- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Protocol for Primary Microglial Culture Preparation
Published: Vol 6, Iss 21, Nov 5, 2016 DOI: 10.21769/BioProtoc.1989 Views: 26591
Reviewed by: Soyun KimPengpeng LiAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Selective Enrichment and Identification of Cerebrospinal Fluid-Contacting Neurons In Vitro via PKD2L1 Promoter-Driven Lentiviral System
Wei Tan [...] Qing Li
Nov 20, 2025 1373 Views
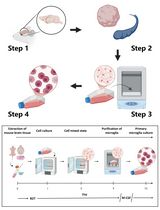
Revisiting Primary Microglia Isolation Protocol: An Improved Method for Microglia Extraction
Jianwei Li [...] Guohui Lu
Dec 5, 2025 1561 Views
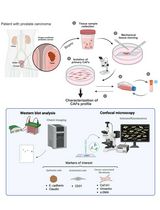
Non-Enzymatic Isolation of Cancer-Associated Fibroblasts From Human Prostate Tumor Explants
Giulia Gangarossa [...] Paola Chiarugi
Mar 5, 2026 116 Views
Abstract
Primary microglia, in either mono-culture or co-culture with neurons or astrocytes, are a powerful tool for studying mechanisms underlying microglial inflammatory responses and cell type-specific interactions in the central nervous system (CNS). This protocol provides the details of how to prepare high purity primary microglia from newborn mouse pups. The overall steps include brain cell dissociation, mixed glial cell culture, and microglia isolation.
Background
In recent years, neuroinflammation has become a hotspot area in neuroscience studies. Inflammatory responses, such as glial activation and cytokine upregulation, were observed in brains of patients with various neurological diseases (Fan et al., 2015; Koshimori et al., 2015; Garden and Campbell, 2016). Neuroinflammation is considered not only a consequence of pathological changes in the brain but also a contributor to disease progression (Schwartz et al., 2013). In addition, the physiological functions of inflammatory pathways, the importance of which were previously underestimated, are being revealed as surprisingly versatile. For instance, activation of the complement signaling pathway is commonly observed in the central nervous system (CNS) in neurological diseases and is suspected to be involved in disease pathophysiology (Michailidou et al., 2015; Loeffler et al., 2008). Now we know that it also plays essential function in the developmental regulation of synaptic refinement (Stevens et al., 2007). Along with the increasing attention on inflammation, interest in microglial function during development, neuroprotection, and pathogenesis continues growing. Microglia are resident innate immune cells of myeloid lineage located in the brain and are critical components of the immune system in the CNS. The activation of microglia in some neurological diseases may directly participate in pathogenic processes. For instance, TREM2 mutations, which affects only microglia, are a genetic risk factor for Alzheimer’s disease (Yuan et al., 2016; Wang et al., 2015). At the same time, developmental roles of microglia are being revealed. For example, synaptic maturation during early development requires microglia and this regulation may underline the pathogenesis of developmental diseases such as autism (Edmonson et al., 2016; Stephan et al., 2012). Tools for studying microglia include in vivo models (e.g., microglia-deficient PU.1 knockout mice [McKercher et al., 1996]) and in vitro systems such as immortalized microglial cell lines and primary microglial culture. While in vivo tools are powerful for demonstrating systematic microglial function, in vitro tools are ideal for mechanistic characterization due to the easy manipulation of experimental factors. Compared to immortalized microglial cell lines, primary microglia better mimic in vivo microglial properties (Stansley et al., 2012). The altered gene expression upon stimulation may be better presented in primary microglia than in microglial cell lines (Stansley et al., 2012; Henn et al., 2009). Here we described a protocol for establishing high purity primary microglial culture derived from neonatal mice and the method has yielded robust results in our work (Lian et al., 2016). Dissociated cells are collected through enzymatic digestion of collected brains and seeded to grow mixed glial culture. Microglia growing on top of a confluent astrocyte layer are purified through mechanical tapping of mixed glial culture.
