- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
In vitro Fluorescent Matrix Degradation Assay for Entamoeba histolytica
Published: Vol 6, Iss 6, Mar 20, 2016 DOI: 10.21769/BioProtoc.1769 Views: 8979
Reviewed by: Fanglian HeAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Preparation of Mosquito Salivary Gland Extract and Intradermal Inoculation of Mice
Michael A. Schmid [...] Laura D. Kramer
Jul 20, 2017 19254 Views

Rearing of Culex spp. and Aedes spp. Mosquitoes
Elizabeth Kauffman [...] Laura D. Kramer
Sep 5, 2017 16812 Views
An Optimized P. berghei Liver Stage–HepG2 Infection Model for Simultaneous Quantitative Bioimaging of Host and Parasite Nascent Proteomes
James L. McLellan [...] Kirsten K. Hanson
Mar 5, 2024 1921 Views
Abstract
Fluorescent matrix degradation assay is a popular and widely used assay in the field of invadopodium biology (Artym et al., 2009). Matrix remodeling and degradation can be observed under both physiological and pathological conditions. Cancer cells extensively remodel and degrade the underlying matrix by employing actin-rich protrusive structures called invadosomes. Similar structures are formed by the protozoan parasite Entamoeba histolytica (E. histolytica), upon coming in contact with fibronectin, a major component of the host (extracellular matrix) ECM. Here, we describe a similar assay to measure matrix degradation by Entamoeba histolytica.
Materials and Reagents
- Sterile disposable pipettes (10 ml and 5 ml)
- Aluminum foil
- Four well tissue culture plates (Thermo Fisher Scientific, NuncTM, catalog number: 176740 )
- Round Glass coverslips (12 mm) (Bellco Glass, catalog number: 1943-10012A )
- Parafilm
- Entamoeba histolytica HM1:IMSS strain (a kind gift from Prof. Alok Bhattacharya, School of Life Sciences, Jawaharlal Nehru University, New Delhi, India)
- Fluorescently labeled fibronectin (CYTOSKELETON, catalog number: FNR02 )
- Glutaraldehyde (25% stock solution in dH2O, EM grade) (Sigma-Aldrich, catalog number: G5882 )
- Sodium Borohydride (NaBH4) (Sigma-Aldrich, catalog number: 452882 )
- Poly-L-lysine (Sigma-Aldrich, catalog number: P8920 )
- 70% ethanol (Merck Millipore Corporation)
- Alexa 568 Phalloidin (Thermo Fisher Scientific, Molecular ProbesTM, catalog number: A12380 )
- 1% BSA (Sigma-Aldrich, catalog number: A-7906 ) in PBS
- Mowiol mounting medium (Sigma-Aldrich, catalog number: 81381 )
- Complete BI medium (incomplete supplemented with 15%, Adult Bovine Serum)
- GasPakTM EZ Anaerobe Pouch System (BD Biosciences, catalog number: 260683 )
- Sterile dH2O
- Sodium chloride (NaCl) (Sigma-Aldrich, catalog number: S7653 )
- KCl
- Potassium phosphate monobasic (Na2HPO4) (Sigma-Aldrich, catalog number: P5655 )
- Potassium phosphate dibasic trihydrate (KH2PO4) (Sigma-Aldrich, catalog number: P5504 )
- NaOH
- Hilyte 488 Fibronectin
- Biosate peptone BBL (BD Biosciences, catalog number: 211862 )
- Glucose (Dextrose) (Sigma-Aldrich, catalog number: D9434 )
- L-cysteine (Sigma-Aldrich, catalog number: C1276 )
- L-ascorbic acid (Sigma-Aldrich, catalog number: A5960 )
- Ferric ammonium citrate (Sigma-Aldrich, catalog number: F5879 )
- 10x phosphate-buffered saline (PBS) (see Recipes)
- 0.01% poly-L-lysine solution (see Recipes)
- 0.5% glutaraldehyde solution (see Recipes)
- 5 mg/ml sodium borohydride (see Recipes)
- 4% paraformaldehyde (PFA) (see Recipes)
- 1% BSA in PBS (see Recipes)
- 0.1% Triton-X-100 (see Recipes)
- 100 μg/ml Hilyte488 Fibronectin (FNR02) (see Recipes)
- Basal incomplete medium (see Recipes)
- Complete medium (see Recipes)
Equipment
- Laminar air flow hood
- 37 °C incubator
- Forceps
- Moist chamber
- Vacuum Pump and glass Pasteur pipettes with tubings
- Table top centrifuge with swinging bucket rotor
Procedure
Coating of glass coverslips with fluorescent matrix
- Clean the glass coverslips with acetone, air dry and sterilize them by autoclaving before use.
- Further, handle the coverslips under sterile conditions inside the laminar air flow hood. Transfer the coverslips to a four well tissue culture plate using ethanol wiped forceps.
- Coat each coverslip with 0.01% poly-L-lysine using 80 μl of poly-L-lysine solution per coverslip for 20 min at room temperature. Air dry the solution under the hood.
