- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
In vivo Bioluminescence Imaging of Luciferase-labeled Cancer Cells
Published: Vol 6, Iss 6, Mar 20, 2016 DOI: 10.21769/BioProtoc.1762 Views: 36717
Reviewed by: Vivien Jane Coulson-ThomasAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
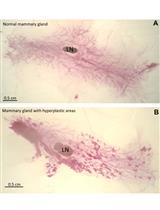
Mouse Mammary Gland Whole Mount Preparation and Analysis
Cornelia Tolg [...] Eva A. Turley
Jul 5, 2018 17017 Views
A Tumor-admixture Model to Interrogate Immune Cell–dependent Tumorigenesis
Jordan T. Noe [...] Robert A. Mitchell
Mar 5, 2023 2815 Views
An Optimized Protocol for Simultaneous Propagation of Patient-derived Organoids and Matching CAFs
Jenny M. Högström and Taru Muranen
Jan 20, 2025 4513 Views
Abstract
Over the past decade, in vivo bioluminescent imaging has emerged as a non-invasive and sensitive tool for studying ongoing biological processes within living organisms (Contag et al., 1997; Contag et al., 1998). Based on the detection and quantitation of the photons produced by the oxidation of luciferin by luciferase enzymes (Harvey, 1927), this technique has proved to be particularly useful in analyzing cancerous cells and monitoring tumor growth (Edinger et al., 1999; Sweeney et al., 1999; Vidal et al., 2015), providing a cost-effective insight into how the disease progresses in vivo, without the need of serial sacrifice of animals. This protocol describes in detail the procedure of obtaining luciferase-tagged tumors in immunocompromised mice that can be studied by bioluminescent imaging through the use of an IVIS Spectrum imager.
Keywords: Bioluminescent imagingMaterials and Reagents
- FalconTM Standard 10 cm2 Tissue Culture Dishes (Corning, catalog number: 353003 )
- Syringe Filter 0.45 μm strainer (Corning, catalog number: CLS431225 )
- 22Rv1 prostate cancer cells (ATCC, catalog number: CRL-2505 )
- NOD.Cg-Prkdcscid Il2rgtm1Wjl/SzJ (NSGTM) mice (Jackson Lab, catalog number: 00557 )
- Phoenix-Ampho cells (ATCC, catalog number: CRL-3213 )
- pLenti CMV Puro LUC (w168-1) (Addgene, catalog number: 17477 )
- Helper pCMV-VSV-G (Addgene, catalog number: 8454 ) and pMD2.G (Addgene, catalog number: 12259 )
- One Shot Stbl3 Competent E. coli (Thermo Fisher Scientific, InvitrogenTM, catalog number: C7373-03 )
- LB Broth Medium (Thermo Fisher Scientific, BioReagentsTM, catalog number: BP1426-2 )
- Ampicillin (Sigma-Aldrich, catalog number: A9393 )
- QIAprep Spin Miniprep Kit (QIAGEN, catalog number: 27106 )
- RPMI 1640 Medium (Thermo Fisher Scientific, GibcoTM, catalog number: 11875119 )
- Dulbecco’s Phosphate-Buffered Saline (DPBS) 1x (Corning, catalog number: 21-031 )
- jetPEI® DNA Transfection Agent (Polyplus-transfection, catalog number: 101-10N )
- Penicillin-Streptomycin antibiotic (Thermo Fisher Scientific, GibcoTM, catalog number: 15140122 )
- Polybrene® (Santa Cruz Biotechnology, catalog number: 134220 )
- Isoflurane (Baxter, catalog number: 1001936040 )
- Matrigel® Growth Factor Reduced (GFR) Basement Membrane Matrix (Corning, catalog number: 354230 )
- D-luciferin (PerkinElmer, catalog number: 770504 )
Equipment
- Thermo-block (Thermo Fisher Scientific, model: IsotempTM Digital and Analog Dry Bath Incubator )
- Shaker incubator (Eppendorf AG, New Brunswick Scientific, model: Innova 44R Incubator/Shaker )
- Table-top microcentrifuge (Eppendorf AG, model: centrifuge 5415R )
- Spectrophotometer (Thermo Fisher Scientific, model: SPECTRONICTM 200 Spectrophotometer )
- Incubator (Thermo Fisher Scientific, model: Heracell 150i )
- Vortex (Scientific Industries, model: Vortex Genie 2 )
- IVIS Spectrum imager (PerkinElmer, model: IVIS Spectrum Preclinical In Vivo Imaging System )
Software
- Caliper LifeScience Living Image® in vivo imaging software (PerkinElmer, catalog number: 128110) (Lumina/Kinetic/XR/100, Living Image V4.1)

Figure 1. Schematic representation of the steps required to successfully tag cells with luciferase and perform in vivo imaging
Procedure
- Bacterial transformation
Artificially constructed plasmids are the most commonly used vectors for introducing foreign DNA into bacterial cells. In this protocol, we used the commercial plasmid Lenti CMV Puro LUC, which contains the firefly luciferase gene, and was transformed into a culture of Stbl3 competent cells.- Transfer 50 μl of Stbl3 competent cells in a 1.5 ml Eppendorf tube placed on ice.
