- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Methods for Detecting Microbial Methane Production and Consumption by Gas Chromatography
(*contributed equally to this work) Published: Vol 6, Iss 7, Apr 5, 2016 DOI: 10.21769/BioProtoc.1779 Views: 11069
Reviewed by: Valentine V TrotterLaura Molina-GarcíaAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Soluble and Solid Iron Reduction Assays with Desulfitobacterium hafniense
Lucrezia Comensoli [...] Edith Joseph
Sep 5, 2018 6737 Views
In vivo Quantification of Alkanes in Escherichia coli
Tabinda Shakeel [...] Syed Shams Yazdani
Apr 20, 2020 4808 Views
A SsrA/NIa-based Strategy for Post-Translational Regulation of Protein Levels in Gram-negative Bacteria
Gonzalo Durante-Rodríguez [...] Pablo I. Nikel
Jul 20, 2020 4394 Views
Abstract
Methane is an energy-dense fuel but is also a greenhouse gas 25 times more detrimental to the environment than CO2. Methane can be produced abiotically by serpentinization, chemically by Sabatier or Fisher-Tropsh chemistry, or biotically by microbes (Berndt et al., 1996; Horita and Berndt, 1999; Dry, 2002; Wolfe, 1982; Thauer, 1998; Metcalf et al., 2002). Methanogens are anaerobic archaea that grow by producing methane gas as a metabolic byproduct (Wolfe, 1982; Thauer, 1998). Our lab has developed and optimized three different gas chromatograph-utilizing assays to characterize methanogen metabolism (Catlett et al., 2015). Here we describe the end point and kinetic assays that can be used to measure methane production by methanogens or methane consumption by methanotrophic microbes. The protocols can be used for measuring methane production or consumption by microbial pure cultures or by enrichment cultures.
Materials and Reagents
- Balch tubes (Bellco Glass Inc., catalog number: 2048-00150 )
- Butyl Rubber Stoppers (Bellco Glass Inc., catalog number: 2048-11800A )
- 11 mm Aluminum Seal Crimps (Wheaton, catalog number: 224176-01 ) (Figure 1A)
- 20 mm Aluminum Seal Crimps (Wheaton, catalog number: 224178-01 )
- 18 G and 22 G BD PrecisionGlide Needle (Becton Dickinson, catalog number: 305195 )
- 22 G BD PrecisionGlide Needle (Becton Dickinson, catalog number: 305155 )
- Hamilton Gas Tight Syringe [1705 Sl 50 μl Syr (22s, 2”, 2) L] (Hamilton Company, catalog number: 80956 ) (Figure 1)
- Autosampler vials (National Scientific, catalog number: G4012-1W )** (Figure 1A)
- 11 mm straight plug stopper, natural red rubber (Wheaton, catalog number: 224100-030 )** (Figure 1A)
- UHP 99.99% Methane Gas Tank (Airgas, model: LW908 )
- Nalgene Labtop cooler (Sigma-Aldrich, catalog number: C2312-1EA )
- Nalgene Labtop cooler, Jr. (Sigma-Aldrich, catalog number: C2437-1 EA )
- 1 ml, 5 ml and 10 ml BD TB Syringes (Becton Dickinson, catalog number: 309624 , 309632 and 309640 )
- 15 ml sterile polypropylene Falcon conical centrifuge tubes (Corning, catalog number: 352196 )
- Sterile PVDF syringe filter (17 mm diameter, 0.2 μm pore size) (Thermo Fisher Scientific, catalog number: F25136 )
- 20 ml BD Luer Lok Disposable syringe (Becton Dickinson, catalog number: 302830 )
- 16 x 132 mm mm Type 1 glass A borosilicate glass test tubes (Bellco Glass Inc., catalog number: 2011-16125 )**
- 16 mm KAP-UTS test tube caps, various colors (Bellco Glass Inc., catalog number: 2007-16005 )**
- 1 ml micropipette (Mettler-Toledo, Rainin, model: L-1000 )*
- 200 μl micropipette (Mettler-Toledo, Rainin, model: L-200 )*
- 20 μl micropipette (Mettler-Toledo, Rainin, model: L-20 )*
- 20 μl HydroLogix SoftFit-L Pipet Tips (VWR International, catalog number: 89031-366 )*