- Wash the coverslips by adding 500 μl of sterile 1x PBS gently to the side of the well and incubating for 5 min each. The poly-L-lysine coated side should face up during the washes. After each wash aspirate it gently using a sterile glass pasteur pipette attached to a vacuum pump and repeat the washes for three times.
- After the third PBS wash, add 500 μl of ice cold 0.5% glutaraldehyde to each well such that the coverslips are immersed and incubate under dark at room temperature for 15 min.
- Aspirate the glutaraldehyde solution gently by tilting the plate to an angle such that coverslips are not disturbed and again wash with sterile 1x PBS. Repeat the washes for three times.
- Make a humid chamber using tissue paper and 70% ethanol. Neatly make a stack of tissue paper by folding it over and moisten it by spraying with 70% ethanol. Place the stack in a sterile Petri dish to make a humid chamber. Place a neatly cut parafilm over the moist tissue paper stack and press it gently so that it sticks well.
- Now, place appropriate number of drops of 50 μl of prewarm (37 °C, preferably in a water bath) fluorescently labeled fibronectin (100 μg/ml) over the parafilm according to the number of coverslips being coated.
- Carefully lift the coverslips from the wells holding at the edges using a pair of forceps and place them with their poly-L-lysine coated side facing downwards on the drop of the fluorescent fibronectin on the parafilm. While placing the coverslips, ensure that there is no air bubble trapped beneath them. If so, gently press the coverslips with the forceps to eliminate the trapped air bubbles.
- Incubate under safe light in a 37 °C incubator for an hour.
- After an hour, remove the coverslips from the incubator. Now coverslips should be handled in safe light. Lift the coverslips carefully one-by-one with the help of the forceps without scratching the matrix and place them with their coated side facing up in a fresh tissue culture plate.
- Again wash them gently with 1x PBS as done previously in step 4. Meanwhile, prepare fresh 5 mg/ml solution of sodium borohydride by dissolving it in 1x PBS. Add 500 μl of sodium borohydride per well such that the coverslips are immersed in it. Incubate under dark at room temperature for 15 min. As sodium borohydride bubbles upon dissolution, make sure that the bubbles do not settle on the coverslips by gently rocking them back and forth.
- Wash the coverslips with 1x PBS for three times and then finally sterilize by soaking them into 70% ethanol for 10 min at room temperature. Further wash the coverslips with sterile 1x PBS and then finally add 500 μl of complete BI medium to each well and incubate at 37 °C in an incubator for an hour.
- Coverslips are now ready to be used. If you do not want to use them immediately, sterilize them using 70% ethanol and then store them at 4 °C under dark for up to a week.
Sample preparation for the assay
- Take a confluent tube of Entamoeba histolytica HM1: IMSS strain in logarithmic phase and remove the used medium. Add 10 ml of fresh complete BI medium and transfer the glass tube on ice for 5 min so as to detach the cells. The culture volume in the glass tube is 13 ml.
- Now gently tap the tube to completely detach the trophozoites and transfer in a 15 ml conical tube. Take a small aliquot to count the number of cells using haemocytometer. Spin them down at 500 x g for 5 min at room temperature. Now resuspend the cell pellet in warm complete BI medium to a concentration of 40 x 104 cells/ml.
- Add 0.5 ml of cells i.e. 2 x 105 cells per well over the fluorescently matrix coated coverslips and place the tissue culture plate inside a GasPak EZ pouch to maintain anaerobic conditions. Place the whole setup in 37 °C incubator.
- Incubate for 36-48 h at 37 °C under dark.
- Close to the end of incubation, warm an aliquot of 1x PBS and 4% PFA at 37 °C for further use.
Preparing samples for fluorescence microscopy
- Aspirate the medium by tilting the plate to an angle without disturbing the coverslips. Now wash the cells with warm sterile 1x PBS and fix the cells immediately using 4% PFA. Incubate at room temperature for 15 min under dark.
- After fixation, wash the sample with 1x PBS and permeabilize with 0.1% Triton-X-100 at room temperature for 10 min. Wash again and block the samples with 1% BSA in PBS for an hour at room temperature.
- To label with Alexa 568 Phalloidin, dilute phalloidin in a ratio of 1:100 in the blocking solution (1% BSA in PBS).
- Transfer the coverslips carefully to a humid chamber again and add 50 μl of Alexa 568 Phalloidin to each coverslip and incubate under dark at room temperature for an hour.
- Subsequently, wash the samples by again placing them back in the tissue culture plate and adding 1x PBS.
- After the final wash, mount the sample using mounting medium. Let the samples air dry at room temperature under dark.
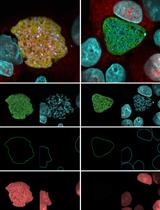
- Examine the fluorescence of the matrix and the samples using epifluorescence or confocal microscope with appropriate light source and filter sets. Since the matrix layer formed is very thin the samples should be focused accurately so that the entire field of view has uniform fluorescence. The degradation of the matrix can be observed as loss of fluorescence around the cells and would appear as a dark halo surrounding the cells.

