- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Documentation of Floral Secretory Glands in Pleurothallidinae (Orchidaceae) Using Scanning Electron Microscopy (SEM)
Published: Vol 6, Iss 22, Nov 20, 2016 DOI: 10.21769/BioProtoc.2021 Views: 10468
Reviewed by: Samik BhattacharyaNing LiuAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
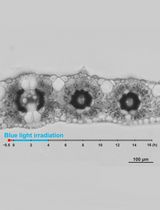
Live Leaf-Section Imaging for Visualizing Intracellular Chloroplast Movement and Analyzing Cell–Cell Interactions
Yuta Kato [...] Mitsutaka Taniguchi
Aug 5, 2025 2377 Views

Live-Cell Monitoring of Piecemeal Chloroplast Autophagy
Masanori Izumi [...] Shinya Hagihara
Nov 5, 2025 1723 Views
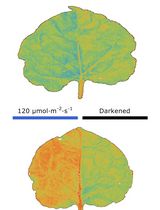
Chloroplast Movement Imaging Under Different Light Regimes With a Hyperspectral Camera
Paweł Hermanowicz [...] Justyna Łabuz
Dec 20, 2025 792 Views
Abstract
A clear, step by step description of the treatment of orchid flowers, subtribe Pleurothallidinae, with Critical Point Drying for SEM is presented. It shows that a simple, short fixation and dehydration method prior to Critical Point Drying is sufficient to obtain good results.
Background
Pleurothallidinae (Orchidaceae) have relatively small flowers that exhibit a wide variety of ornaments on the perianth parts, especially on the sepals and lip. It is clear that most of those play an important role in attracting and arresting pollinators, which in most cases are known to be small flies. However, it is difficult to judge just from looking at photographs from the stereoscopic microscope if the observable thickenings, papillae or hairs are actually secretory glands. Scanning Electron Microscopy (SEM) can be a powerful tool to explore these flowers in greater detail, aiding to locate, compare and identify those glands (Karremans et al., 2015).
Materials and Reagents
- Snap cap vials (10 ml) (VWR, catalog number: 548-0621 )
- Gloves (nitrile, powder free)
- Petri dish
- Double sided ultra-smooth carbon adhesive tape, 12 mm diameter (Electron Microscopy Sciences, catalog number: 77827-12 )
- Plastic box or desiccator with dry silica gel
- Ethanol
Note: For steps using diluted ethanol use a 96% non denatured ethanol with distilled water, for steps using pure ethanol use a min. of 99.9%, water free ethanol. - Acetone, HPLC Plus, for HPLC, ≥ 99.9% (Sigma-Aldrich, catalog number: 650501-1L )
- Carbon dioxide, technical grade (in gas cylinder with feed pipe)
- Argon gas 4.6, technical quality
Equipment
- Aluminium specimen mount, JEOL(Ted Pella, catalog number: 16231 )
- Soft pincet (e.g., light forceps, [VWR, catalog number: 232-2119 ])
- SEM specimen mount tweezers (Electron Microscopy Sciences, catalog number: 76800 )
- Platina/palladium (Pt/Pd) target, 57 x 0.1 mm (Pt/Pd ratio, 80:20)
- Automated critical point dryer (Leica Microsystems, model: EM CPD300 )
- Rotary mixer (Ted Pella, Pelco®, model: R1 )
- Holders and baskets for Leica critical point dryer (e.g., Fine mesh specimen holder with mesh specimen baskets or filter disc holder with 4 wells)
- Sputter coater (Quorum Technologies, model: Q150T S ) with rotating holder for 6 SEM stubs (Q150T S/E/ES Sample Preparation System Instruction Manual), film thickness monitor and RV3 Rotary Vane pump (Edwards, model: A65201903 )
- Field emission scanning electron microscope (JEOL, model: JSM-7600F ) (Figure 1)
Note: This product is discontinued, but any standard scanning electron microscope will do (for example JEOL, model: JSM-IT100 InTouchScopeTM)
Figure 1. Field emission scanning electron microscope in place and ready for use
Software
- Jeol PC-SEM version 2.1.0.3
Procedure
- Preparing the samples for critical point drying
- Carefully harvest fresh flowers using a conventional pincet. Take care of collecting only young, undamaged flowers. It is important to collect material from plants that are in good, healthy condition (for example the same day of anthesis; Figure 2). Place the flower in a plastic tube containing ethanol (70%), covering it completely. Leave it to soak for at least one week (this is highly dependent on the size of the material, and it is likely that a shorter period of a few days will also be enough for Pleurothallidinae flowers). The condition of the cell structures observed in SEM is mostly dependent on the quality of the collected material.