- 200 μl HydroLogix SoftFit-L Pipet Tips (VWR International, catalog number: 89031-388 )*
- 1 ml HydroLogix SoftFit-L Pipet Tips (VWR International, catalog number: 89031-430 )*
- Envision paper towels (Georgia-Pacific, catalog number: 23504 )
- Hamilton gastight tapered syringe as injection needle (Hamilton Company, catalog number: 5181-8809 ) (Figure 1)
- UHP Air Gas Tank with regulator plumbed to GC (Matheson Tri-Gas®, model: SG SPPULW700 )
- UHP Helium Gas Tank with regulator plumbed to GC (Matheson Tri-Gas®, model: SG SPPULW800P )
- UHP Nitrogen Gas Tank with regulator plumbed to GC (Matheson Tri-Gas®, model: LW 411P )
- UHP Hydrogen Gas Tank with regulator plumbed to GC (Matheson Tri-Gas®, model: SG SPPULW500P )
- ddH2O
- Mupirocin (Sigma-Aldrich, catalog number: M7694 )**
- Sodium hydroxide (NaOH) anhydrous pellets (Sigma-Aldrich, catalog number: S8045-500 G )
- HS culture medium
- 3-N-(morpholino) propanesulfonate (MOPS) (pH 6.8)
- UHP 100% nitrogen (Matheson Tri-Gas®, model: SG SPPULW411 )
- 50 mM methanol
- General preparation of anaerobic solutions (see Recipes)
- 200x mupirocin stock (see Recipes)
- Plain medium (no C source) (see Recipes)
- 2x C medium (see Recipes)
Notes:- *Materials and reagents are only used for anaerobic condition.
- ** Materials and reagents can be used for both anaerobic and aerobic conditions.
- *Materials and reagents are only used for anaerobic condition.
Equipment
- 250 ml beaker (Thermo Fisher Scientific, catalog number: FB-100-250 )
- Anaerobic chamber or glove box with (Coylab, model: type B ) (Figure 2)
- Agilent 7890A Gas Chromatograph with Flame Ionization detector (Agilent Technologies, model: G3440A )
- Agilent Autosampler (Agilent Technologies, model: G4513A )
- Agilent Technologies OpenLAB CDS ChemStation Edition Rev C.01.02
- GS-CarbonPLOT GC Column (Agilent Technologies, catalog number: 113-3132 )
- Agilent Liner 4mm ID tap GW (Agilent Technologies, catalog number: 5062-3587 )
- Merlin Microseal High-Pressure Replacement Septum (Restek Corporation, catalog number: 22812 )
- IEC Medilite Microcentrifuge (Thermo Fisher Scientific, catalog number: 004480F ) (Figure 2)
Note: Equipment is used for anaerobic condition. - Spectronic 20D+ (Thermo Fisher Scientific, catalog number: 14-385-129 )
Software
- Agilent OpenLAB CDS ChemStation software
Procedure
- Preparing a standard curve
- Prep autosampler vials by flushing with air. Stopper, crimp and label vials (Figure 1A).
- Open gas valve and regulator of 99.99% methane gas tank. Regulator is fitted with a stem that allows a needle to fit into the tip (Figure 3). The stem is fitted with an air tight septa to not allow undesired methane gas to leak out of the tank.
- Using a 50 µl gastight Hamilton syringe (Reno, NV), insert the needle into the end of the regulator (Figure 3). Be sure that the stopcock is in the off position before inserting the needle. Using two fingers on either side of the needle, help guide needle in to prevent the needle from bending.
- Once the Hamilton syringe is inserted past the septa, open the stopcock and withdraw more than the desired amount of methane (i.e., If needing 50 µl of CH4, withdraw about 55 µl). Be careful not to allow plunger of the syringe to come out of syringe completely.
- Close the stopcock and withdraw the needle from end of the regulator (Figure 2B-2C).
- Open the stopcock and dispel gas from syringe to the desired volume.
- Allow gas to equilibrate to atmospheric pressure and then close the stopcock.
- Inject methane gas into stoppered/crimped autosampler vial.
- Run methane capture method on Agilent GC (details described below).