Figure 1. Rab21 mediated fibronection degradation by amoebic trophozoites. A. Trophozoites overexpressing Rab21CA mutant were plated on a Hilyte 488 fibronectin coated glass coverslip and incubated for 48 h at 37 °C under dark. Cells were fixed, permeabilized and stained with Alexa 568 Phalloidin and DAPI. B. Undamaged Hilyte 488 Fibronectin. Scale bar, 10 µm.
Notes
- Always prepare fresh 0.5% glutaraldehyde solution. As this solution is hazardous, avoid contact with skin and eyes.
- Sodium borohydride should also be prepared fresh just before use. Upon dissolution, sodium borohydride releases free H2 gas, so it should always be prepared in conical tubes with enough free space to avoid spills and bumping of the solution. Do not tightly cap the tubes containing the solution.
- Passage your cells regularly and do not allow cells to grow to 100% confluency. Use cells which are 70-80% confluent for the experiment.
Recipes
- 10x phosphate-buffered saline (PBS)
Dissolve 80 g NaCl with 2 g KCl, 14.4 g Na2HPO4, 2.4 g KH2PO4
Adjust pH to 7.4 with NaOH
Add dH2O to 1,000 ml
Filter sterilize (0.2 μm)
Stored at room temperature - 0.01% poly-L-lysine solution
Dilute 0.1% solution of poly-L-lysine in sterile dH2O in a 1:10 ratio - 0.5% glutaraldehyde solution
Add 0.2 ml of stock solution to 10 ml of sterile 1x PBS - 5 mg/ml sodium borohydride
Weigh 100 mg of sodium borohydride
Dissolve in 20 ml of 1x PBS just before use - 4% PFA
Weigh 4 g of PFA
Dissolve in 100 ml of pre-warmed 1x PBS with constant stirring - 1% BSA in PBS
Weigh 0.5 g of BSA and dissolve in 50 ml of 1x PBS - 0.1% Triton-X-100
Make a stock solution of 10% Triton-X-100 in 1x PBS
Add 0.5 ml of 10% Triton-X-100 in 50 ml of 1x PBS - 100 μg/ml Hilyte488 Fibronectin (FNR02)
Hilyte 488 Fibronectin comes as a desiccated powder and is stored at 4 °C under dark Dissolve the powder at a final concentration of 1 mg/ml using sterile dH2O
Make aliquots and stored at -20 °C - Basal incomplete medium
Dissolve:
3 g of Biosate peptone BBL
1 g of Glucose
200 mg of NaCl
60 mg of KH2PO4
100 mg of K2HPO4
100 mg of L-cysteine
20 mg of L-ascorbic acid and 2.28 mg of Ferric ammonium citrate in dH2O
Adjust the pH to 6.8 using 8 N NaOH
Add dH2O to 88 ml
Sterilized by autoclaving at 121 °C at 15 psi for 15 min - Complete medium
Add 15 ml of adult bovine serum (filter sterilized) and 2 ml of 100x vitamin mixture (filter sterilized) to 88 ml of basal incomplete medium
Acknowledgments
We would like to thank Prof. Alok Bhattacharya (Jawaharlal Nehru University, New Delhi, India) and Dr. Sandipan Ganguly (National Institute of Cholera and Enteric Diseases, Kolkatta, India) for providing us with the Entamoeba histolytica HM1: IMSS strain. We are grateful to Professor Stefan Linder (University of Hamburg, Hamburg, Germany) for helpful discussion and suggestions. We are also thankful to Drs. Vira V. Artym, Kenneth M. Yamada and Susette C. Mueller as the protocol for E. histolytica has been adapted from their published protocol for studying cancer cell invasion in Methods in Molecular Biology. We sincerely thank Ms. Lekha Vinod Shah, final year Dual BS-MS degree program student at IISER Bhopal, Department of Biological Sciences, who shot and edited the video for the protocol.We are thankful to the Central Instrumentation Facility at IISER Bhopal, India. This work was funded by intramural funds from IISERB and funds from Max Planck Gesellschaft (MPG), Germany and Department of Science and Technology (DST), India. ME was funded by Senior Research Fellowship provided by University Grant Commission, Government of India.
References
- Artym, V. V., Yamada, K. M. and Mueller, S. C. (2009). ECM degradation assays for analyzing local cell invasion. Methods Mol Biol 522: 211-219.
- Emmanuel, M., Nakano, Y. S., Nozaki, T. and Datta, S. (2015). Small GTPase Rab21 mediates fibronectin induced actin reorganization in Entamoeba histolytica: implications in pathogen invasion. PLoS Pathog 11(3): e1004666.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Emmanuel, M. and Datta, S. (2016). In vitro Fluorescent Matrix Degradation Assay for Entamoeba histolytica. Bio-protocol 6(6): e1769. DOI: 10.21769/BioProtoc.1769.
Category
Microbiology > Microbe-host interactions > In vivo model > Protozoan
Cell Biology > Cell imaging > Fluorescence
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link