- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Cell Surface Protein Detection to Assess Receptor Internalization
Published: Vol 6, Iss 20, Oct 20, 2016 DOI: 10.21769/BioProtoc.1968 Views: 10864
Reviewed by: HongLok LungHsin-Yi ChangAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
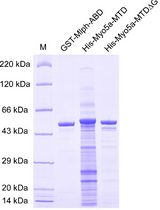
Determination of Dissociation Constants for the Interaction of Myosin-5a with its Cargo Protein Using Microscale Thermophoresis (MST)
Rui Zhou [...] Xiang-Dong Li
Feb 5, 2025 1773 Views
Cell-Sonar, an Easy and Low-cost Method to Track a Target Protein by Expression Changes of Specific Protein Markers
Sabrina Brockmöller [...] Simone Rothmiller
Feb 5, 2025 1673 Views
SiMPull-POP: Quantification of Membrane Protein Assembly via Single Molecule Photobleaching
Ryan J. Schuck [...] Rajan Lamichhane
Jan 5, 2026 344 Views
Abstract
The migration of membrane receptors upon exposure to different stimulants/inhibitors is of great importance. Among others, the internalization of membrane receptors affects their accessibility to ligands and cell responsiveness to environmental cues. Experimentally, receptor internalization can be used as a measure of their activation. In our studies, we employed this approach to explore cross-talk between a seven transmembrane domain receptor for neuropeptide Y (NPY), Y5R, and a tyrosine kinase receptor for brain-derived neurotrophic factor (BDNF), TrkB. To this end, we measured the internalization of Y5R upon stimulation with the TrkB ligand, BDNF. Upon treatment with BDNF, the cells were exposed to a membrane impermeable, biotinylation reagent that selectively labels surface proteins. Subsequently, the biotinylated membrane proteins were affinity-purified on columns with avidin resins and analyzed by Western blot. Differences in the fraction of receptors present on the cell surface of control and ligand-treated cells served as a measure of their internalization and response to particular stimuli.
Keywords: Membrane receptorsBackground
Cell membrane receptor internalization in response to external stimuli can be measured using two major strategies – microscopic and biochemical. The most common approach is the use of microscopy – either in real-time or on fixed cells. In the first approach, the cells expressing receptors labelled with fluorescent tags (e.g., fused to the fluorescent proteins) are examined in live cells by time-lapse confocal microscopy. Alternatively, cells expressing fluorescently labeled receptors can be exposed to the desired stimuli and then fixed at a pre-defined time. Subsequently, sub-cellular localization of these receptors (i.e., membrane vs. intracellular fraction) is examined by fluorescence microscopy and compared with the untreated control. The advantage of time-lapse microscopy is the ability to examine the same cells at different time points and directly assess changes in the receptor distribution upon stimulation (Czarnecka et al., 2015). However, since this assessment has to be performed under high magnification, the number of cells that can be analyzed is limited and the response is not always uniform among the cells. On the other hand, fixing the cells upon stimulation allows for examining a larger cell population and for analysis of the native, not-labeled receptors, if combined with fluorescently labeled ligands or immunocytochemistry (Bohme et al., 2008; Fabry et al., 2000). However, in this case, the analysis of the receptor sub-cellular localization is usually qualitative and the time of exposure may not be optimal, as the changes are not examined in real time.
The biochemical approach takes advantage of cell-impermeable biotinylation reagents that selectively cross-link extracellular domains of cell surface receptors. The biotin-labeled cell membrane proteins are then affinity-purified and the receptor of interest can be selectively detected by Western blot (Czarnecka et al., 2015). This approach allows for quantitative analysis of the cells as a whole population and does not require fusion with a fluorescent protein that may potentially change the behavior of the tested receptors. However, as with microscopic analysis of fixed cells upon treatment, the time of exposure to the ligand remains to be determined. Therefore, in our study, we combined time-lapse confocal microscopy, which allowed us to perform the initial assessment of the internalization rate and determine the time of ligand exposure allowing for detecting maximal changes in receptor sub-cellular localization, and the subsequent selective isolation of cell surface receptors at this time point to achieve quantitative results and confirm microscopic observations (Czarnecka et al., 2015). This strategy was successful in demonstrating neuropeptide Y (NPY) Y5R receptor internalization upon stimulation with non-cognate ligand, brain-derived neurotrophic factor (BDNF), and therefore proving the interactions between NPY and BDNF systems.