- Suggested volumes of 99.99% methane to inject into standard vials: 5 µl, 10 µl, 20 µl, 30 µl, 40 µl, and 50 µl (Figure 4).
Table 1. Gas chromatograph “Methane” method settingsInlet Mode Splitless Purge flow 60 ml/min at 0.75 min Heater 250 °C Pressure 31.529 psi Septum purge flow 3 ml/min standard Column Column type GS CarbonPLOT Flow 6.5 ml/min Pressure 31.529 psi Ave. velocity 85.265 cm/sec Constant flow Yes Post run 7.3213 ml/min ALS injection volume 2 µl Oven Temperature 145 °C Hold time 3 min FID detector Heater 300 °C H2 flow 30 ml/min Air flow 400 ml/min Makeup flow (N2) 25 ml/min Flame On
- Prep autosampler vials by flushing with air. Stopper, crimp and label vials (Figure 1A).
- Calculations
- Table S1. GC Methane Calculations guides the user in making a methane standard to determine the amount of moles of methane produced by a culture. Within the spreadsheet, the “Variables” tab contains constants and variables used in the calculation. The constants used conform to Beadle Center, University of Nebraska-Lincoln, but can be modified to accommodate other atmospheric and temperature conditions.
- A standard curve needs to be generated each time the FID detector is turned on. Once the standard has been created, export the following data to Microsoft Excel: Date and Time, Sample_Name, Vial #, Retention Time (min), Height, Area, and Injection_DataFileDirectory. Copy and paste the exported data into the attached spreadsheet’s “Standard Data” tab, making sure numbers do not transfer as text and everything is in the correct column. If using Agilent OpenLAB CDS ChemStation software, a report layout can be created that only includes these fields in the order of the spreadsheet.
- Once the “Standard Data” tab is filled out, fill out the “Constants” tab with injection volume, vial volume, and the volumes of methane gas added to each standard vial, corresponding to what was entered on the “Standards” tab. The spreadsheet will complete the calculations. The standard curve created by default is in Peak Area vs Methane (nmoles). It is important to note that this standard curve is the amount of methane injected into the GC (2 µl injected from each autosampler vial). This can be used in a direct comparison to 2 µl sampled in the same way elsewhere. The number of nmoles has not been multiplied by the dilution factor that would represent the number of moles in the entire autosampler vial. (For a 1.99 ml autosampler vial, the dilution factor is 995.)
- When using the standard curve created by the attached spreadsheet, data obtained directly from the GC can be compared. When completing the End Point Assay, the peak areas obtained from each sample can be directly plugged into the standard curve equation. In a Kinetic Assay, the volume needs to be adjusted to account for the vial volume displaced by the cell suspension (intact resuspended cells). Once the amount of methane detected by the GC is calculated, dilution factors must be used to calculate the total amount of methane in the headspace of the given culture.
- Table S1. GC Methane Calculations guides the user in making a methane standard to determine the amount of moles of methane produced by a culture. Within the spreadsheet, the “Variables” tab contains constants and variables used in the calculation. The constants used conform to Beadle Center, University of Nebraska-Lincoln, but can be modified to accommodate other atmospheric and temperature conditions.
- End point assay
- Grow pure or enrichment cultures in Balch tubes to the desired optical density. For example, for pure cultures of Methanosarcina acetivorans C2A grown at 35 °C in HS medium with 125 mM methanol as carbon and energy source with a 1:100 inoculum early stationary phase (OD600 = ~ 0.9) is reached between 50 and 75 h (Catlett et al., 2015).
- Prepare autosampler vials by flushing with air. Stopper, crimp, and label vials.
- In the fume hood, take culture tubes and insert an 18 G needle three quarters of the way through the blue butyl stopper (Figure 5A). Do not push the needle through the stopper completely.
- Using a gas tight Hamilton syringe, insert the syringe needle into the 18 G needle (Figure 5B). Be sure that the Hamilton syringe stopcock is closed at this time (Figure 2C). The 18 G needle is used as a guide to help puncture the butyl stopper with the Hamilton syringe needle while keeping the syringe needle straight.
- Push the Hamilton syringe needle through the butyl blue stopper until the end of the needle becomes visible in the headspace of the culture tube (Figure 5B). Once again, do not push the 18 G needle completely through the stopper.