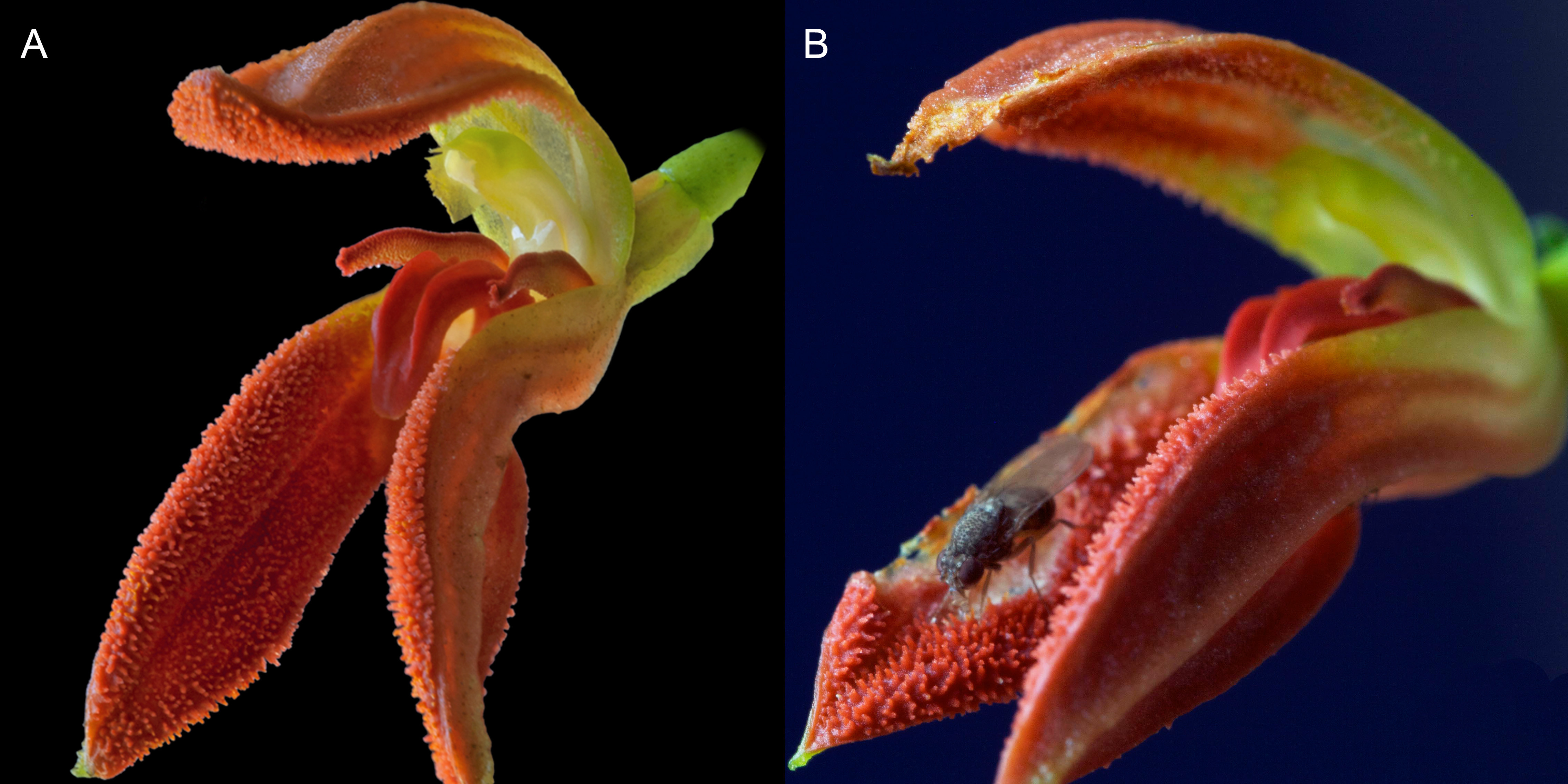

Figure 2. Flowers of Specklinia remotiflora. A. Shortly after anthesis, flower in healthy condition. B. Damaged flower, a few days after anthesis, not recommended for visualization with SEM. - Place flowers directly from ethanol 70% into snap cap vials (10 ml) with acetone pure grade. Cover the flower completely. Try to leave as little time as possible between cutting the flowers from the plant and placing them in ethanol 70%.
