- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Sample Preparation for X-ray Micro-computed Tomography of Woody Plant Material and Associated Xylem Visualisation Techniques
Published: Vol 6, Iss 6, Mar 20, 2016 DOI: 10.21769/BioProtoc.1767 Views: 10706
Reviewed by: Samik BhattacharyaPengpeng LiArsalan Daudi

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
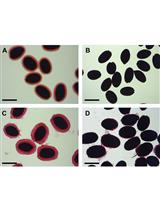
Analysis of Monosaccharides from Arabidopsis Seed Mucilage and Whole Seeds Using HPAEC-PAD
Gillian H. Dean [...] George W. Haughn
Dec 20, 2019 5777 Views
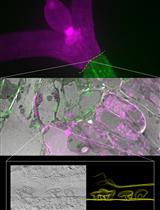
Targeting Ultrastructural Events at the Graft Interface of Arabidopsis thaliana by A Correlative Light Electron Microscopy Approach
Clément Chambaud [...] Lysiane Brocard
Jan 20, 2023 3012 Views

Closed Systems to Study Plant–Filamentous Fungi Associations: Emphasis on Microscopic Analyses
Vasiliki Skiada and Kalliope K. Papadopoulou
Feb 20, 2025 2901 Views
Abstract
Variation in the tissue structure of short rotation coppice (SRC) willow is a principle factor driving differences in lignocellulosic sugar yield yet much of the physiology and development of this tissue is unknown. Traditional sectioning can be both difficult and destructive in woody tissue; however, technology such as three dimensional X-ray micro-computational tomography (μCT) scanning can be used to move biological researchers beyond traditional two dimensional assessment of tissue variation without having to destructively cut cells. This technology does not replace classical microscopic techniques but rather can be carefully integrated with traditional methods to improve exploration of the world of plant biology in three dimensions. The procedures below outline preparation of willow for 3D X-ray μCT and associated xylem staining and visualisation techniques, in particular secondary xylem programmed-cell-death (PCD) delay during gelatinous fibre (g-fibre) development. Many of the staining techniques here are transferable to other woody species such as poplar and Eucalyptus.
Keywords: Micro-computed tomographyMaterials and Reagents
- Glass slides
- Cover slips
- 48 well plates
- Razor blades
- Short rotation coppice (SRC) willow trees [cultivar Resolution-pedigree: {S. viminalis. x (S. viminalis. x S. schwerinii SW930812)] x [S. viminalis. x (S. viminalis. x S. schwerinii ‘Quest’)]}
- 37% Formaldehyde (36.5-38% in H2O) (Sigma-Aldrich, catalog number: F8775 )
- Acetic acid (≥ 99.7%) (Sigma-Aldrich, catalog number: 320099 )
- Ethanol
- FAA: 3.7% formaldehyde, 5% acetic acid and 47.5% ethanol
- Oasis® floral foam (OASIS Floral Products) (http://oasisfloralproducts.com/)
- Safranin-O solution (Sigma-Aldrich, catalog number: HT90432 )
- DPX Mountant for histology (Sigma-Aldrich, catalog number: 06522 )
Note: This product is named “Slide mounting medium (p-Xylene-bis(N-pyridinium bromide))” at Sigma-Aldrich. - Distilled water
- β-D-glucosyl Yariv reagent (Biosupplies Australia Pty Ltd., catalog number: 100-2 )
- Histoclear clearing agent (National Diagnostics, catalog number: HS-200 )
- Chlorazol black E/direct black 38 (Sigma-Aldrich, catalog number: C1144 )
Equipment
- Reichert Sliding Microtome
- Tweezers (for handling section)
- Light microscope
Procedure
X-ray μCT can also potentially be used for a broader range of crops, and a more diverse set of biological questions. In most cases, care should be taken to air-dry material prior to scanning; water saturation will reduce cell wall contrast while forced drying beyond fibre saturation point runs the risk of inducing cell collapse, altering the plant’s tissue architecture. An example of this wider potential for the technology in plant research is provided in Figure 1 and Video 1.
Figure 1. Miscanthus internode 3D render. X-ray 3D Computational tomography example images of mature Miscantus x giganteus stem.
- Stem sampling
The research where 3D X-ray μCT scanning was used to assess xylem fibre PCD (Brereton et al., 2015) used short rotation coppice (SRC) willow trees (cultivar Resolution-pedigree: [S. viminalis. x (S. viminalis. x S. schwerinii SW930812)] x [S. viminalis. x (S. viminalis. x S. schwerinii ‘Quest’)] grown in pots for 12 weeks. At week six the trees were tipped at a 45° angle to the horizontal and held in place for a further 6 weeks before harvesting (with the stem tied to a straight bamboo stem every two days to maintain the 45° angle). This alteration to the vector of gravitational stimulus elicits a unique change in tissue development known as the reaction wood response (Andersson-Gunneras et al., 2003; Brereton et al., 2011; Pitre et al., 2007; Wardrop and Dadswell, 1955).- Harvest a sample whole stem segment of 6-8 cm from the midpoint of the stem.