Materials and Reagents
- Cell scrapers (Corning, Falcon®, catalog number: 353085 )
- 150 mm tissue culture-treated dishes (Corning, catalog number: 430599 )
- 50 ml centrifuge conical tubes (Corning, catalog number: 430829 )
- 15 ml centrifuge conical tubes (Corning, catalog number: 430791 )
- 1.5 ml centrifuge tube
- Nitrocellulose membrane (GE Healthcare, catalog number: 10600011 )
- SH-SY5Y neuroblastoma cells
- Puncture foil
- RPMI media (ATCC, catalog number: 30-2001 )
- Fetal bovine serum (FBS) (Thermo Fisher Scientific, GibcoTM, catalog number: 10437-028 )
- Geneticin (G 418 disulfate salt) (Sigma-Aldrich, catalog number: A1720 )
- Fungizone (Amphotericin B) (Thermo Fisher Scientific, GibcoTM, catalog number: 15290018 )
- Penicillin-streptomycin (Thermo Fisher Scientific, GibcoTM, catalog number: 15140122 )
- Protease inhibitor cocktail (Sigma-Aldrich, catalog number: P8340-1 ml )
- Bio-Rad protein assay dye reagent concentrate (Bio-Rad Laboratories, catalog number: 5000006 )
- Cell surface protein isolation kit (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 89881 ) containing the following reagents
- Sulfo-NHS-SS-biotin
- Phosphate buffered saline (PBS) pack to be reconstituted in ultrapure water
- Quenching solution
- Tris buffered saline (TBS) pack to be reconstituted in ultrapure water
- Lysis buffer
- Immobilized NeutrAvidin gel
- Wash buffer
- Columns and caps
- No-weigh bithiothreitol (DTT)
- Ultra-pure water (Sigma-Aldrich, catalog number: W4502 )
- SDS-PAGE sample buffer (PierceTM lane marker non-reducing) (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 39001 )
- 4-20% Tris-glycine gels (Thermo Fisher Scientific, InvitrogenTM, catalog number: EC60252BOX )
Note: This product has been discontinued. - Blotting-grade blocker (nonfat dry milk) (Bio-Rad Laboratories, catalog number: 1706404 )
- TweenTM 20, Fisher BioReagentsTM (Thermo Fisher Scientific, Fisher Scientific, catalog number: BP337-100 )
- Goat polyclonal anti Y5R antibody (Everest Biotech, catalog number: EB06769 )
- Donkey anti-goat antibody, horseradish peroxidase conjugated (Santa Cruz Biotechnology, catalog number: sc-2020 )
- SuperSignalTM West Pico chemiluminescent substrate (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 34080 )
Equipment
- Centrifuge for 15 ml tubes (BD, AdamsTM, model: DYNACTM Centrifuge 0101 )
Note: Equipment discontinued. Can be replaced by any centrifuge holding 15 ml tubes. - Centrifuge for Eppendorf tubes (Eppendorf, model: Centrifuge 5417 C )
Note: Equipment discontinued. Can be replaced by any centrifuge holding 1.5 ml tubes. - Rocking platform (Boekel Scientific, catalog number: 260350 )
- Sonicator (Thermo Fisher Scientific, model: Sonic Dismembrator 550 )
Note: Equipment discontinued. Can be replaced by any other sonicator.
Procedure
- Treatment
- Plate the cells to be tested in a 150 mm culture dish to obtain 80-90% confluency the following day (do not exceed 4 x 107 cells per labeling reaction).
Note: In our study, we used SH-SY5Y neuroblastoma cells stably transfected with TrkB receptor cultured in RPMI media supplemented with 10% fetal bovine serum, 0.3 mg/ml of geneticin, 0.5 µg/ml fungizone, 100 U/ml penicillin and 100 µg/ml streptomycin (Ho et al., 2002). The cells were plated at a density of 2 x 106 cells/plate. - On the following day, replace the standard cell culture media with media that does not contain serum (serum free media).
- 24 h later treat cells as desired.
Note: In our study, we treated the cells with NPY at a concentration of 10-7 M and BDNF at 1 ng/ml for 8 min (Czarnecka et al., 2015). Untreated cells served as a control.
- Plate the cells to be tested in a 150 mm culture dish to obtain 80-90% confluency the following day (do not exceed 4 x 107 cells per labeling reaction).
- Biotinylation
- Dissolve the contents of 1 vial containing 12 mg of Sulfo-NHS-SS-biotin in 48 ml of ice-cold PBS to obtain a final concentration of 0.25 mg/ml biotin in the biotin-labeling solution.
- Upon the desired treatment, remove media and wash cells twice with 10 ml of ice-cold PBS per dish. Remove PBS immediately after washing.
Note: Do not allow PBS to remain in contact with the cells for more than 5 sec to prevent rounding and detachment of cells. - Add 12 ml of biotin-labeling solution to each dish.
