf sections fit in the square Petri dish used.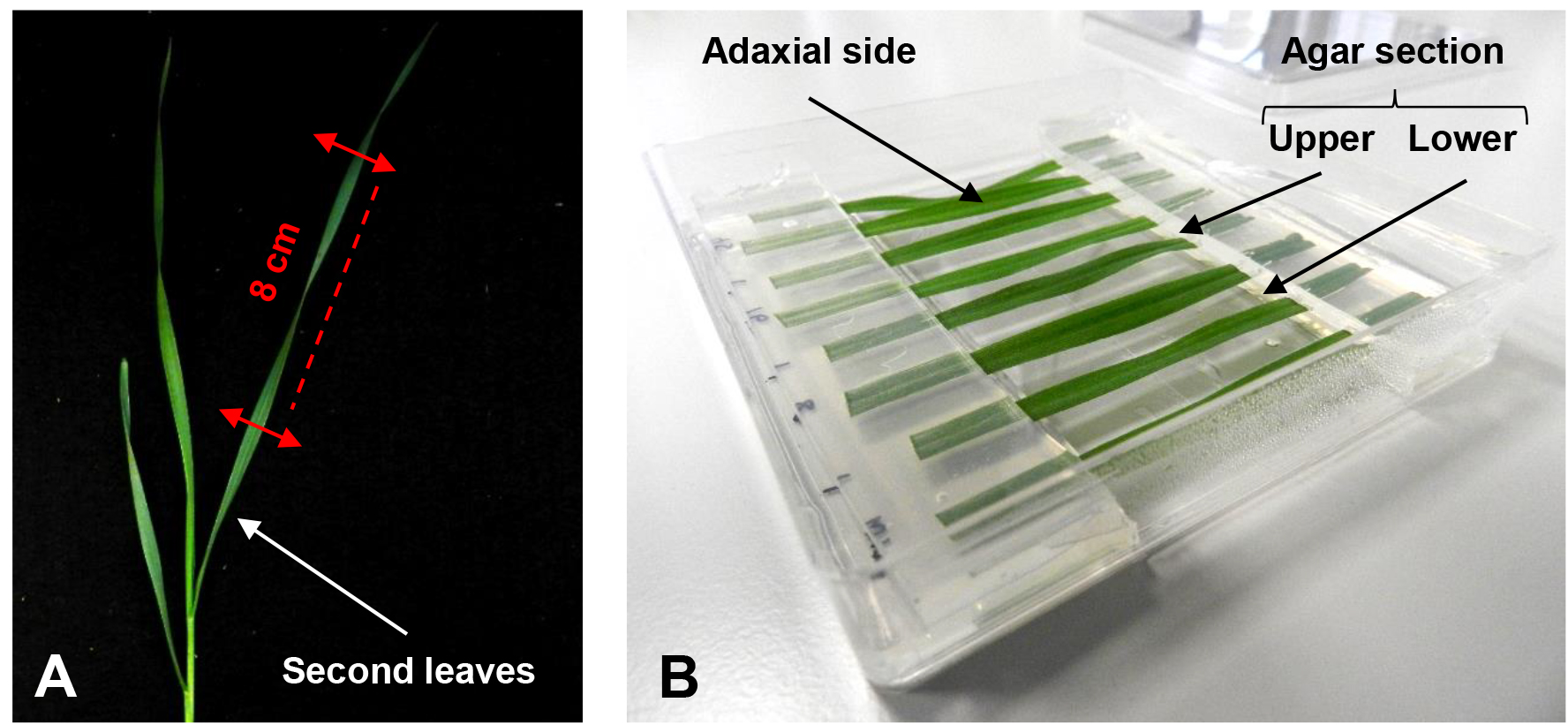
Figure 1. Setup of the detached leaf assay. A. Section of a wheat second leaf at the 3-leaf-stage to be used for the assay. B. Different leaf sections are placed in a square Petri dish with their cut ends held between an upper and lower agar section. - Immediately after cutting, place leaf sections with the adaxial side facing upwards on the surface of a square Petri dish. Put the cut ends between a sandwich of 1 % plant agar pH 5.7 containing 0.5 mM benzimidazole (Figure 1B, Video 1).