- Add between 100 ng and 500 ng of plasmid into the tube and resuspend with Stbl3 competent cells (pipet to resuspend, do not use vortex).
- Incubate the tube on ice for 30 min, gently resuspend every 10 min.
- Place the tube in a thermo-block for exactly 45 sec at 42 °C, and immediately put back into ice for 2 min.
- Add 800 μl of LB-Medium with no antibiotic into the tube, and gently resuspend the sample with a pipet (do not use vortex).
- Grow up bacteria in a shaker incubator at 37 °C speed 200 rpm for 2-3 h.
- Plate 100 µl of transformation mixture on LB-agar plates with 100 μg/ml of ampicillin.
- Place plate in an incubator at 37 °C overnight.
- Pick a single colony and inoculate a flask containing 50 ml of LB-Medium + 100 μg/ml of ampicillin.
- Incubate overnight at 37 °C in a shaker incubator at a speed of 200 rpm.
- Transfer 50 μl of Stbl3 competent cells in a 1.5 ml Eppendorf tube placed on ice.
- DNA extraction
There are several methods that can be used to purify plasmid DNA depending on the size of the bacterial culture and its corresponding plasmid yield, and a handful of kits available from varying manufacturers to do it. Here, we perform a minipreparation (miniprep) using the QIAprep Spin Miniprep Kit, which allows the isolation of a small-scale sample of up to 20 μg plasmid DNA from the transformed bacteria. Briefly, in this protocol the isolation of DNA from bacteria relies upon the use of columns and centrifugation steps in which DNA is sheared, extracted, and precipitated. Follow protocol of the manufacturer. - Transfection of DNA into viral packaging cells
Depending on the plasmid, several packaging cells can be used to generate the desired virus. In this protocol we used Phoenix-Ampho cells, which are a second-generation retrovirus producer cell line that is also capable of a long-term stable production of lentiviruses when helper DNA (CMV and DMV) is present. Helper DNA encodes viral genes that increase viral replication and packaging.- Seed 3 x 106 Phoenix-Ampho cells in 8 ml of RPMI supplemented with 10% FBS and 1% PenStrep in a 10 cm2 culture plate 24 h before transfection.
- In a first 1.5 ml Eppendorf tube, add 6 μg of DNA, 3 μg of each helper (CMV and DMV) and fill up to 250 μl of 150 mM NaCl (per sample to transfect).
- In a second 1.5 ml tube add 24 μl of jetPEI and 226 μl of 150 mM NaCl (per sample to transfect).
- Vortex the tubes for 10 sec.
- Add the Jetpei + NaCl into the Eppendorf tube with the DNA and vortex for 10 sec.
- Incubate mixture for 15 min at room temperature.
- Remove media of plate and add 9.5 ml of fresh RPMI supplemented with 10% FBS and 1% PenStrep to Phoenix-Ampho cells (see step C1).
- Add 500 μl drop by drop, of the mixture (step C5) to the plate and incubate overnight at 37 °C.
- After 24 h change the medium and add fresh RPMI supplemented with 10% FBS and 1% PenStrep.
- After 48 h, collect and filter the medium with a 0.45 μm strainer into a 50 ml Falcon tube.
Note: Viral concentration can be measured after this step by serial dilution to obtain quantitative information. Various techniques can be used to quantify virus concentration which include traditional methods (e.g. plaque assay) and commercial kits (e.g., ELISA or Q-PCR).
- Seed 3 x 106 Phoenix-Ampho cells in 8 ml of RPMI supplemented with 10% FBS and 1% PenStrep in a 10 cm2 culture plate 24 h before transfection.