- Place culture dishes on a rocking or orbital shaker and gently agitate for 30 min at 4 °C (cells must be covered with the biotin-labeling solution).
- Add 600 µl of quenching solution (kept and applied at 4 °C) to each culture plate to quench reaction. Ensure even coverage of the cells by gently tilting the plate and subsequently proceed to the next step.
- Gently scrape cells into solution and transfer the contents to corresponding 50 ml conical tubes. Rinse each plate with a single 4 ml volume of ice-cold TBS. Add the cells remaining in the rinse solution to previously collected cells in the 50 ml conical tubes corresponding to the appropriate treatment group.
- Centrifuge cells at 500 x g for 3 min at 4 °C.
- Discard supernatant.
- Add 5 ml of ice-cold TBS and re-suspend cells by gentle pipetting.
- Centrifuge cells again for 3 min at 4 °C.
- Discard supernatant.
- Dissolve the contents of 1 vial containing 12 mg of Sulfo-NHS-SS-biotin in 48 ml of ice-cold PBS to obtain a final concentration of 0.25 mg/ml biotin in the biotin-labeling solution.
- Cell lysis
- Supplement the desired amount of lysis buffer (number of samples x 370 µl + 10% excess by volume) with protease inhibitor cocktail at a 1:100 ratio.
- Re-suspend cells in 370 µl of lysis buffer by pipetting and then transfer to a tube that is fitting for the sonication process (in our case 15 ml tube).
- Sonicate (1.5 power) 5 x 1-sec bursts.
- Incubate cells for 30 min on ice, and during that period:
- Vortex every 5 min for 5 sec.
- Sonicate additionally every 10 min on ice.
- Transfer clarified supernatant to a 1.5 ml centrifuge tube. At this stage:
- Measure protein concentration using the Bradford method (Bio-Rad protein assay) according to the manufacturer’s protocol.
- Normalize protein concentrations for all samples and use equal volumes of these normalized lysates for subsequent steps (approx. 400 µl).
Note: Our protein of interest is expressed at low levels; therefore we used the highest possible protein concentration, normalized to the least concentrated sample. However, we suggest to adjust the protein amount used in the assay according to the expression level of tested protein based on previous experiments, or to establish the proper concentration in a separate pilot experiment.
- Set aside 30 µl of each normalized lysate to be run on a gel as a loading control. Use the remaining volume (approx. 370 µl) for the isolation of biotinylated cell surface proteins (step D6 below).
- Supplement the desired amount of lysis buffer (number of samples x 370 µl + 10% excess by volume) with protease inhibitor cocktail at a 1:100 ratio.
- Isolation of labeled proteins
- Insert a column into a collection tube.
- Add 350 µl of NeutrAvidin agarose gel to each column and cap the column.
- Centrifuge for 1 min at 1,000 x g, and then discard flow-through (FT).
- Add 400 µl of wash buffer to the gel.
- Centrifuge for 1 min at 1,000 x g, and then discard FT.
- Apply bottom cap to each column and add cell lysate to agarose; apply top cap to each column (wrap each cap in parafilm, as shown in Figure 1).
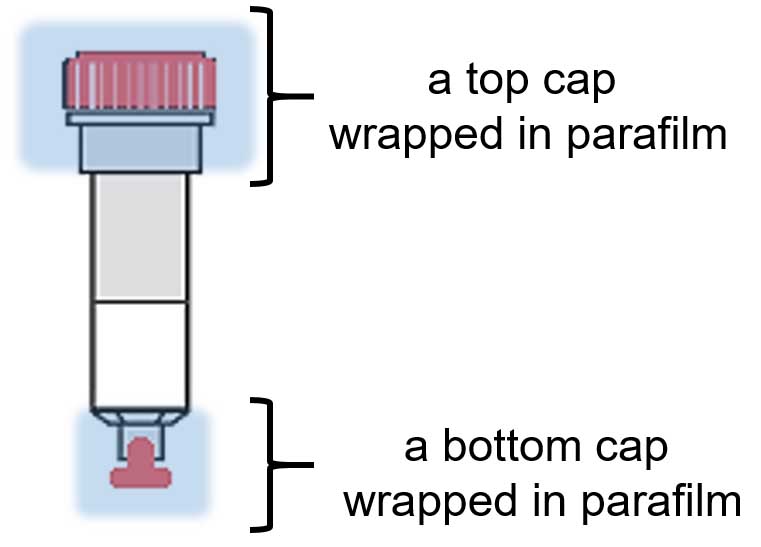
Figure 1. The sketch of a single column with applied bottom and top caps (in red) wrapped in parafilm (blue)
- Incubate for 60 min at room temperature, mixing throughout by placing columns on a rotator/rocking platform.