- Turn slowly in a rotary mixer for 30 min at room temperature.
- Replace the used for fresh acetone, once again covering the flowers immediately. Flowers should be exposed to air as short as possible because this damages the cells.
- Turn slowly for 30 min once again.
- Submerge the holders/baskets for critical point drying in a Petri dish with acetone. Put flowers in the holders. Remember not to let them dry up! Add a small (5 mm) piece of paper with (pencil written) sample number to each sample well.
- Proceed to critical point drying following the Leica EM CPD300 protocol (Leica EM CPD300 application booklet) (Figure 3).
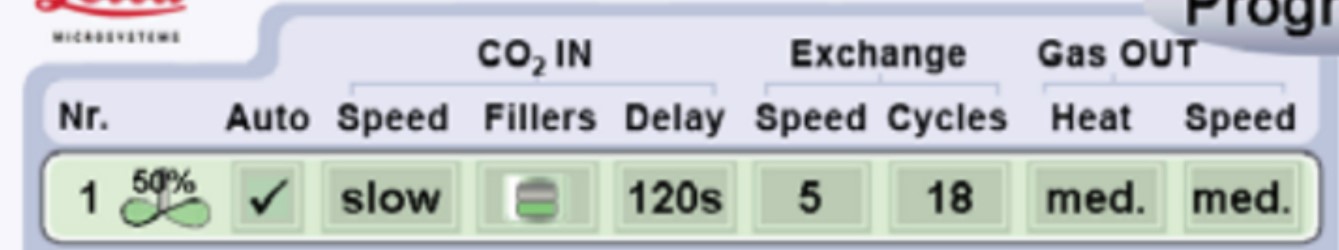
Figure 3. Settings used for critical point drying using the Leica EM CPD300
- Carefully harvest fresh flowers using a conventional pincet. Take care of collecting only young, undamaged flowers. It is important to collect material from plants that are in good, healthy condition (for example the same day of anthesis; Figure 2). Place the flower in a plastic tube containing ethanol (70%), covering it completely. Leave it to soak for at least one week (this is highly dependent on the size of the material, and it is likely that a shorter period of a few days will also be enough for Pleurothallidinae flowers). The condition of the cell structures observed in SEM is mostly dependent on the quality of the collected material.
- Placing and displaying the samples for coating
- Take samples out of the dryer. They should be papery, light, brittle and practically colorless. Be careful handling them, they can easily get damaged.
- Place the aluminum specimen mounts in acetone for 60 sec to clean them. Take them out and place on a tissue, allowing the acetone to evaporate completely by exposing to air for a few minutes. Handle stubs with gloves from now on. Paste carbon adhesive double sided tape on one (top) end. Label stubs on the bottom according to samples
Note: You might need more than one stub per sample depending on size of the flowers. - Handle flowers carefully using a soft pincet. Consider that every part touched with the pincet will be damaged, therefore hold them from a point which is not interesting to visualize later. In most pleurothallids the rachis, pedicel or ovary will do as they may not be of interest for the visualization.