- Open the stopcock of the Hamilton syringe. Quickly and steadily withdraw the Hamilton syringe plunger to the desired volume of headspace. (When using the standard curve described above, it is best to withdraw 50 µl of headspace from the culture.) Once at the desired volume of gas, quickly turn the stopcock of the Hamilton syringe, sealing the gas inside the syringe. Do not let the gas escape to equilibrate with atmospheric pressure (as in section A). Depending on the pressure in the growing culture, the plunger on the Hamilton syringe may continue to move as the gas expands in the syringe.
- Withdraw the Hamilton syringe and 18 G needle from the culture’s butyl stopper.
- With the Hamilton stopcock still closed, insert the Hamilton syringe needle into a stoppered and crimped autosampler vial.
- Open the stopcock and inject all the gas from the syringe into the autosampler vial.
- Close the stopcock and withdraw the Hamilton syringe from the autosampler vial.
- Once the headspace from all the cultures has been collected, place the vials in the autosampler.
- Run the “Methane” method on the GC (Table 1).
- Grow pure or enrichment cultures in Balch tubes to the desired optical density. For example, for pure cultures of Methanosarcina acetivorans C2A grown at 35 °C in HS medium with 125 mM methanol as carbon and energy source with a 1:100 inoculum early stationary phase (OD600 = ~ 0.9) is reached between 50 and 75 h (Catlett et al., 2015).
- Kinetic assay
- Before beginning, have the following materials and reagents completely anaerobic and ready in the chamber: Mupirocin stock (3.5 mg/ml) (or appropriate protein synthesis inhibitor depending on the susceptibility profile of the organisms you are assaying), sterile 16 mm test tubes, autosampler vials, autosampler stoppers and crimps, IEC Medilite Microcentrifuge. Prepare a stock of plain medium (no carbon source), and a second stock of growth medium with twice the concentration of carbon source (2x C medium). It is recommended to store these stocks in the anaerobic chamber.
- Grow cultures in Balch tubes to desired optical density (exponential phase OD600 = ~0.3-0.5). Record OD of the culture.
- Bring into the anaerobic chamber: Balch tube cultures in a 4 °C Nalgene Labtop Cooler, sterile labeled microcentrifuge tubes, autosampler crimper, liquid waste container, solid waste container, autosampler vial rack, and forceps.
- Measure the amount of plain medium and 2x C medium required for the experiment (Table 2). Add protein synthesis inhibitor to desired concentration. For methanogens, add mupirocin to 70 µM.
- Once items are brought into the chamber, gently resuspend any settled cells in the 10 ml culture. Withdraw 5 ml using a syringe and place in a sterile test tube. Withdraw the remaining 5 ml and place in a separate test tube.
- Spin down cell cultures in the centrifuge for 5 min at 1,228 x g.
- Decant supernatant and resuspend each pellet in 5 ml/test tube of plain medium. Use the syringe to disrupt the pellet and spin gently to avoid lysing cells.
- Spin the resuspended cells for 5 min in the centrifuge at 1,228 x g.
- As the centrifuge is running, place autosampler vials on an autosampler vial rack. Seven vials are required to assay one strain: five vials for sample replicates, a medium-only control (medium without cells) and a no substrate cells-only control (cells in plain medium only, no carbon source).
- For 500 µl cell suspensions, Add 250 µl 2x C medium to the five sample replicates and to the medium-only control. Add 250 µl of plain medium to both the medium-only and to the no-cells controls (Table 2).
- When the centrifuge has halted, immediately decant supernatant. Tap the test tube onto a piece of paper towel to remove all residual media.
- Use 2 ml plain medium to resuspend and combine pellets into one tube. This should be 2 ml total resuspension for each 10 ml Balch tube culture. Keep cold in a 4 °C Nalgene Labtop Cooler.
- Add 250 µl of cell suspension to the sample replicates and the cells-only control.
- Place 200 µl of leftover cell suspension into a labeled microcentrifuge tube. This will be used to measure protein concentration by Bradford assay.
- Crimp and label vials. Forceps can be helpful in placing stoppers and crimps onto each vial.
- Remove the vials from the chamber and place in a 35 °C incubator for 5 min before placing vials in the GC autosampler.