- Add protease inhibitor cocktail to the desired amount of the wash buffer calculated according to the following formula: number of samples x 4 washing steps x 400 µl + an excess of 10% by volume.
- Remove bottom caps and place columns in their collection tubes.
- Centrifuge columns for 1 min at 1,000 x g, and then discard FT.
- Return columns to collection tubes, remove the top caps and add 400 µl of wash buffer to each column.
- Apply both caps to columns and mix content by inverting columns.
Note: To simplify this step, in place of a bottom cap, a piece of parafilm can be held (a fresh piece needs to be used for each sample in each step). - Remove bottom caps and centrifuge for 1 min at 1,000 x g, and then discard FT.
- Remove top caps.
- Repeat steps D11-D14 three times.
- Replace bottom cap on columns.
- Insert a column into a collection tube.
- Protein elution
- Puncture foil covering one No-weigh DTT microtube with a pipette tip, and add 50 µl of ultrapure water to obtain 1 M DTT.
- Prepare the loading buffer by combining the desired volume of SDS-PAGE sample buffer (number of samples x 150 µl + 10% excess by volume) with 1 M DTT solution at a 1:20 ratio to obtain a final concentration of 50 mM DTT.
- Add 150 µl of SDS-PAGE sample buffer containing DTT to each column (the resins must be covered), and then cap columns from top and bottom.
- Incubate columns for 60 min at room temperature on a rocking platform.
Note: Alternatively, as mentioned in the original protocol provided in the kit, samples can be heated in a heat block for 5 min at 95 °C. However, it will lead to the recovery of some NeutrAvidin protein monomer (15 kDa) in the eluates. - Remove a column’s top cap first, and then the bottom cap; place columns in new collection tubes and apply a top cap.
Note: The sequence of caps removal is particularly important in the case of elution at a high temperature (step E4 above), as it prevents eluate leakage from the tubes. - Centrifuge for 2 min at 1,000 x g.
- Analyze eluates and input samples by Western blot (see below). Store samples at -20 °C if not used immediately.
- Puncture foil covering one No-weigh DTT microtube with a pipette tip, and add 50 µl of ultrapure water to obtain 1 M DTT.
- Protein separation and target protein identification by Western blot
- Denature eluates by incubation in boiling water for 2 min.
Note: Eluates are already in SDS-PAGE sample buffer used to elute the biotinylated proteins from columns, as described in the ‘Protein elution’ section. - In addition, prepare the previously obtained lysates for Western blotting by adding SDS-PAGE loading buffer with reducing agent (50 mM DTT) and denature samples, as described in point 1 above.
- Separate both fractions (namely, eluates and lysates) on 4-20% Tris-glycine gels and transfer to a nitrocellulose membrane according to the standard procedures.
- Block the nitrocellulose membrane with 5% milk in Tris-buffered saline with 0.2% of Tween 20 (TBST).
- Blot with the desired primary and secondary antibody to detect the surface receptor of interest.
Note: In our study, we used goat polyclonal anti-Y5R antibody diluted at 1:1,000 in 5% milk/TBST followed by donkey anti-goat-horseradish peroxidase-linked antibody diluted at 1:25,000. - Develop immunoblots with SuperSignal West Pico chemiluminescent substrate. Optionally, the more sensitive Femto chemiluminescent substrate can be used if a weak signal is expected.
- Denature eluates by incubation in boiling water for 2 min.
Data analysis
In the procedure of cell surface protein extraction, cells are exposed to a membrane impermeable, biotinylation reagent. Subsequently, cells are lysed, and the crosslinked membrane proteins are affinity-purified on columns with avidin resins. Kit functionality was validated by the manufacturer, based on the lack of representative intracellular proteins (Heat shock protein 90, Hsp90; calnexin) in the eluates analyzed by Western blot. Our experiments confirmed this observation. While extracellular membrane proteins, such as Y5R, were readily detectable in both lysate and eluate fractions, the eluates were markedly depleted in intracellular proteins, such as cytoplasmic p42/44 mitogen-activated protein kinase (MAPK) and a mitochondrial marker, apoptosis-inducing factor (AIF) (Figure 2). Of note, based on our protocol, the eluates are 5 fold concentrated by volume, as compared to the lysates.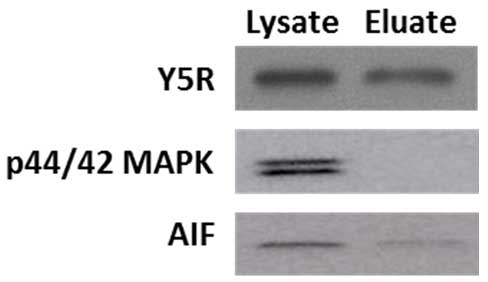
Figure 2. Cell surface protein preparation is depleted in intracellular proteins. A cell membrane protein, Y5 receptor (Y5R), is readily detectable in the original lysates and eluates containing cell surface proteins, while the latter fraction is depleted in cytoplasmic p42/44 mitogen-activated protein kinase (MAPK) and mitochondrial apoptosis-inducing factor (AIF). The proteins loaded on the gel correspond to 0.04 and 0.2 of the original lysate volume for the lysate and eluate fraction, respectively, indicating that the eluate fraction is 5 times concentrated.