Note: This is deliberately chosen in preference to the traditional set breast height of 1.3 meters in order to allow developmentally comparable analysis between genotypes of different morphologies. - Cut two small triangular notches into the wood 3-4 cm along the segment using a razor blade (making sure to reach through the bark to the secondary xylem), both notches should be at asymmetric positions from a transverse perspective (see Figure 2).

Figure 2. Chlorazol black and Safranin O staining. Classical sectioning and staining with light microscopy can be used to help navigate and orientate tissue architecture of the stem (Panel A). In this context, chlorazol black E (counter-stained with safranin O) is used to stain the g-layer of g-fibres in tension wood of a tipped stem sample of SRC willow (cultivar resolution)-this can then be aligned with images from the μCT scan (Panel B). Note, there is variation throughout the entirety of the 3D stem (longitudinal variation in tissue as well as transverse) so multiple notches in the stem biomass being scanned (one is visible here as a single triangle cut in panel A) are an important tool to help navigate the 3D volume data from the scan alongside natural asymmetry of the tissue. Scale bar = 4 mm. - Cut the segment in half to create two 3-4 cm segments. One 3-4 cm stem segment (stem A) will be used for X-ray μCT scanning while the other (stem B) will be used for associated staining and light or confocal microscopy. The notches will assist in navigation of the segments (allowing overlay and alignment) and can be removed in silico for 3D µCT render (stem A) or during sectioning (stem B).
- To make sectioning, staining and X-ray μCT scan differentiation simpler, bark can be removed at this point from both stem segments by manual peeling or with a scalpel.
- After harvesting stem segments should be “fixed” in FAA-formaldehyde, acetic acid, alcohol solution (3.7% formaldehyde, 5% acetic acid and 47.5% ethanol).
- Harvest a sample whole stem segment of 6-8 cm from the midpoint of the stem.
- Drying and pre-μCT scan prep
- After fixation of Stem A for > 3 days, the segment should be removed from FAA solution and briefly washed in H2O 4 days prior to 3D X-ray µCT scanning.
- Air-drying of stem A should be performed at room temperature (no greater than 30 °C) for over 4-5 days in a clean environment with relatively good air turnover. Forced drying at higher temperatures can pull H2O from the cell wall (beyond fibre saturation point) resulting in cell wall collapse. In very dry climates (low humidity) this can also occur during air-drying.
Note: Air-drying of stem samples seems an essential step in creating contrast between the three elements within the biological system under scrutiny: the cell wall, the protoplast and void space. The important factor of air-dried samples being that “full” cells, tubular in shape, are easily distinguished from intracellular void space (which was not the case when “wet”, as intracellular void space is full of water (Figure 3).
Figure 3. Wet and air-dry biomass 2D computational tomography scan examples. 3D X-Ray µCT models are rendered from >1,000 2D images, such as those from a tipped stem sample of SRC willow (cultivar resolution) shown above. A represents a 2D slice of a sample scanned “wet” whereas B represents a 2D slice of the same sample after having been air-dried for 3-4 days. Greater contrast between cell walls, cells with a protoplast (having not completed programmed cell death) and void volume is visible in the air-dried sample. - Once stem A is air-dried the sample can be secured in a low-density media, usually standard Oasis® floral foam (a phenolic based polymer, care should be taken if specifically studying cellulose, as such low densities are hard to differentiate) and positioned for scan with the structure/s of interest in the centre of the scan volume.
- After fixation of Stem A for > 3 days, the segment should be removed from FAA solution and briefly washed in H2O 4 days prior to 3D X-ray µCT scanning.
- Infiltration
- As X-ray absorption increases with atomic number of atoms within a voxel, stem samples can potentially be vacuum infiltrated with heavy metals before 3D X-ray µCT scanning. Such as with phosphotungstate (Staedler et al., 2013).
- As X-ray absorption increases with atomic number of atoms within a voxel, stem samples can potentially be vacuum infiltrated with heavy metals before 3D X-ray µCT scanning. Such as with phosphotungstate (Staedler et al., 2013).
- Post-μCT scan
- Once stem A has been scanned (Brereton et al., 2015) a destructive density measurement can be taken. Basic density (Biermann, 1996; Jourez et al., 2001) is best suited for biomass density quantification as it incorporates tissue structural variation often meaningful in terms of physiology.
- Vacuum infiltrate stem A with water before green volume is measured via water displacement (Biermann, 1996). Vacuum infiltration can be performed by pulling a vacuum over submerged wood samples for around 24 h depending on the material used. Wood oven dry weight is then measured after drying overnight at 105 °C.