- Do six runs of a seven-vial sequence using the “Methane” method on Agilent GC (Table 1). Each vial should be run in order before repeating the sequence for the next measurement, resulting in 20-40 min for methane to accumulate in each vial between measurements.
- To measure protein concentration of the cell suspensions, spin down the 200 µl cell suspension saved in step 14 at 1,500 x g for 3 min. Remove supernatant and resuspend with 200 µl ddH2O. Lyse cells and perform a Bradford with Coomassie reagent and 2 mg BSA standard. Methanogen cells grown in HS medium are easily lysed by resuspension in ddH2O by osmotic shock, but organisms grown in low-osmolarity medium or that have cell walls may require boiling and/or freeze-thaw and vortexing or sonicating to fully lyse.
- Graph the methane peak area vs. time for each sample (Figure 6). Use the slopes to calculate the amount of methane produced or consumed with time (Table S1.).
Table 2. Kinetic assay medium volumes and controls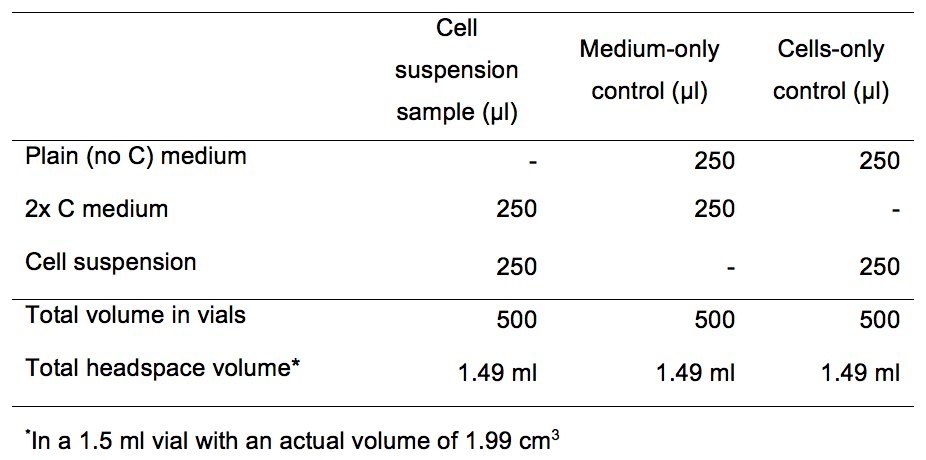