Note: You can have multiple flowers dried so that you can have the different organs represented once. - Place flower on tape. You will need to open up the flowers as much as possible to allow visualisation. Flat flowers like Platystele and Stelis can just be placed entirely (Figure 4), flat on the tape, more tubular flowers like Acianthera and Specklinia have to be dismembered (Figure 5). In those cases, after the synsepal is fixed you can remove the dorsal sepal and petals. It is important to touch the segments as little as possible, to place them on the tape in final position (you will not be able to change position after it was placed), and to consider all possible viewing points you would like.
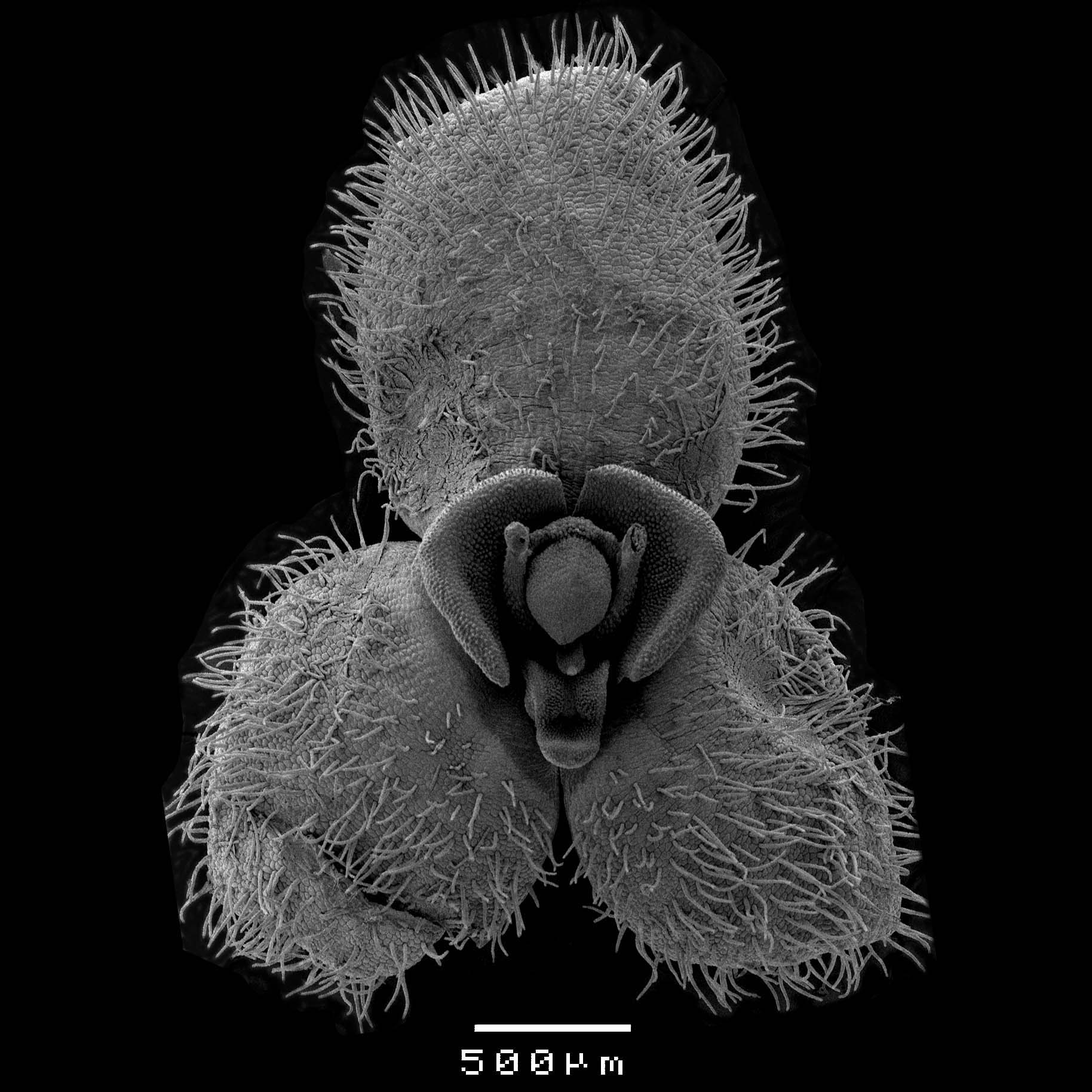
Figure 4. Small and flat flowers like that of Stelis can be displayed completely. Scale bar = 500 μm.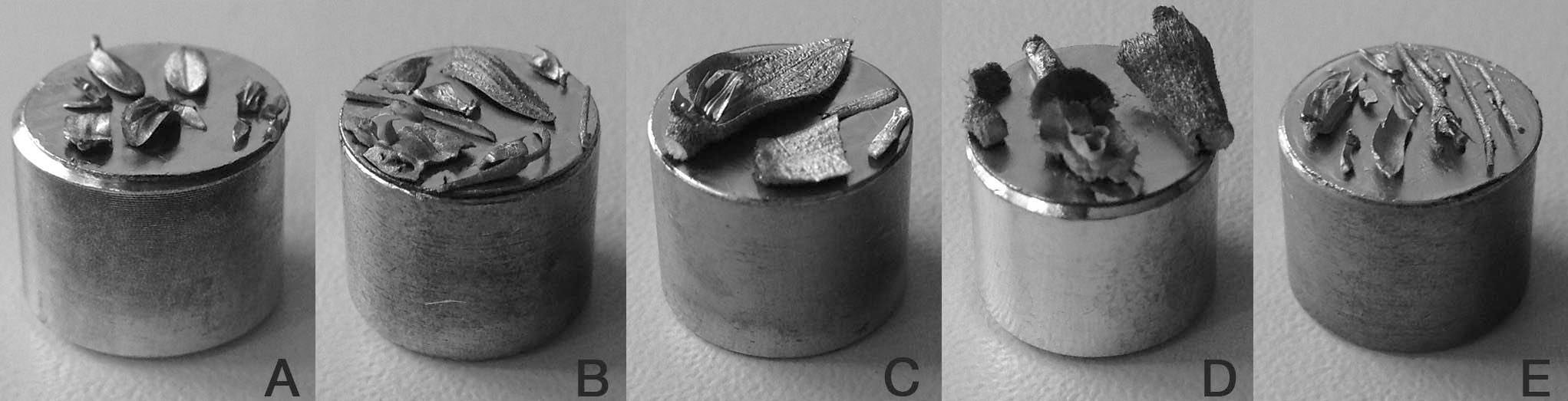
Figure 5. Stubs showing diverse Pleurothallidinae flowers, dismembered and placed according to the visualization needs. A. Lankesteriana sp. B, E. Specklinia spp. C, D. Echinosepala spp. - Store stubs in a Petri dish, and in a closed plastic box or desiccator with some dry silica gel until they will be sputter-coated. Samples can be stored for years at room temperature in a relatively dry place. In places where the humidity is high samples can be stored together with silica gel. If necessary sputter-coating can be repeated before the re-visualization.
- Take samples out of the dryer. They should be papery, light, brittle and practically colorless. Be careful handling them, they can easily get damaged.
- Sputter-coating the samples
- Open Argon cylinder allowing gas to reach the Q150T S sputter coater (Q150T S/E/ES Sample Preparation System Instruction Manual). Turn the machine on.
- Carefully remove the glass chamber and place the stubs in the holder distributing them evenly so that it is balanced when rotating. Place the glass chamber back on. Lower the cover taking care that it does not touch the samples. Select a ‘Profile’ on the operating screen. It is recommended that you use a coating thickness of PT/Pd 20 nm because Pleurothallidinae flowers are quite irregular. (A PT/Pd 10 nm would probably be enough otherwise.) Run the profile (Figure 6). Anthers and pollinaria often become charged with electrons during observation in the SEM, which disrupts the image acquisition. A thicker coating may help prevent this problem.