Materials and Reagents
- 15 ml centrifuge tubes (Corning, catalog number: 430052 )
- 50 ml centrifuge tubes (Corning, catalog number: 430290 )
- 12-well plates (Corning, Costar®, catalog number: 3737 )
- New born pups (mouse, P0-P2)
- Poly-D-lysine (PDL) (Sigma-Aldrich, catalog number: P6407-5MG )
- Ethanol
- Dulbecco’s modified Eagle medium (DMEM) (Thermo Fisher Scientific, GibcoTM, catalog number: 11995065 )
- Fetal bovine serum (FBS) (GE Healthcare, HycloneTM, catalog number: SH30088.03 )
- 10,000 U/ml penicillin-streptomycin (Pen/Strep) (Thermo Fisher Scientific, GibcoTM, catalog number: 15140122 )
- Hanks’ balanced salt solution (HBSS) (Thermo Fisher Scientific, GibcoTM, catalog number: 24020117 )
- 1 M HEPES buffer solution (Thermo Fisher Scientific, GibcoTM, catalog number: 15630080 )
- Glucose (Thermo Fisher Scientific, Fisher Scientific, catalog number: D16-3 )
- Trypsin, powder (Thermo Fisher Scientific, GibcoTM, catalog number: 27250018 )
- Trypsin inhibitor (Sigma-Aldrich, catalog number: T6522-100MG )
- Deoxyribonuclease I (DNase I) (Sigma-Aldrich, catalog number: DN25-100MG )
- Culture medium (500 ml) (see Recipes)
- Dissection medium (500 ml) (see Recipes)
- 2.5% trypsin (20 ml) (see Recipes)
- 1 mg/ml trypsin inhibitor (20 ml) (see Recipes)
- 10 mg/ml DNase (20 ml) (see Recipes)
Equipment
- Vented cap T-75 culture flask (Corning, catalog number: 3276 )
- Dissection tools
- Fine scissors (Fine Science Tools, catalog number: 14060-09 )
- Spring scissors (Fine Science Tools, catalog number: 15009-08 )
- Curved standard forceps (Fine Science Tools, catalog number: 11052-10 )
- Fine forceps (Fine Science Tools, catalog number: 11370-40 )
- Centrifuge machine (Eppendorf, model: 5702 )
- Hemocytometer
- Ventilation hood (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 1323 )
- CO2 cell culture incubator (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 50144906 )
- 37 °C water bath (Thermo Fisher Scientific, Thermo ScientificTM, model: TSGP02 )
Procedure
Depending on the experimental design, the number of microglial cells required for experiments varies. Below we list the steps of processing 3 newborn mouse pups to generate mixed glial cultures in two T-75 flasks. In the mixed culture, astrocytes form a confluent cell layer at the bottom and microglia grow on top of the astrocytic layer. The total amount of primary microglia generated from two T-75 flasks should be enough to seed four 12-well plates at a density of 50,000 cells/cm2.
- Coat two T-75 culture flasks with 7 ml each of 10 μg/ml PDL for 2 h. Wash the flask bottom with distilled water 3 times before use.
Note: You can coat more flasks than needed and the unused coated flasks can be stored at 4 °C for months. We usually keep the flasks in plastic wrap to avoid contamination. - Collect new born pups from breeding cages. Keep the pups on a 37 °C heating plate to maintain body temperature. In the meantime, prepare tools and reagents needed for the culture experiment. Spray dissection tools and work space with 75% ethanol. Warm up culture medium (No. 16 in ‘Materials and Reagents’ and No. 1 in ‘Recipes’) in 37 °C water bath.
