- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
DNA Damage Induction by Laser Microirradiation
Published: Vol 6, Iss 23, Dec 5, 2016 DOI: 10.21769/BioProtoc.2039 Views: 21910
Reviewed by: HongLok LungMarco Di GioiaAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
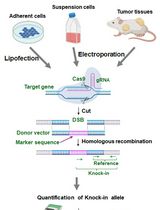
Assay for Site-Specific Homologous Recombination Activity in Adherent Cells, Suspension Cells, and Tumor Tissues
Yuki Yoshino [...] Natsuko Chiba
Apr 5, 2025 2497 Views
Protocol for Quantifying γH2AX Foci in Irradiated Cells Using Immunofluorescence and Fiji Software
Lu Deng [...] Lingying Wu
Aug 20, 2025 2517 Views

A Quantitative DNA Fiber Assay to Monitor Replication Fork Progression, Protection, and Restart
Debanjali Bhattacharya and Ganesh Nagaraju
Feb 5, 2026 330 Views
Abstract
Genome instability can lead to cell death, senescence and cancerous transformation. Specific repair pathways have evolved to prevent accumulation of DNA lesions. Studying these highly dynamic and specific repair pathways requires precise spatial and temporal resolution, which can be achieved through a combination of laser microirradiaiton and live cell microscopy. DNA lesions are introduced at pre-determined sub-nuclear sites and repair can be analyzed in real time in living cells when using fluorescently tagged repair proteins (Mortusewicz et al., 2008). Alternatively, laser microirradiation can be combined with immunofluorescence analysis to study recruitment of endogenous proteins to laser-induced DNA damage tracks that can be visualized by positive controls like, e.g., γH2AX that mark sites of DNA breaks.
Keywords: MicroirradiationBackground
The genomic integrity of mammalian cells is constantly challenged by DNA damage introduced through external and internal sources. Amongst the most common DNA lesions are oxidized bases, double strand breaks, single strand breaks, inter- and intra-strand crosslinks and UV adducts. Various DNA damage signalling and repair pathways have evolved to deal with these lesions. For DNA repair to be fast, precise and efficient, numerous proteins involved in sensing, signalling and repairing specific DNA lesions have to be coordinated in space and time. Furthermore, DNA is organized into higher order chromatin structures and thus for DNA lesions to be accessible to DNA repair enzymes, chromatin has to be remodeled. Laser microirradiation in combination with advanced live cell microscopy allows studying these highly dynamic processes in the context of living cells (Mortusewicz et al., 2008). The protocol described here uses a 405 nm laser that should be readily available at most confocal or spinning disk microscopes to induce DNA damage in living cells and should therefore be cost effective and feasible in most standard cell biology laboratories. Using different sensitization methods (e.g., Hoechst versus BrdU sensitization) and laser energies, the ratio between double strand breaks, single strand breaks, oxidative lesions and UV damage can be modified to the experimental needs.
Materials and Reagents
- Live cell microscopy compatible Petri dish or chambers (e.g., Ibidi, catalog number: 35 mm µ-Grid )
- Adherent cell lines, e.g., U2OS
- If no CO2 control is available at your microscope, use CO2 independent medium without phenol red (e.g., Leibovitz's L-15 medium or CO2 independent medium, Thermo Fisher Scientific, GibcoTM, catalog number: 18045088 ) supplemented with 10% FBS (Thermo Fisher Scientific, GibcoTM, catalog number: 10082139 ) and antibiotics (Thermo Fisher Scientific, GibcoTM, catalog number: 15140-122 ). Alternatively, HEPES (Sigma-Aldrich, catalog number: H4034 ) can be added to your medium of chose.
- 5-bromo-2’-deoxycytidine (e.g., Santa Cruz Biotechnology, catalog number: 1022-79-3 )
- Hoechst 33342 (Thermo Fisher Scientific, Molecular ProbesTM, catalog number: H1399 )
- 4% formaldehyde in PBS (Santa Cruz Biotechnology, catalog number: 30525-89-4 )
- Triton X-100 (Sigma-Aldrich, catalog number: 234729 )
Note: This product has been discontinued. - Mouse-anti-γH2AX (EMD Millipore, catalog number: 05-636 )
- Donkey anti-Mouse IgG (H+L) secondary antibody, Alexa Fluor® 488 (Thermo Fisher Scientific, InvitrogenTM, catalog number: A-21202 )
- Bovine serum albumin (BSA) (Sigma-Aldrich, catalog number: A4503-500g )
- Tween 20 (Sigma-Aldrich, catalog number: P1379-500ML )
- Optional: plasmids encoding for fluorescently tagged proteins of interest, e.g., GFP-Timeless (Figure 1). Fluorescently tagged proteins known to be involved in different DNA repair pathways, like PARP-1, XRCC1 or 53BP1, can be used as controls to set up optimal conditions.