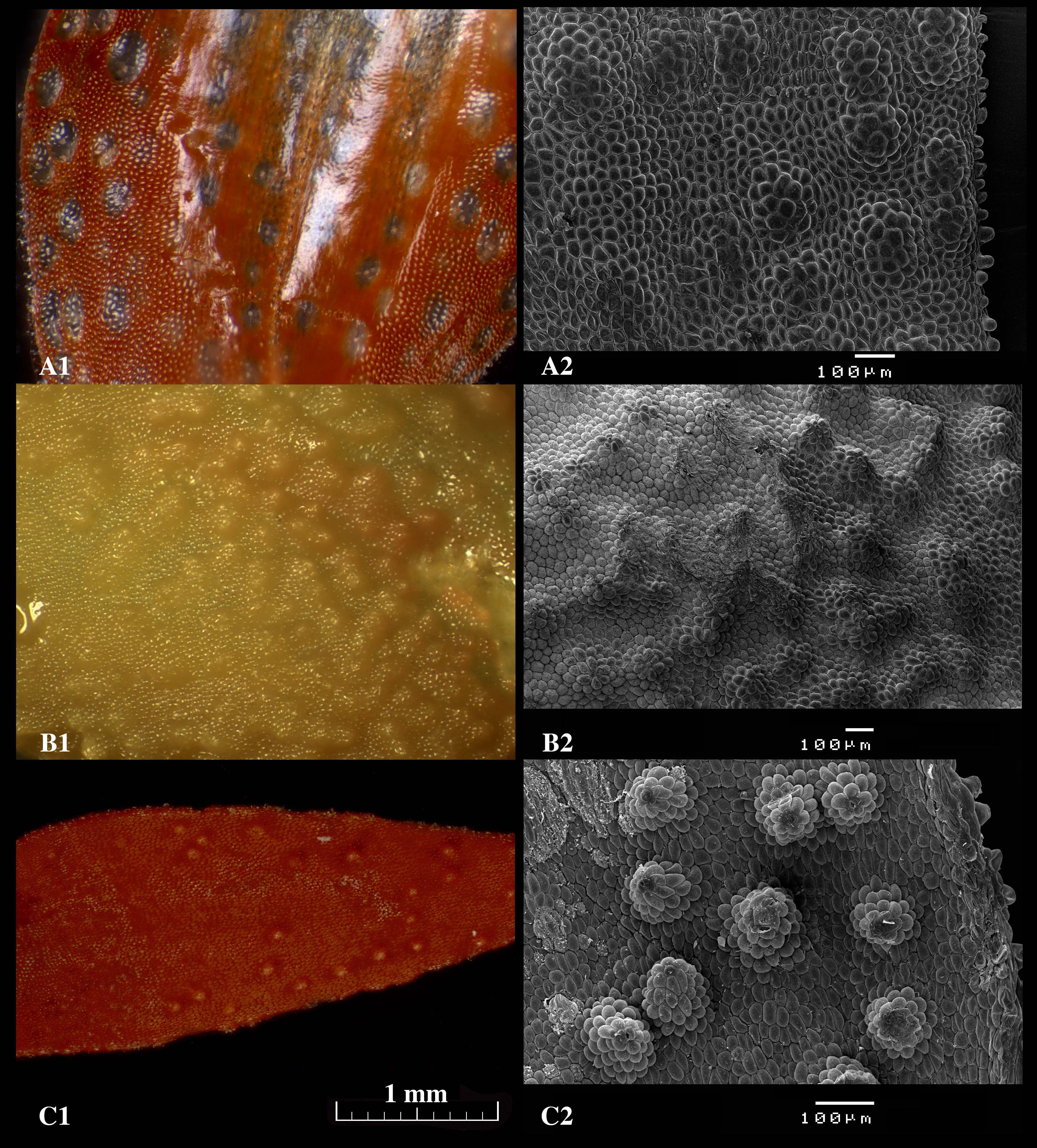
Figure 6. Micrographs of the glandular areas of the sepals of Specklinia species taken without coating using a stereoscopic microscope (1) and with coating using the scanning electron microscope (2). A. S. barbae; B. S. pfavii; C. S. tribuloides. Scale bars = 1 mm (A1, B1 and C1); Scale bars = 100 μm (A2, B2 and C2). - After the run has completely finished the vacuum should allow for the opening of the cover. The samples can be removed using a pincet and returned to the Petri dish.