- Before beginning, have the following materials and reagents completely anaerobic and ready in the chamber: Mupirocin stock (3.5 mg/ml) (or appropriate protein synthesis inhibitor depending on the susceptibility profile of the organisms you are assaying), sterile 16 mm test tubes, autosampler vials, autosampler stoppers and crimps, IEC Medilite Microcentrifuge. Prepare a stock of plain medium (no carbon source), and a second stock of growth medium with twice the concentration of carbon source (2x C medium). It is recommended to store these stocks in the anaerobic chamber.
Representative data
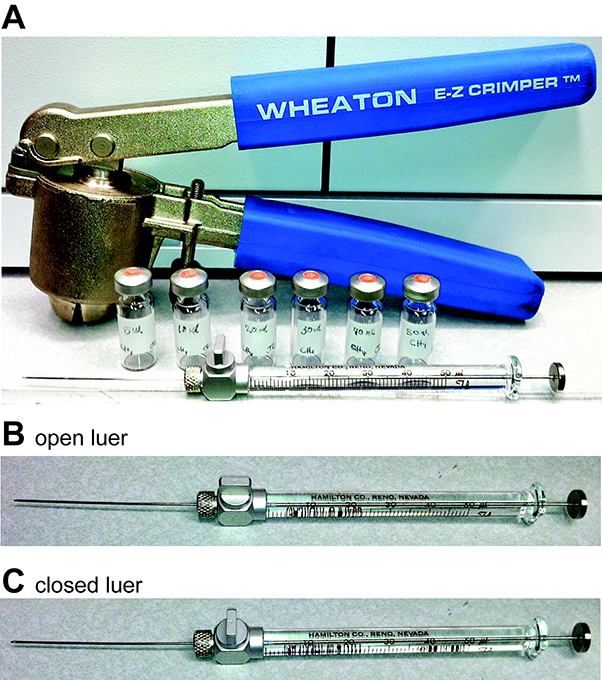

Figure 1. Crimper, gastight autosampler vials and gas-tight Hamilton syringes. A. Crimpers are used to seal autosampler vials using aluminum crimps and rubber stoppers. Hamilton syringes showing open (B) and closed (C) luer fittings.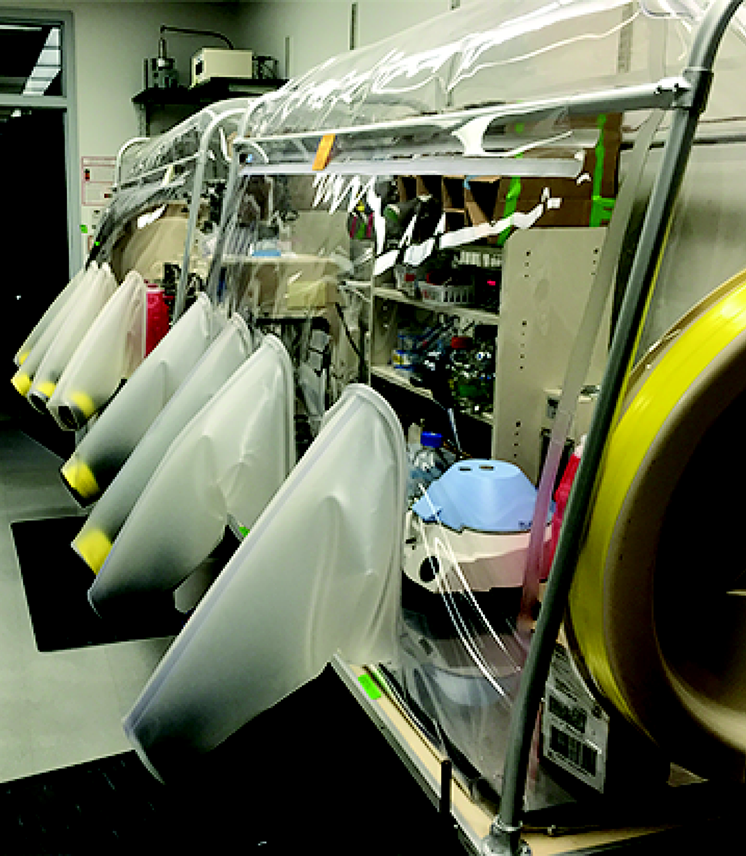
Figure 2. Dual-sided custom Coy anaerobic chamber showing Medlite clinical centrifuge (blue lid)
Figure 3. Methane gas tank fitted with a septa 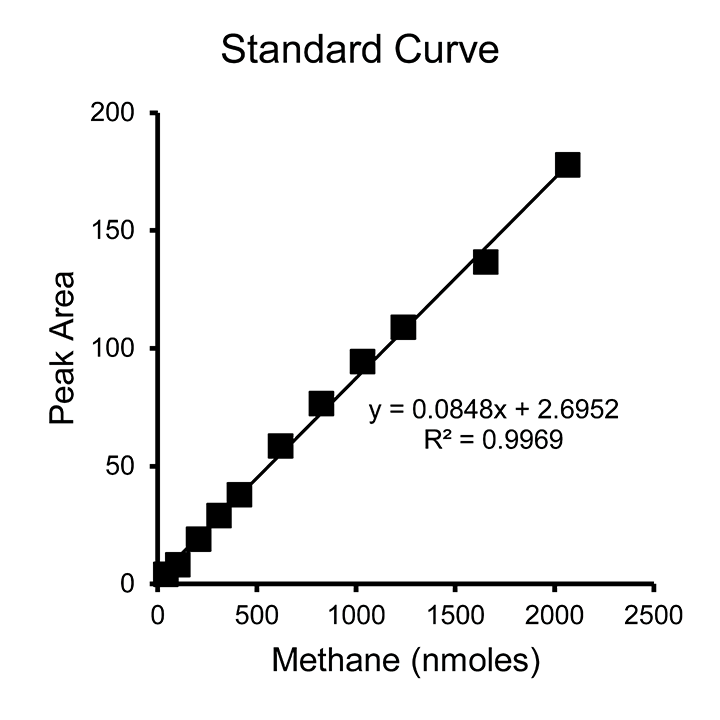
Figure 4. Example standard curve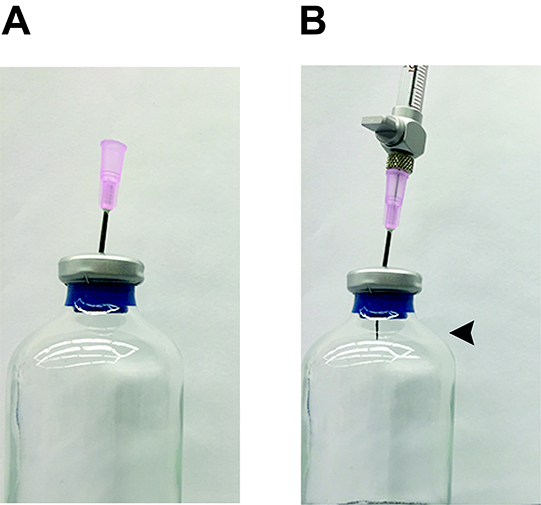
Figure 5. Method for preparing gas standards and sampling gas headspace. A. An 18 G needle is pushed into the stopper about three-quarters of the way through. This acts as a guide for the Hamilton needle to push