- Optional: transfection reagent of choice, e.g., self-made PEI solution or commercial distributor
Equipment
- Inverted confocal or spinning disk microscope equipped with a 405 nm laser and an environmental chamber or insert to control temperature and CO2/humidity for long-term live cell experiments, e.g., Zeiss LSM710 or LSM780 confocal laser scanning microscope equipped with a UV-transmitting Plan-Apochromat 63x/1.40 Oil DIC M27 or Plan-Apochromat 40x/1.30 Oil DIC M27 objective, respectively
- Cell incubator
Software
- Microscope software, e.g., ZEN from Zeiss (Zeiss)
- Microsoft Excel (Microsoft)
- Image J (https://imagej.nih.gov/ij/)
Procedure
- Seed 4-9 x 104 U2OS cells/ml in a 35 mm µ-Grid dish (Ibidi) 2-3 days before starting the microirradiation experiment. Use dishes with Grid to relocate cells after microirradiation and immunofluorescence staining.
- Optional: transfect cells with plasmid encoding a fluorescently tagged protein of interest 24-48 h before microirradiation experiment. In our hands jetPEI gives good transfection efficiencies in U2OS cells.
- Sensitization of cells for DNA damage induction by laser micorirradiation
- BrdU sensitization: incubate cells in medium containing 10 µM BrdU for at least 24 h prior to microirradiation experiment. BrdU is an analog of thymidine, which gets incorporated into DNA during DNA replication and upon irradiation with the 405 nm laser will lead to induction of DNA damage through, e.g., radical formation.
- Hoechst sensitization: incubate cells in medium containing 10 µg/ml Hoechst for 10 min prior to microirradiation experiment. Hoechst is a cell permeable DNA dye that binds to the minor groove of DNA. As Hoechst is excited by UV light, localized microirradiation with a 405 nm laser will generate DNA damage. Use either Hoechst or BrdU for sensitization.
- Add prewarmed, CO2 adjusted live cell medium supplemented with 10% FBS and antibiotics without phenol-red to cells to minimize autofluorescence.
- DNA damage induction followed by live cell imaging (Figure 1)
- Spot irradiation: For DNA damage induction in live cells expressing a fluorescently tagged protein of interest irradiate a diffraction-limited spot within the nucleus with a 405 nm diode laser (Xie et al., 2015) (405 nm laser set to 70%, 1 iteration, zoom 5, pixel dwell time 177.32 µs).
- Line irradiation: For DNA damage induction in live cells expressing a fluorescently tagged protein of interest irradiate a stripe of 5-pixel width with a 405 nm diode laser (Xie et al., 2015) (405 nm laser set to 24%, 1 iteration, zoom 5, pixel dwell time 12.61 µs).
- Live cell imaging: Before and after microirradiation record confocal image series of one mid z-section at 2 sec time interval (typically 6 pre-irradiation and 60 post-irradiation frames, frame size of 512 x 512 pixels, pixel size of 90 nm), using the laser line according to the excitation spectrum of the chosen reporter protein (e.g., 488 nm Ar laser for GFP/tagged Timeless)
- DNA damage induction followed by immunofluorescence staining
- Laser microirradiation: Use the FRAP module of the ZEN software (line scan, 405 nm laser set to 92%, 50 iterations) or the tile scan mode (Xie et al., 2015; Baldeyron et al., 2011) (3 x 3 tiles, image size 128 x 128, scan speed 177.32 µs, every 7th line scanned, 405 nm laser set to 30%).
- Fixation and immunofluorescence:
- Fix cells in 4% formaldehyde for 10 min after different repair times.
- 1 x wash in PBS
- Permeabilize with 0.5% Triton X-100 in PBS for 5 min
- 1 x wash in PBS
- Incubate in blocking solution for 40 min
- 1 x wash in PBS
- Add primary antibody, e.g., Mouse-anti-γH2AX diluted in blocking buffer and incubate overnight at 4 °C.
- 3 x wash in PBST for 5 min
- Add secondary antibody, e.g., Donkey anti-Mouse IgG (H+L) secondary antibody, Alexa Fluor® 488 diluted in 4% BSA/PBS and incubate for 1 h at RT
- 3 x wash in PBST for 5 min
- Add fresh PBS and store at 4 °C
- Laser microirradiation: Use the FRAP module of the ZEN software (line scan, 405 nm laser set to 92%, 50 iterations) or the tile scan mode (Xie et al., 2015; Baldeyron et al., 2011) (3 x 3 tiles, image size 128 x 128, scan speed 177.32 µs, every 7th line scanned, 405 nm laser set to 30%).
Data analysis
For evaluation of the recruitment kinetics, fluorescence intensities at the irradiated region are corrected for background and for total nuclear loss of fluorescence over the time course and normalized to the pre-irradiation value. For the quantitative evaluation of microirradiation experiments, data of at least 10-20 nuclei from three independent experiments should be averaged and the mean curve and the standard error of the mean calculated and displayed using Microsoft Excel software.