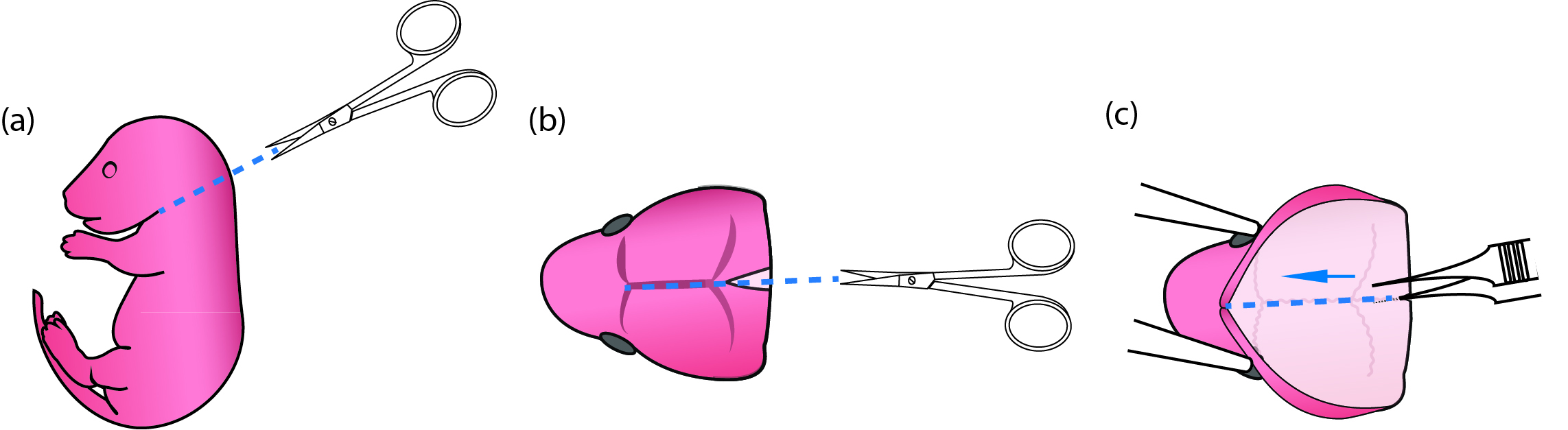
Figure 1. Process of newborn mouse pups for primary microglial culture. (a) Use fine scissors to cut off the pup head. (b) Use fine scissors to cut open the scalp along the midline from the posterior end to the middle point between the two eyes. (c) When the thin skull was exposed, put one end of the fine forceps beneath the skull but above the brain tissue and pull toward the snout along the midline so that the brain could then be easily scooped out using curving forceps. - Remove pups from the heating plate. Decapitate and place the heads into a 6 cm Petri dish containing 5 ml cold dissection media (Figure 1a) (No. 17 in ‘Materials and Reagents’ and No. 2 in ‘Recipes’). Use fine scissors to cut open the scalp along the midline starting posteriorly and ending near the snout (Figure 1b). Place one sharp tip of the fine forceps beneath the skull at the posterior end of a brain, and cut the skull by pushing the end from posterior to anterior (Figure 1c). Scoop out the brain using curving forceps and immerse the brains in 5 ml cold dissection media in a new Petri dish.
Note: Newborn pups have a transparent and soft skull. Using scissors to cut the skull may damage the fragile brain tissue underneath. - Put the Petri dish containing the brains under a dissection microscope. Carefully remove the meninges (readers may refer to the video presented by Bowyer et al., 2012 for this step) and collect the cortices and hippocampi. If you use 3 pups, you will get 6 halves. Put 3 halves per Petri dish with 5 ml dissection media and mince the tissue into small pieces using spring scissors.
- Transfer the contents of each dish to a 50 ml tube. Wash the dish with dissection media to collect any remaining tissue on the dish and repeat the transfer. Fill the 50 ml tube to reach a final volume of 30 ml dissection media.
- Add 1.5 ml 2.5% trypsin to each tube and incubate in the 37 °C water bath for 15 min. Swirl frequently.
- Add 1.2 ml 1 mg/ml trypsin inhibitor and incubate for 1 min. Add 750 μl 10 mg/ml DNase to digest the sticky DNA released from dead cells.
- Centrifuge the tube at 400 x g for 5 min. Aspirate the supernatant and triturate the pellet with 5 ml warm culture media using a 1 ml pipet tip. Transfer the homogenous cell suspension to a 15 ml tube. If undissociated tissue chunks remain, let them settle and repeat the trituration and transfer step using 3 ml media.
- Centrifuge the 15 ml tubes at 400 x g for 5 min. Aspirate the supernatant and resuspend the pellet with 5 ml warmed culture media.
- Count the cell density using hemocytometer.
- Plate each tube of cells into one coated T-75 flask at the density of 50,000 cells/cm2. Add culture media to reach a volume of 15 ml in the flasks. Put seeded flasks into a CO2 cell culture incubator with 5% CO2, 100% humidity at 37 °C.
- Change the culture medium the next day to remove cell debris and then change culture media every 5 days.