through the stopper and not bend. B. A Hamilton syringe is inserted into an 18 G needle and pushed through the rest of the stopper. Once the end of the Hamilton syringe needle is through the end of the stopper (arrow), headspace can be extracted. Do not push the Hamilton syringe so far into the stopper that the 18 G needle is pushed through the stopper, as this will allow headspace gas to quickly escape.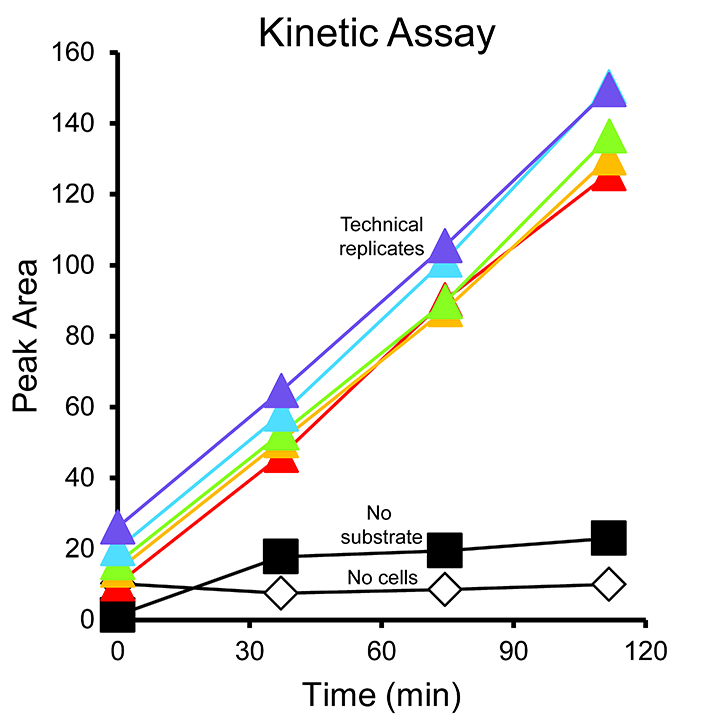
Figure 6. Example kinetic assay results
Recipes
Notes:
- Preparing anaerobic solutions is a technical skill. For general instructions, please refer to Wolfe and Metcalf (2010).
- Some Methanosarcina species of methanogens can be grown in medium of different osmolarity (low-salt, LS, or high-salt, HS) culture medium (Sowers et al., 1993). If growing other methanogens or methanotrophs, use the appropriate medium recipe for the organism(s) of interest.
- 200x mupirocin stock
- Under anaerobic conditions, stopper and crimp an empty Balch tube. Autoclave to sterilize.
- Dissolve 35 mg mupirocin in 0.5 ml 1 M NaOH in a 15 ml Falcon tube.
- Complete volume to 10 ml with ddH2O. The final concentration of mupirocin in the 200x stock solution is 3.5 mg/ml (14 mM).