- Open Argon cylinder allowing gas to reach the Q150T S sputter coater (Q150T S/E/ES Sample Preparation System Instruction Manual). Turn the machine on.
- Visualization and photographs
- Turn the JEOL JSM-7600F field emission scanning electron microscope on, following the microscope manual.
- Use the multiple 12 mm stub holder and place up to six stubs tightly. Always use powder free gloves when handling the holder and the stubs.
- Follow the microscope manual to place the samples inside using the specimen exchange rod.
- Follow the microscope manual to visualize the material. Set the parameters for visualization: Set the work distance (Z) to 15 mm, and set the acceleration voltage to 10 KV. Probe current is 6 and detector mode is SB.
Note: The work distance may be varied, a shorter distance gives higher resolution but lower focus depth, a higher work distance will give lower resolution but higher focus depth. Accelerating voltage can be varied, at higher kV the resolution becomes better but there is more chance of charging. It is advised not to use more than 15 kV for biological samples. Lower kV can be useful if you need to avoid charging (e.g., 5 kV) or if you want to work without coating your material (e.g., 2 kV). However, a lower kV gives less resolution and noisier images. For our work with this specific SEM we always use Probe current 6 and a detector mode SB for our biological samples. This setting combines collection of side scattered and backscattered electrons, which gives good image results. - When working with magnifications higher than 500x you can switch from ‘Low Mag’ to ‘High Mag’ mode.
- Save pictures as bitmap (Figure 7).
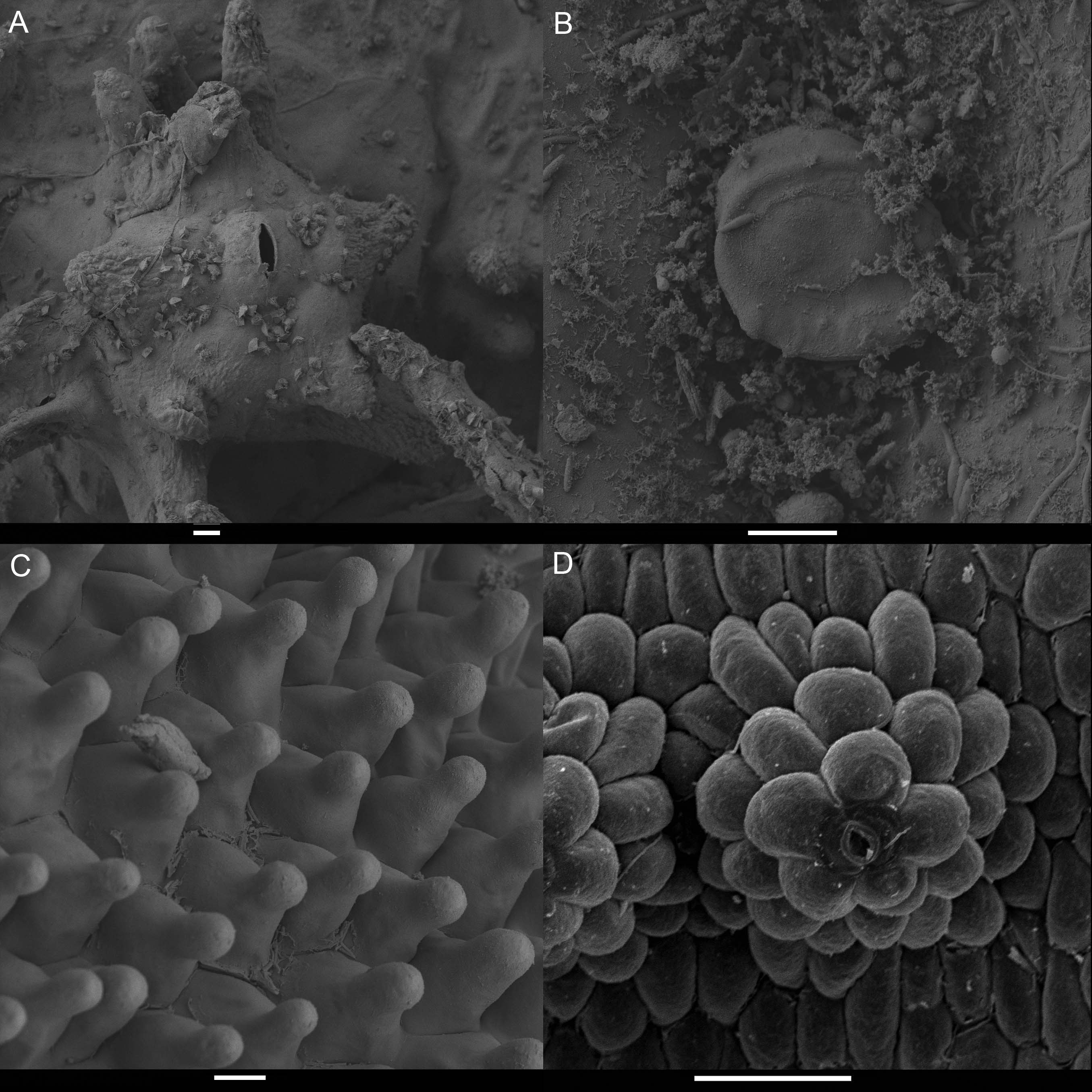