- In 5-7 days, astrocytes at the bottom of the flask form a confluent cell layer (Figure 2). Microglia and some oligodendrocytes grow on top of the astrocytic layer.

Figure 2. Astrocytes form a connective confluent layer at the bottom in the mixed glial culture. Arrows point to representative astrocytes. Scale bar = 100 μm. - To collect microglia, vigorously tap the flasks on the bench top and collect the floating cells in conditioned culture media (No need to change media before tapping). The resulting cells are purified microglia. Use a hemocytometer to count the floating cell density and seed the cells at 50,000 cells/cm2 in PDL-coated culture vessels. After 2 h, check that the microglia have attached to the bottom under a microscope. Aspirate medium and replace with fresh culture medium. The microglial cells are ready to use the next day (Figure 3).
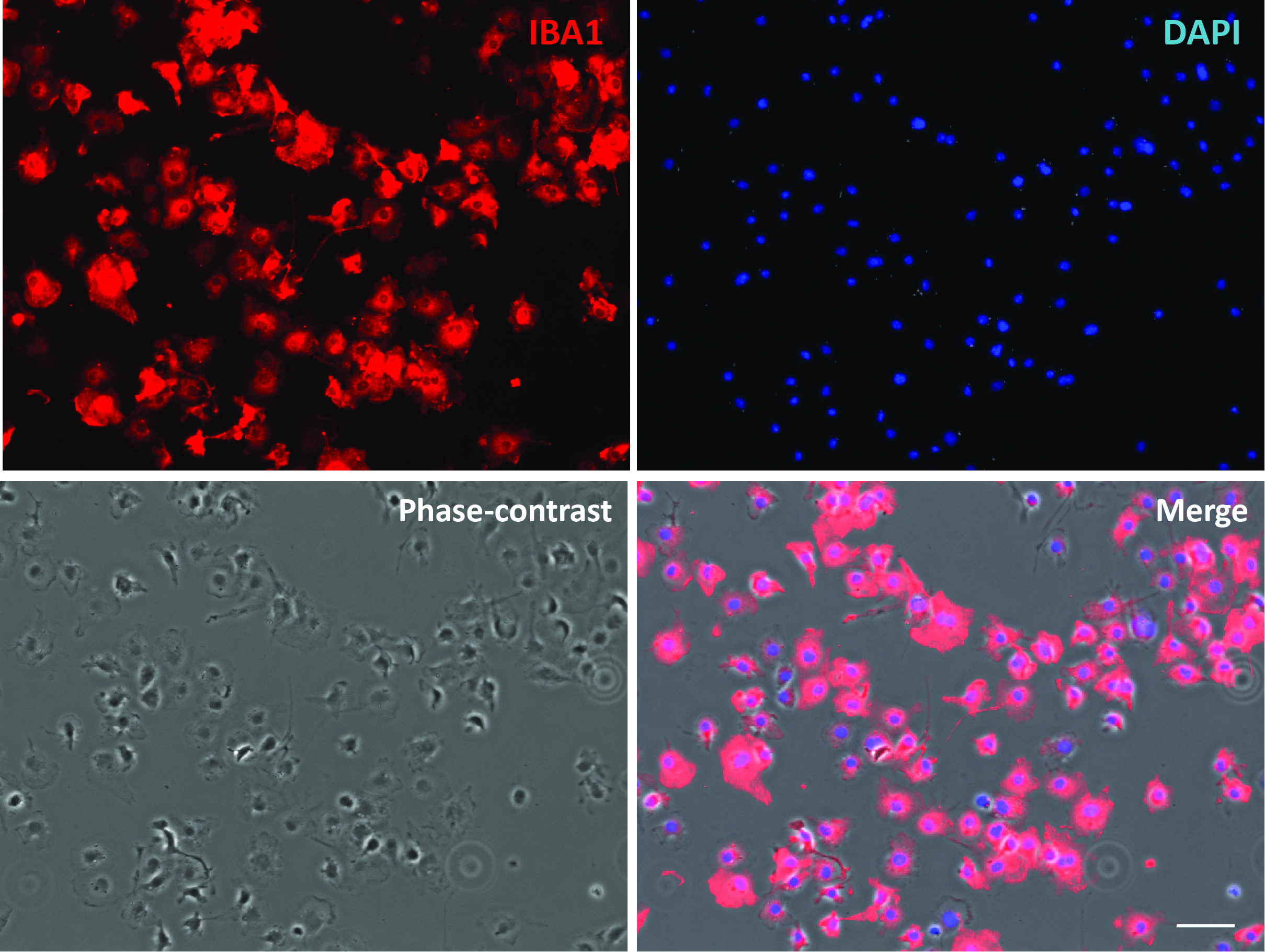
Figure 3. Purified primary microglial culture. Cells were stained with microglial marker protein Iba1. Phase-contrast image was taken to show the cell body. Scale bar = 50 μm.