Note: Removing the agar from the center of the plate prevent excessive fungal growth at the point of leaf inoculation, additionally removed agar section that could be used to form the sandwich. Benzimidazole is used to delay the leaf senescence.Video 1. Placement of the leaf cut ends between agar sections - Puncture the center of each leaf section with the tip of a glass Pasteur pipette and treat with a 4 μl droplet of 0.02 % (v/v) Tween-20 solution with 106 conidia/ml of F. graminearum strain GZ3639 prepared as previously described (Ali et al., 2015).
- Add 2 ml of water into the plate under the leaf sections to keep high humidity and seal the plates with a piece of Parafilm before to incubate at 20 °C under a 16 h light/8 h dark cycle in a growth room.
Note: Try to place your plates always at the same level and position in your growth room or growth cabinet. The level of condensation inside your plates may differ and affect reproducible disease development. - Analyse the leaf sections at 4 days post-inoculation (Figure 2).
Note: You can follow as well the progress of infection during a time course. In that case photograph your leaf section every day and process analysis as described below (Data analysis). - Repeat the experiment at least three times, each time including 6 plates per treatment, and each plate including two leaf sections per wheat genotype.
Data analysis
- Place the digital camera on the copy stand to calibrate the height and the light to illuminate completely the square petri dish from the side at a 45° angle.
Note: It’s important to make sure that the lighting is even on all the leaf sections particularly on the infected area. Uneven lighting will affect an accurate delineation of the infected area and thus, its measurement. For more details about how to use a copy strand go to: http://www.jiscdigitalmedia.ac.uk/toolkit/digitisation-equipment/the-copy-stand - Remove the cover plates and photograph the leaf sections with a ruler placed inside the field of view. Camera settings were as follows: ISO 160, F-stop 3.4, shutter speed 1/30.
Note: The quality of the image is an important determinant of accurate measurement. Adjust the camera settings according to your light conditions. It’s important to make sure the infected area is in focus to allow your image to be sharp and thus your measurement to be accurate. - Collect all the pictures of your experiment save in TIFF format and measure the disease leaf area using Fiji software (Schindelin et al., 2012).
- Open all your individual pictures in Fiji and create a stack of all your images (‘Image’ > ‘Stacks’ > ‘Image to stacks’).
- Convert your RGB images to 8-bit binary images (‘Image’ > ‘Type’ > ‘8-bit’).
- Set the scale using a known distance of your photographed ruler. Select the button Straight and draw a line along a known distance of your ruler. Set the scale (‘Analyse’ > ‘Set scale’). Add the known distance and select the unit of your choice (e.g., cm).
Note: To make sure your scale is correct, reuse the tool Straight and measure a known distance of your ruler and compare your result (‘Analyse’ > ‘Measure’). - Crop your images to focus your analysis only on the leaf sections. Select the button ‘Rectangular’ and define your area of interest. Make sure that all your leaf sections from your different images are included in your selection before cropping the image (‘Image’ > ‘Crop’).
- To perform an automatic particles analysis, the first step is to set a threshold for the image. This tells the particle analysis what are the objects of interest (infected area) as distinct from the background (non-infected area). Manually calibrate the threshold until the black area fits within your infected area (‘Image’ > ‘Adjust’ > ‘Threshold’) (Figure 2).
- To automatically measure the infected area, use the tool ‘Analyse particles’ (‘Analyse’ > ‘Analyse particles’). Set the minimum and maximum area in the box ‘Size’ (Figure 2). Select inside the box ‘Show’ the setting ‘Overlay outlines’ and tick the boxes ‘Display results’, ‘Summarize’ and ‘In situ show’ (Figure 2).
Note: To know what minimum size to choose, you can manually measure the smallest particle in your pictures with the freehand selection tool. - The results will appear in a table and the corresponding particles measured will be identified on your images. The results from the table can be copy-and-pasted into an Excel file.