- Using sterile technique, fit a 22 G needle to a sterile 17 mm diameter 0.2 µm pore size PVDF syringe filter using a plastic 20 ml syringe. Slowly push the solution through the filter, through the needle, and into the sterile anaerobic Balch tube.
- Use vacuum-vortex technique to make the mupirocin stock solution anaerobic (Wolfe and Metcalf, 2010).
- Store at 4 °C for up to a month.
- Under anaerobic conditions, stopper and crimp an empty Balch tube. Autoclave to sterilize.
- Plain medium (no C source)
Follow the same culture medium recipe as usual, but omit the carbon source. For methanogens that cannot produce methane from CO2 such as Methanosarcina acetivorans, the normal HS medium is prepared. For methanogens and autotrophs that can fix CO2, carbonate, bicarbonate, and CO2 gas should also be eliminated from the recipe and replaced with a buffer at the appropriate pH. For example, for Methanosarcina species that can produce methane from CO2, the bicarbonate in the normal HS medium recipe is replaced with 50 mM 3-N-(morpholino) propanesulfonate (MOPS) (pH 6.8), and the medium is sparged and dispensed into Balch tubes under 100% nitrogen. - 2x C medium
Add twice the concentration of carbon source to Plain medium (no C source). For example, when assaying Methanosarcina grown on HS medium, if the desired final concentration in the assay is 50 mM methanol, add 100 mM methanol to Plain HS medium to make 2x C HS medium.
Acknowledgments
This material is based upon work supported by the National Science Foundation under Grants IOS-1449525 and MCB-1449014, by the Water Environment Research Foundation grant NTRY6R14, and by the Nebraska Center for Energy Sciences Cycle 8 award to N. Buan. M. Smith was supported by an American Society for Microbiology Undergraduate Research Fellowship and a Pepsi UCARE Fellowship. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the funding sources. The authors declare no competing interests.
References
- Berndt, M. E., Allen, D. E. and Seyfried, W. E. (1996). Reduction of CO2 during serpentinization of olivine at 300 °C and 500 bar. Geology 24: 351-354.
- Catlett, J. L., Ortiz, A. M. and Buan, N. R. (2015). Rerouting cellular electron Flux to increase the rate of biological methane production. Appl Environ Microbiol 81(19): 6528-6537.
- Dry, M. E. (2002). The fischer-tropsch process: 1950-2000. Catal Today 71, 227-241.
- Horita, J. and Berndt, M. E. (1999). Abiogenic methane formation and isotopic fractionation under hydrothermal conditions. Science 285(5430): 1055-1057.
- Metcalf, W. W., Griffin, B. M., Cicchillo, R. M., Gao, J., Janga, S. C., Cooke, H. A., Circello, B. T., Evans, B. S., Martens-Habbena, W., Stahl, D. A. and van der Donk, W. A. (2012). Synthesis of methylphosphonic acid by marine microbes: a source for methane in the aerobic ocean. Science 337(6098): 1104-1107.
- Sowers, K. R., Boone, J. E. and Gunsalus, R. P. (1993). Disaggregation of methanosarcina spp. and growth as single cells at elevated osmolarity. Appl Environ Microbiol 59(11): 3832-3839.
- Thauer, R. K. (1998). Biochemistry of methanogenesis: a tribute to Marjory Stephenson. 1998 Marjory Stephenson Prize Lecture. Microbiology 144 (Pt 9): 2377-2406.
- Wolfe, R. S. (1982). Biochemistry of methanogenesis. Experientia 38 198-201.
- Wolfe, R. S. and Metcalf, W. W. (2010). A vacuum-vortex technique for preparation of anoxic solutions or liquid culture media in small volumes for cultivating methanogens or other strict anaerobes. Anaerobe 16(3): 216-219.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Aldridge, J. T., Catlett, J. L., Smith, M. L. and Buan, N. R. (2016). Methods for Detecting Microbial Methane Production and Consumption by Gas Chromatography. Bio-protocol 6(7): e1779. DOI: 10.21769/BioProtoc.1779.
Category
Microbiology > Microbial metabolism > Other compound
Biochemistry > Other compound > Alkane
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link