- Calculated as (Biermann, 1996; Jourez et al., 2001): Basic density = oven dry mass/volume.
- Once stem A has been scanned (Brereton et al., 2015) a destructive density measurement can be taken. Basic density (Biermann, 1996; Jourez et al., 2001) is best suited for biomass density quantification as it incorporates tissue structural variation often meaningful in terms of physiology.
- Sectioning and staining
Staining of the transverse sections from Stem B allows for different tissue types to be visualised and quantified within the stem. By clearly following each stem segment and navigating natural asymmetry (as well as notches cut into the stems), the in silico 3D X-ray µCT render, g-layers of g-fibres [chlorazol black E (Brereton et al., 2015; Brereton et al., 2011; Brereton et al., 2012; Robards and Purvis, 1964)], β-galactosyl Yariv reagent (Tryfona et al., 2012; Kitazawa et al., 2013), secondary cell wall (safranin O) and remnants of cell contents (Coomassie and β-galactosyl Yariv) can be aligned and compared in situ. The in situ β-galactosyl Yariv reagent and Coomassie staining seem to be without published precedence but are simple and effective. Coomassie staining is particularly effective for macro navigation (across a whole transverse section) of cell content remnants using light microscopy, as classical viability staining can be ineffective in woody tissue (such as: Tunnel and NBT) due to cell disruption during the sectioning process. Staining duration and concentrations are only suggestions here as cell wall structure will vary depending on the plant studied and the growth conditions.- Once Stem B is secured in the Reichert sledge microtome, adjust angle of blade to the correct plane (this may require some trial and error depending on the material), dampen the sample with water and cut the thickness of < 30 µm.
- A 48 well plate is useful to perform multiple staining pipelines simultaneously on a common set of sections.
- Using tweezers throughout, first place a section in H2O to hydrate for 2 min before using section for either chlorazol black E staining, β-galactosyl Yarive reagent in situ staining or Coomassie in situ staining (cell wall antibodies can also be integrated with analysis here).
Chlorazol black E staining (example Figure 2)- Counter-stain by placing sections in a well containing 1% safranin O (aq) for 2-3 min.
- Wash in wells containing H2O x2 for 30 sec.
- Place section in progressive dehydration series: 50% ethanol for 30 sec, 75% ethanol for 30 sec, 100% ethanol x2 for 1 min.
- Place section in well containing 1% chlorazol black E (in methoxyethanol) for 4 min to stain g-layers.
- Wash in well containing 100% ethanol x2 for 1 min.
β-galactosyl Yariv reagent staining (example Figure 4)- Place section in 0.1 mg/ml dilute β-galactosyl Yariv reagent in 0.15 M NaCl and incubate at room temperature for 4 h on gently shaking platform.
- Wash as necessary in wells containing H2O.
- Move onto step E9.

Figure 4. In situ β-galactosyl Yariv reagent staining. β-galactosyl Yariv reagent has long been used to condense arabinogalactan proteins (AGPs). As the g-layer of g-fibres in tension wood are rich in Fasciclin-like arabinogalactan protein (Lafarguette et al., 2004; Andersson-Gunneras et al., 2006), β-galactosyl Yariv reagent works well as an in situ stain for g-layers in tension wood of SRC willow stem as well as a protoplast stain (live cells will also include AGPs). Light microscopy of a tipped stem sample of SRC willow (cultivar resolution) shows A patterning of tensions wood g-layers and delayed PCD well aligned those of Figures 2. The same sample magnified using light microscopy to x63 illustrates: B. Tension wood (TW), C Opposite wood (OW) and D TW/OW interface show how clear the in situ β-galactosyl Yariv reagent staining can differentiate these cellular components. Scale bar = 4 mm.
Coomassie staining (example Figure 5)- Place section in 1% Coomassie solution and incubate at room temperature for 4 h on gently shaking platform.
- Wash as necessary in wells containing H2O.
- Move onto step E9.
Slide construction- Rinse staining section in Histoclear clearing agent for 1 min. At this stage apply DPX to slide to slightly set (1 min or so).
- Place section on to the DPX and apply cover slip, lowering gently. Apply pressure to force out air bubbles, place some weight (a few grams) on top and leave to set for 24 h.

Figure 5. In situ Coomassie staining. A. Stitched 25 μm transverse sections stained with 1% Coomassie solution and counter stained with safranin O. B. Stitched 25 μm transverse sections without counterstain, remnants of cell contents is visible if present. While z-stacked images are difficult to assemble clearly using Coomassie staining, the process was invaluable to assess remnants of cell content using light microscopy (by eye, as one can move slightly through 3D space using light microscopes) and led to the identification of the same PCD patterning using 3D X-ray μCT. Scale bar = 4 mm.