We used the above procedure to provide evidence for cross-talk between NPY receptor Y5R and BDNF receptor, TrkB. To this end, SHSY5Y neuroblastoma cells stably transfected with TrkB receptor were treated with NPY or BDNF for 8 min (Czarnecka et al., 2015; Ho et al., 2002). The time of stimulation and ligand concentrations were selected based on previous time-lapse microscopy experiments (Czarnecka et al., 2015). Upon ligand treatment, the surface proteins were labelled, immunoprecipitated and analyzed by Western blot for Y5R. As shown in Figure 3, NPY stimulation resulted in a modest decrease in the surface fraction of Y5R, while BDNF treatment triggered more profound Y5R internalization. These data were in agreement with the results of parallel experiment in which receptor internalization was monitored by time-lapse microscopy (Czarnecka et al., 2015).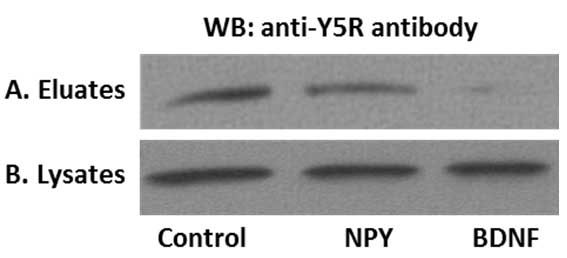
Figure 3. Representative results of the surface protein immunoprecipitation experiment. A. Y5 receptor (Y5R) cell membrane expression in SY5Y/TrkB transfectants. Under control conditions Y5R was present on the cell membrane and the addition of NPY, as well as BDNF decreased the cell membrane pool of Y5R. B. The total Y5R levels do not differ in cell lysates, confirming equal protein input.
Acknowledgments
This work was supported by National Institutes of Health (NIH) grants: 1RO1CA123211, 1R03CA178809, R01CA197964 and 1R21CA198698 to JK. The protocol adapted from Czarnecka et al. (2015).
References
- Bohme, I., Stichel, J., Walther, C., Morl, K. and Beck-Sickinger, A. G. (2008). Agonist induced receptor internalization of neuropeptide Y receptor subtypes depends on third intracellular loop and C-terminus. Cell Signal 20(10): 1740-1749.
- Czarnecka, M., Trinh, E., Lu, C., Kuan-Celarier, A., Galli, S., Hong, S. H., Tilan, J. U., Talisman, N., Izycka-Swieszewska, E., Tsuei, J., Yang, C., Martin, S., Horton, M., Christian, D., Everhart, L., Maheswaran, I. and Kitlinska, J. (2015). Neuropeptide Y receptor Y5 as an inducible pro-survival factor in neuroblastoma: implications for tumor chemoresistance. Oncogene 34(24): 3131-3143.
- Fabry, M., Langer, M., Rothen-Rutishauser, B., Wunderli-Allenspach, H., Hocker, H. and Beck-Sickinger, A. G. (2000). Monitoring of the internalization of neuropeptide Y on neuroblastoma cell line SK-N-MC. Eur J Biochem 267(17): 5631-5637.
- Ho, R., Eggert, A., Hishiki, T., Minturn, J. E., Ikegaki, N., Foster, P., Camoratto, A. M., Evans, A. E. and Brodeur, G. M. (2002). Resistance to chemotherapy mediated by TrkB in neuroblastomas. Cancer Res 62(22): 6462-6466.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Czarnecka, M. and Kitlinska, J. (2016). Cell Surface Protein Detection to Assess Receptor Internalization. Bio-protocol 6(20): e1968. DOI: 10.21769/BioProtoc.1968.
Category
Cancer Biology > General technique > Biochemical assays > Protein analysis
Biochemistry > Protein > Interaction > Protein-protein interaction
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