Data analysis
Getting good quality primary microglial culture is the basis for experiments such as phagocytosis, RNA and protein analysis upon various treatments, and immunocytochemical staining. However, the statistical analysis of experiments after primary microglial culture is beyond the scope of this protocol and therefore, the data analysis process is not discussed here.
Notes
- This protocol described microglial preparation from newborn pups (P0-P2). Older pups could also be used since glial cell are not post-mitotic like neurons, but new born pups give better yield.
- Most of the procedure should be done in a sterile ventilation hood. Exceptions include the handling of the pups, the brain dissection, centrifugation, and cell counting. Use sterile tubes and dishes. Spray tools and outside surface of tubes and dishes with 70% ethanol before carrying them into the hood. Except for the 37 °C incubations in the water bath and culture incubator, all steps are performed at room temperature and should be completed in a timely manner to enhance viability of cells.
- If the mixed glial cells reach confluency but microglial cells are not needed immediately, the mixed glial culture can be passaged. Mixed glial cultures can also be frozen and stored for long term in freezing media composed of DMEM with 20% FBS and 10% DMSO in liquid nitrogen. When the new passage or recovered frozen cells reach confluency, microglia can grow on top of the confluent cell layer and be purified by tapping.
- Astrocytes and microglia grow more vigorously than oligodendrocytes in this culture condition. After tapping the mixed glial culture, the collected floating cells may contain some oligodendrocytes. However, microglial cells have much stronger attaching capability than oligodendrocytes. After seeding the floating cells, microglia attach to the culture vessel bottom much more efficiently than oligodendrocytes. At 2 h after seeding before fresh media is added, aspiration of the old media will remove unattached contaminating oligodendrocytes.
Recipes
- Culture medium (500 ml)
450 ml DMEM
50 ml FBS
Optionally, you can add 5 ml Pen/Strep
Filter and store at 4 °C - Dissection medium (500 ml)
450 ml 1x HBSS
5 ml 1 M HEPES
3 g glucose powder
5 ml Pen/Strep solution
Filter and store at 4 °C - 2.5% trypsin (20 ml)
0.5 g trypsin powder dissolved in 20 ml HBSS
Filter, aliquot, and store at -20 °C - 1 mg/ml trypsin inhibitor (20 ml)
0.02 g trypsin inhibitor
Filter, aliquot, and store at -20 °C - 10 mg/ml DNase (20 ml)
0.2 g DNase I powder dissolved in 20 ml HBSS
Filter, aliquot, and store at -20 °C
Acknowledgments
We thank the R01s from the NIH (AG032051, AG020670, and NS076117 to H.Z.) for supporting this work.
References
- Bowyer, J. F., Thomas, M., Patterson, T. A., George, N. I., Runnells, J. A. and Levi, M. S. (2012). A visual description of the dissection of the cerebral surface vasculature and associated meninges and the choroid plexus from rat brain. J Vis Exp(69): e4285.
- Edmonson, C. A., Ziats, M. N. and Rennert, O. M. (2016). A non-inflammatory role for microglia in autism spectrum disorders. Front Neurol 7: 9.
- Fan, Z., Okello, A. A., Brooks, D. J. and Edison, P. (2015). Longitudinal influence of microglial activation and amyloid on neuronal function in Alzheimer’s disease. Brain 138(Pt 12): 3685-3698.
- Garden, G. A. and Campbell, B. M. (2016). Glial biomarkers in human central nervous system disease. Glia 64(10): 1755-1771.