- Once Stem B is secured in the Reichert sledge microtome, adjust angle of blade to the correct plane (this may require some trial and error depending on the material), dampen the sample with water and cut the thickness of < 30 µm.
Acknowledgments
The author was financially supported by BioFuelNet Canada. X-ray μCT scanning was performed in collaboration with Farah Ahmed and Dan Sykes at the Natural History Museum with funding from BBSRC Sustainable Bioenergy Centre (BSBEC), working within the BSBEC BioMASS (http://www.bsbec-biomass.org.uk/) Programme. In situ staining techniques were developed in collaboration with Dr. Michael Jasmine Ray.
References
- Andersson-Gunneras, S., Hellgren, J. M., Bjorklund, S., Regan, S., Moritz, T. and Sundberg, B. (2003). Asymmetric expression of a poplar ACC oxidase controls ethylene production during gravitational induction of tension wood. Plant J 34(3): 339-349.
- Andersson-Gunneras, S., Mellerowicz, E. J., Love, J., Segerman, B., Ohmiya, Y., Coutinho, P. M., Nilsson, P., Henrissat, B., Moritz, T. and Sundberg, B. (2006). Biosynthesis of cellulose-enriched tension wood in Populus: global analysis of transcripts and metabolites identifies biochemical and developmental regulators in secondary wall biosynthesis. Plant J 45(2): 144-165.
- Biermann, C. J. (1996). Handbook of pulping and papermaking, 2nd edn. Academic Press.
- Brereton, N. J., Ahmed, F., Sykes, D., Ray, M. J., Shield, I., Karp, A. and Murphy, R. J. (2015). X-ray micro-computed tomography in willow reveals tissue patterning of reaction wood and delay in programmed cell death. BMC Plant Biol 15: 83.
- Brereton, N. J., Pitre, F. E., Ray, M. J., Karp, A. and Murphy, R. J. (2011). Investigation of tension wood formation and 2,6-dichlorbenzonitrile application in short rotation coppice willow composition and enzymatic saccharification. Biotechnol Biofuels 4: 13.
- Brereton, N. J., Ray, M. J., Shield, I., Martin, P., Karp, A. and Murphy, R. J. (2012). Reaction wood - a key cause of variation in cell wall recalcitrance in willow. Biotechnol Biofuels 5(1): 83.
- Jourez, B., Riboux, A. and Leclercq, A. (2001). Comparison of basic density and longitudinal shrinkage in tension wood and opposite wood in young stems of Populus euramericana cv. Ghoy when subjected to a gravitational stimulus. Canadian Journal of Forest Research-Revue Canadienne De Recherche Forestiere 31, 1676-1683.
- Kitazawa, K., Tryfona, T., Yoshimi, Y., Hayashi, Y., Kawauchi, S., Antonov, L., Tanaka, H., Takahashi, T., Kaneko, S., Dupree, P., Tsumuraya, Y. and Kotake, T. (2013). beta-galactosyl Yariv reagent binds to the beta-1, 3-galactan of arabinogalactan proteins. Plant Physiol 161(3): 1117-1126.
- Lafarguette, F. et al. (2004). Poplar genes encoding fasciclin-like arabinogalactan proteins are highly expressed in tension wood. New Phytologist 164, 107-121.
- Pitre, F. E., Cooke, J. E. K. and Mackay, J. J. (2007). Short-term effects of nitrogen availability on wood formation and fibre properties in hybrid poplar. Trees-Structure and Function 21, 249-259.
- Robards, A. W. and Purvis, M. J. (1964). Chlorazol black E as a stain for tension wood. Biotechnic and Histochemistry 39, 309-315.
- Staedler, Y. M., Masson, D. and Schonenberger, J. (2013). Plant tissues in 3D via X-ray tomography: simple contrasting methods allow high resolution imaging. PLoS One 8(9): e75295.
- Timell, T. E. (1969). The chemical composition of tension wood. Comptes Rendus Biologies 72, 173-181.
- Tryfona, T., Liang, H. C., Kotake, T., Tsumuraya, Y., Stephens, E. and Dupree, P. (2012). Structural characterization of Arabidopsis leaf arabinogalactan polysaccharides. Plant Physiol 160(2): 653-666.
- Wardrop, A. and Dadswell, H. (1955). The nature of reaction wood. IV. Variations in cell wall organization of tension wood fibres. Australian Journal of Botany 3, 177-189.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Brereton, N. J. B. (2016). Sample Preparation for X-ray Micro-computed Tomography of Woody Plant Material and Associated Xylem Visualisation Techniques. Bio-protocol 6(6): e1767. DOI: 10.21769/BioProtoc.1767.
Category
Plant Science > Plant cell biology > Tissue analysis
Plant Science > Plant physiology > Tissue analysis
Cell Biology > Tissue analysis
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