- Image registration and measurement of fluorescence intensities
To compensate for potential cell movements in x and y direction during image acquisition, the TurboReg plug-in from ImageJ can be used. - Open file in ImageJ
- Display ROIs
- Split channels
- Copy the first image of the image series, this will be used as a template for the image registration
- Turbo reg
- Source – original image series
- Target – new image
- Choose Rigid body and set quality to fast
- Batch = movement compensation
- Save new file as tiff
- For analyses define three different regions of interest (ROIs) in ImageJ
- ROIi = irradiated region (spot or line ROI)
- ROIn = outline of the cell nucleus
- ROIb = background region outside of the cell
- Add ROIs to ROI manager (Edit→Selection→add to manager), choose multi measure and save ROIs and measurements (measurement types can be chosen in: Edit→options→input/output. Untick Row, if needed)
- Copy measurements into Microsoft Excel
- Subtract background intensity (ROIb) from fluorescence intensities of nucleus (ROIn) and irradiated region (ROIi): ROIi-b = ROIi - ROIb and ROIn-b = ROIn - ROIb
- To compensate for bleaching during acquisition and set the pre irradiation value to 1 use double normalization: (ROIn-b t0/ROIn-b t)/(ROIi-b t0/ROIi-b t)
Where, ROIi-b t and ROIn-b t is the fluorescence intensity value at time point t and ROIi-b t0 and ROIn-b t0 is the fluorescence intensity at time point zero, respectively (pre-irradiation value)
Figure 1. Determination of protein recruitment to laser-induced DNA damage sites in fixed and live cells. A. Schematic illustration of micorirradiation procedure using either the tile scanning mode to induce DNA damage in a large number of cells followed by fixation and immunofluorescence analysis or line and spot irradiation of single cells expressing fluorescently tagged proteins of interest followed by live cell imaging. B. Representative confocal images after micoirradiation using the tile scanning mode of the Zeiss LSM780. U2OS cells were microirradiated, fixed after indicated time points and stained with antibodies against γH2AX and RPA, which can be used as markers for double strand breaks and ssDNA formations to identify DNA damage sites. C. U2OS cells transfected with expression plasmid encoding GFP/tagged Timeless were microirradiated and recruitment of GFP-Timeless followed in real/time with a time interval of 2 sec.
Notes
To ensure reproducibility between experiments, a laser power meter can be used to measure the 405 nm laser intensity. Alternatively regular immunofluorescence staining with a reference antibody against, e.g., γH2AX as a positive control can be employed.
Recipes
- Fixation solution
4% formaldehyde in PBS - Permeabilization solution
0.5% Triton X-100 in PBS - Blocking solution
4% BSA in PBS - Washing buffer
0.05% Tween 20 in PBS
Acknowledgments
This work was supported by the Helleday foundation. The protocol has been adapted from the following publications: Walter et al. (2003), Mortusewicz et al. (2008), Xie et al. (2015), Mortusewicz et al. (2016), Mortusewicz et al. (2007), and Mortusewicz et al. (2006).
References
- Baldeyron, C., Soria, G., Roche, D., Cook, A. J. and Almouzni, G. (2011). HP1α recruitment to DNA damage by p150CAF-1 promotes homologous recombination repair. J Cell Biol 193(1): 81-95.
- Mortusewicz, O., Amé, J. C., Schreiber, V. and Leonhardt, H. (2007). Feedback-regulated poly (ADP-ribosyl) ation by PARP-1 is required for rapid response to DNA damage in living cells. Nucleic acids research 35(22): 7665-7675.
- Mortusewicz, O., Evers, B. and Helleday, T. (2016). PC4 promotes genome stability and DNA repair through binding of ssDNA at DNA damage sites. Oncogene 35(6): 761-770.
- Mortusewicz, O., Leonhardt, H. and Cardoso, M. C. (2008). Spatiotemporal dynamics of regulatory protein recruitment at DNA damage sites. J Cell Biochem 104(5): 1562-1569.
- Mortusewicz, O., Rothbauer, U., Cardoso, M. C. and Leonhardt, H. (2006). Differential recruitment of DNA Ligase I and III to DNA repair sites. Nucleic Acids Res 34(12): 3523-3532.
- Walter, J., Cremer, T., Miyagawa, K. and Tashiro, S. (2003). A new system for laser-UVA-microirradiation of living cells. J Microsc 209(Pt 2): 71-75.
- Xie, S., Mortusewicz, O., Ma, H. T., Herr, P., Poon, R. Y., Helleday, T. and Qian, C. (2015). Timeless interacts with PARP-1 to promote homologous recombination repair. Mol Cell 60(1): 163-176.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Tampere, M. and Mortusewicz, O. (2016). DNA Damage Induction by Laser Microirradiation. Bio-protocol 6(23): e2039. DOI: 10.21769/BioProtoc.2039.
Category
Cancer Biology > Genome instability & mutation > Cell biology assays > DNA structure and alterations
Cell Biology > Cell imaging > Live-cell imaging
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