- Henn, A., Lund, S., Hedtjarn, M., Schrattenholz, A., Porzgen, P. and Leist, M. (2009). The suitability of BV2 cells as alternative model system for primary microglia cultures or for animal experiments examining brain inflammation. ALTEX 26(2): 83-94.
- Koshimori, Y., Ko, J. H., Mizrahi, R., Rusjan, P., Mabrouk, R., Jacobs, M. F., Christopher, L., Hamani, C., Lang, A. E., Wilson, A. A., Houle, S. and Strafella, A. P. (2015). Imaging striatal microglial activation in patients with Parkinson’s disease. PLoS One 10(9): e0138721.
- Loeffler, D. A., Camp, D. M. and Bennett, D. A. (2008). Plaque complement activation and cognitive loss in Alzheimer’s disease. J Neuroinflammation 5: 9.
- Lian, H., Litvinchuk, A., Chiang, A. C., Aithmitti, N., Jankowsky, J. L. and Zheng, H. (2016). Astrocyte-microglia cross talk through complement activation modulates amyloid pathology in mouse models of Alzheimer’s disease. J Neurosci 36(2): 577-589.
- McKercher, S. R., Torbett, B. E., Anderson, K. L., Henkel, G. W., Vestal, D. J., Baribault, H., Klemsz, M., Feeney, A. J., Wu, G. E., Paige, C. J. and Maki, R. A. (1996). Targeted disruption of the PU.1 gene results in multiple hematopoietic abnormalities. EMBO J 15(20): 5647-5658.
- Michailidou, I., Willems, J. G., Kooi, E. J., van Eden, C., Gold, S. M., Geurts, J. J., Baas, F., Huitinga, I. and Ramaglia, V. (2015). Complement C1q-C3-associated synaptic changes in multiple sclerosis hippocampus. Ann Neurol 77(6): 1007-1026.
- Schwartz, M., Kipnis, J., Rivest, S. and Prat, A. (2013). How do immune cells support and shape the brain in health, disease, and aging? J Neurosci 33(45): 17587-17596.
- Stevens, B., Allen, N. J., Vazquez, L. E., Howell, G. R., Christopherson, K. S., Nouri, N., Micheva, K. D., Mehalow, A. K., Huberman, A. D., Stafford, B., Sher, A., Litke, A. M., Lambris, J. D., Smith, S. J., John, S. W. and Barres, B. A. (2007). The classical complement cascade mediates CNS synapse elimination. Cell 131(6): 1164-1178.
- Stephan, A. H., Barres, B. A. and Stevens, B. (2012). The complement system: an unexpected role in synaptic pruning during development and disease. Annu Rev Neurosci 35: 369-389.
- Stansley, B., Post, J. and Hensley, K. (2012). A comparative review of cell culture systems for the study of microglial biology in Alzheimer's disease. J Neuroinflammation 9: 115.
- Wang, Y., Cella, M., Mallinson, K., Ulrich, J. D., Young, K. L., Robinette, M. L., Gilfillan, S., Krishnan, G. M., Sudhakar, S., Zinselmeyer, B. H., Holtzman, D. M., Cirrito, J. R. and Colonna, M. (2015). TREM2 lipid sensing sustains the microglial response in an Alzheimer's disease model. Cell 160(6): 1061-1071.
- Yuan, P., Condello, C., Keene, C. D., Wang, Y., Bird, T. D., Paul, S. M., Luo, W., Colonna, M., Baddeley, D. and Grutzendler, J. (2016). TREM2 haplodeficiency in mice and humans impairs the microglia barrier function leading to decreased amyloid compaction and severe axonal dystrophy. Neuron 90(4): 724-739.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Lian, H., Roy, E. and Zheng, H. (2016). Protocol for Primary Microglial Culture Preparation. Bio-protocol 6(21): e1989. DOI: 10.21769/BioProtoc.1989.
-
Lian, H., Litvinchuk, A., Chiang, A. C., Aithmitti, N., Jankowsky, J. L. and Zheng, H. (2016). Astrocyte-microglia cross talk through complement activation modulates amyloid pathology in mouse models of Alzheimer’s disease. J Neurosci 36(2): 577-589.
Category
Neuroscience > Cellular mechanisms > Cell isolation and culture
Cell Biology > Cell isolation and culture > Cell isolation
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link