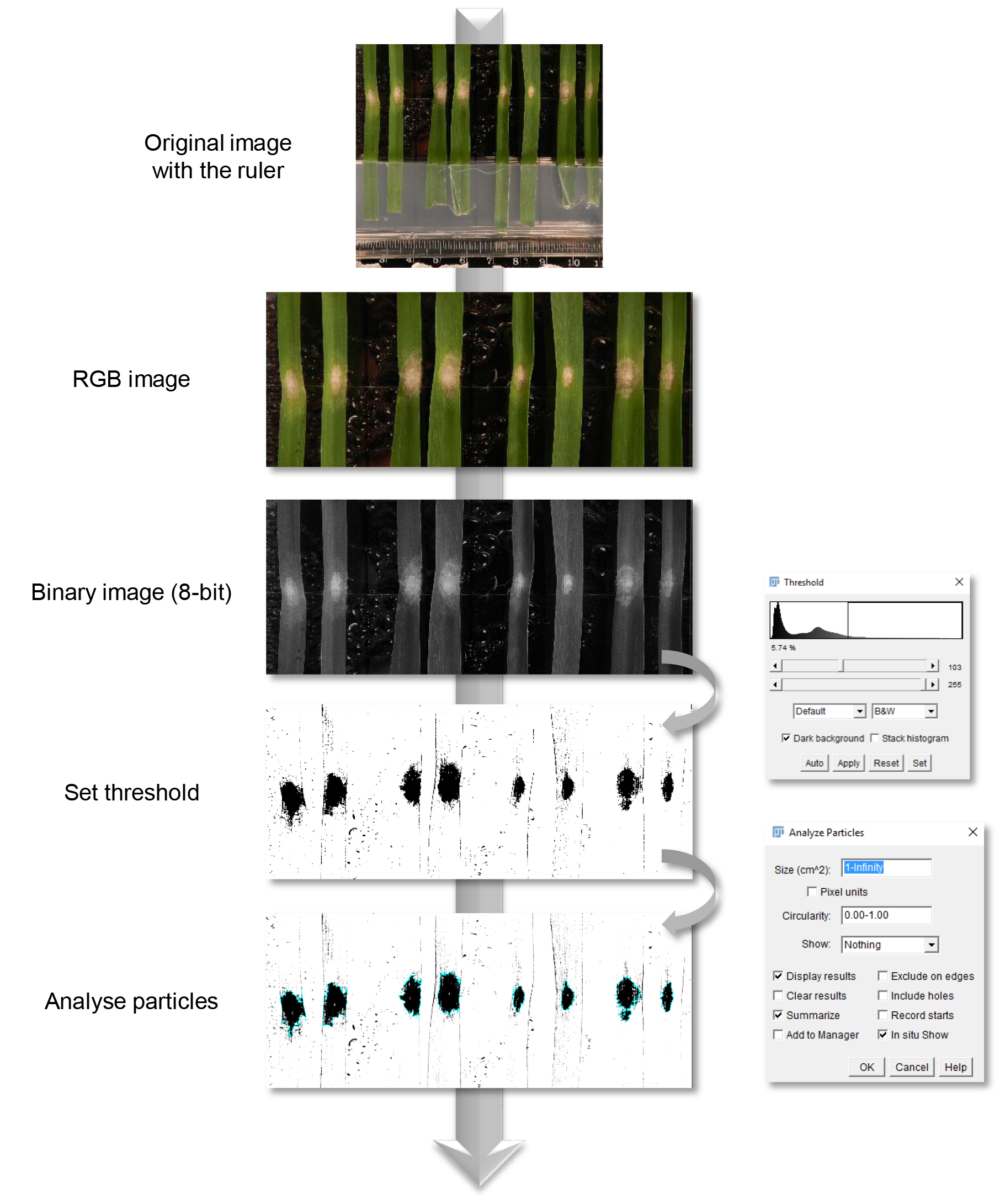

Figure 2. Fiji image analysis pipeline. At four days post-inoculation leaf sections were analysed by image analysis. Corresponding RGB images were converted to binary images. In order to measure automatically the infected area, a threshold was manually set prior to analysis.