Figure 7. Scanning electron micrographs of diverse glands found in the flowers of Pleurothallidinae. A. Possible osmophore on the abaxial surface of the sepals of Echinosepala. B. Sunken trichome on abaxial surface of Condylago (= Stelis). C. Nectaries on the lip of Platystele. D. Nectar secreting stomata on adaxial surface of the sepals of Specklinia. The measurements represent 100, 100, 100 and 50 μm.
- Turn the JEOL JSM-7600F field emission scanning electron microscope on, following the microscope manual.
Notes
- If no fresh flowers are available it is possible to use specimens kept in the liquid collections at herbaria (Procedure A, step 1). Flowers preserved in FAA and its variations will still be suitable for visualization through the method described here.
- When Critical Point Drying (Procedure A, steps 2-6), instead of acetone, you can also use 100% ethanol (absolute). In our experience, this also gives good results.
Acknowledgments
The Alberta Mennega Foundation funded the first author at several stages of the study that resulted in the protocol here described. This protocol has been modified from the original application booklet by Leica Microsystems.
References
- Karremans, A. P., Pupulin, F., Grimaldi, D., Beentjes, K. K., Butot, R., Fazzi, G. E., Kaspers, K., Kruizinga, J., Roessingh, P., Smets, E. F. and Gravendeel, B. (2015). Pollination of Specklinia by nectar-feeding Drosophila: the first reported case of a deceptive syndrome employing aggregation pheromones in Orchidaceae. Ann Bot 116(3): 437-455.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Karremans, A. P., van Heuven, B. J., Langelaan, R. and Gravendeel, B. (2016). Documentation of Floral Secretory Glands in Pleurothallidinae (Orchidaceae) Using Scanning Electron Microscopy (SEM). Bio-protocol 6(22): e2021. DOI: 10.21769/BioProtoc.2021.
Category
Plant Science > Plant cell biology > Cell imaging
Plant Science > Plant cell biology > Tissue analysis
Cell Biology > Cell imaging > Electron microscopy
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