Notes
- If you want to compare different genotypes or treatments, avoid the block effect by randomising the leaves within the Petri dishes.
Acknowledgments
This work was supported by the Science Foundation Ireland (project Nos. 10/IN.1/B3028 and 14/1A/2508). Parts of this protocol were adapted from previously described detached leaf disease experiments (Browne and Cooke, 2004; Perochon et al., 2015). The authors thank Pierre Grandjean for technical assistance and Emmanuel G. Reynaud (UCD) for assistance with image analysis.
References
- Ali, S. S., Gunupuru, L. R. and Doohan, F. M. (2015). Visual assessment of the severity of Fusarium seedling blight (FSB) and fusarium head blight (FHB) disease in barley. Bio-protocol 5(12): e1507.
- Browne, R. A. (2009). Investigation into components of partial disease resistance, determined in vitro, and the concept of types of resistance to Fusarium head blight (FHB) in wheat. Eur J Plant Pathol 123(2): 229-234.
- Browne, R. A. and Cooke, B. M. (2004). Development and evaluation of an in vitro detached leaf assay forc pre-screening resistance to Fusarium head blight in wheat. Eur J Plant Pathol 110: 91-102.
- Li, X., Zhang, J. B., Song, B., Li, H. P., Xu, H. Q., Qu, B., Dang, F. J. and Liao, Y. C. (2010). Resistance to Fusarium head blight and seedling blight in wheat is associated with activation of a cytochrome p450 gene. Phytopathology 100(2): 183-191.
- Perochon, A., Jianguang, J., Kahla, A., Arunachalam, C., Scofield, S. R., Bowden, S., Wallington, E. and Doohan, F. M. (2015). TaFROG encodes a Pooideae orphan protein that interacts with SnRK1 and enhances resistance to the mycotoxigenic fungus Fusarium graminearum. Plant Physiol 169(4): 2895-2906.
- Proctor, R. H., Hohn, T. M. and McCormick, S. P. (1995). Reduced virulence of Gibberella zeae caused by disruption of a trichothecene toxin biosynthetic gene. Mol Plant Microbe Interact 8(4): 593-601.
- Schindelin, J., Arganda-Carreras, I., Frise, E., Kaynig, V., Longair, M., Pietzsch, T., Preibisch, S., Rueden, C., Saalfeld, S., Schmid, B., Tinevez, J. Y., White, D. J., Hartenstein, V., Eliceiri, K., Tomancak, P. and Cardona, A. (2012). Fiji: an open-source platform for biological-image analysis. Nat Methods 9(7): 676-682.
- Shin, S., Kim, K. H., Kang, C. S., Cho, K. M., Park, C. S., Okagaki, R. and Park, J. C. (2014). A simple method for the assessment of Fusarium head blight resistance in Korean wheat seedlings inoculated with Fusarium graminearum. Plant Pathol J 30(1): 25-32.
- Zadoks, J. C., Chang, T. T. and Konzak, C. F. (1974). A decimal code for the growth stages of cereals. Weed Research 14: 415-421.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Perochon, A. and Doohan, F. M. (2016). Assessment of Wheat Resistance to Fusarium graminearum by Automated Image Analysis of Detached Leaves Assay. Bio-protocol 6(24): e2065. DOI: 10.21769/BioProtoc.2065.
- Perochon, A., Jianguang, J., Kahla, A., Arunachalam, C., Scofield, S. R., Bowden, S., Wallington, E. and Doohan, F. M. (2015). TaFROG encodes a Pooideae orphan protein that interacts with SnRK1 and enhances resistance to the mycotoxigenic fungus Fusarium graminearum. Plant Physiol 169(4): 2895-2906.
Category
Plant Science > Plant immunity > Disease bioassay
Microbiology > Microbe-host interactions > In vivo model > Plant
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link