- Viral particle cell infection
The viral particles produced in the previous step are used to infect cells. Virus will infect cells which hold a receptor that the virus can bind to. Therefore, the efficiency of viral infection will greatly depend on the cell type. In this protocol, we infected 70% confluent 22Rv1 prostate cancer cells growing in 10 cm2 culture dishes with 10 ml of media.- Add Polybrene to cultured cells to a final concentration of 10 μg/ml.
Note: Polybrene is an efficient infection reagent used to introduce viral vectors into mammalian cells. Polybrene acts by neutralizing the charge repulsion between virions and the cell surface (Davis et al., 2004). - Using a 5 ml pipet, add 4 ml of filtered viral particles to cultured cells.
- Centrifuge 10 cm2 plate at 250 x g for 5 min.
Note: This step helps the physical contact of viruses with surface of cells increasing infection efficiency. - Place cells in 37 °C, 5% CO2 incubator for 18-24 h.
Note: Infection efficiency can be affected by incubation time. Troubleshooting using different incubation times can be performed to identify most efficient incubation time for a specific cell type. - Aspirate the medium and add 10 ml of fresh RPMI supplemented with 10% FBS and 1% PenStrep. Incubate at 37 °C for 24 h.
- Select infected 22Rv1 prostate cancer cells by adding Puromycin at a final concentration of 2 µg/ml to culture media.
Note: Puromycin concentration will depend on cell type. - Replace medium containing Puromycin every 3 days. Cells that have incorporated the pLenti CMV Puro LUC will grow up under antibiotic selection conditions.
- Check that the cells have incorporated the luciferase plasmid.
Note: Multiple methods can be used to test successful luciferase. Examples are detection of luciferase by PCR or measuring luciferase activity upon D-Luciferin exposure (see Figure 2).
- Add Polybrene to cultured cells to a final concentration of 10 μg/ml.
- Injection of luciferase-tagged cells into immunocompromised mice
In this protocol, we performed subcutaneous injection of luciferase-tagged tumor cells. However, alternative injection sites such as the mouse prostate (orthotropic), which promotes a prostate microenvironment, and subrenal capsule injection, which enhances successful engraftment and preserves tumor heterogeneity, can be performed based on investigator preference (Hidalgo et al., 2014; Siolas et al., 2013).
Note: Conduct all animal procedures in compliance with protocols approved by the institutional animal care committee. This protocol has been conducted at our institution under a specific Animal Care Committee in accordance and compliance with all relevant regulatory and institutional agencies, regulations and guidelines.- Harvest attached cells by removing culture media, washing cells with 10 ml 1x DBPS two times and trypsinizing cells using 2 ml 0.05% Trypsin. Recover 2 ml trypsinized cells to 15 ml tube containing 4 ml RPM1 supplemented with 10% FBS and 1% PenStep.
- Count cells using a hemocytometer.
- Mix 106 cells suspended in 100 μl of RPMI supplemented with 100 μl of extracellular matrix (1:1 ratio) and place on ice.
- Anesthetize mice prior to subcutaneous injection in a chamber supplying 5% (v/v) inhaled isoflurane in 1 L/min of oxygen.
- Using a 25 gauge needle and a 1 ml syringe, inject 250 μl of cell and extracellular matrix suspension subcutaneously into the flanks of immunodeficient NSG mice.
- Harvest attached cells by removing culture media, washing cells with 10 ml 1x DBPS two times and trypsinizing cells using 2 ml 0.05% Trypsin. Recover 2 ml trypsinized cells to 15 ml tube containing 4 ml RPM1 supplemented with 10% FBS and 1% PenStep.
- Bioluminescence imaging
The IVIS Spectrum imager expresses the bioluminescent signal in photons per second and displays it as an intensity map. The luminescence, which is the consequence of the photon flux emitted by the luciferase-expressing cells, directly correlates to the size of the tumor and can be measured at the site of injection using a region of interest (ROI) tool. The optical emission image can be adjusted to provide optimal contrast and resolution without affecting quantitation using the Living Image© in vivo imaging software.- Prepare a fresh stock solution of D-Luciferin at 15 mg/ml in 1x DPBS.
- Anesthetize the mice using isoflurane vaporizer and place them inside the camera box of the IVIS Spectrum imager.
- Run a background image of the mice.
- Place the mice in dorsal recumbency (abdomen face up) and inject 150 mg/kg of D-Luciferin (from step F1) intra-peritoneally (ip) 5 min before imaging. The preferred region of injection is the mouse lower right quadrant of the abdominal region.
- Taking into account the time dependent D-Luciferin uptake by tumor cells, run sequential images of the mice every 2 min until luminescence saturation is reached.
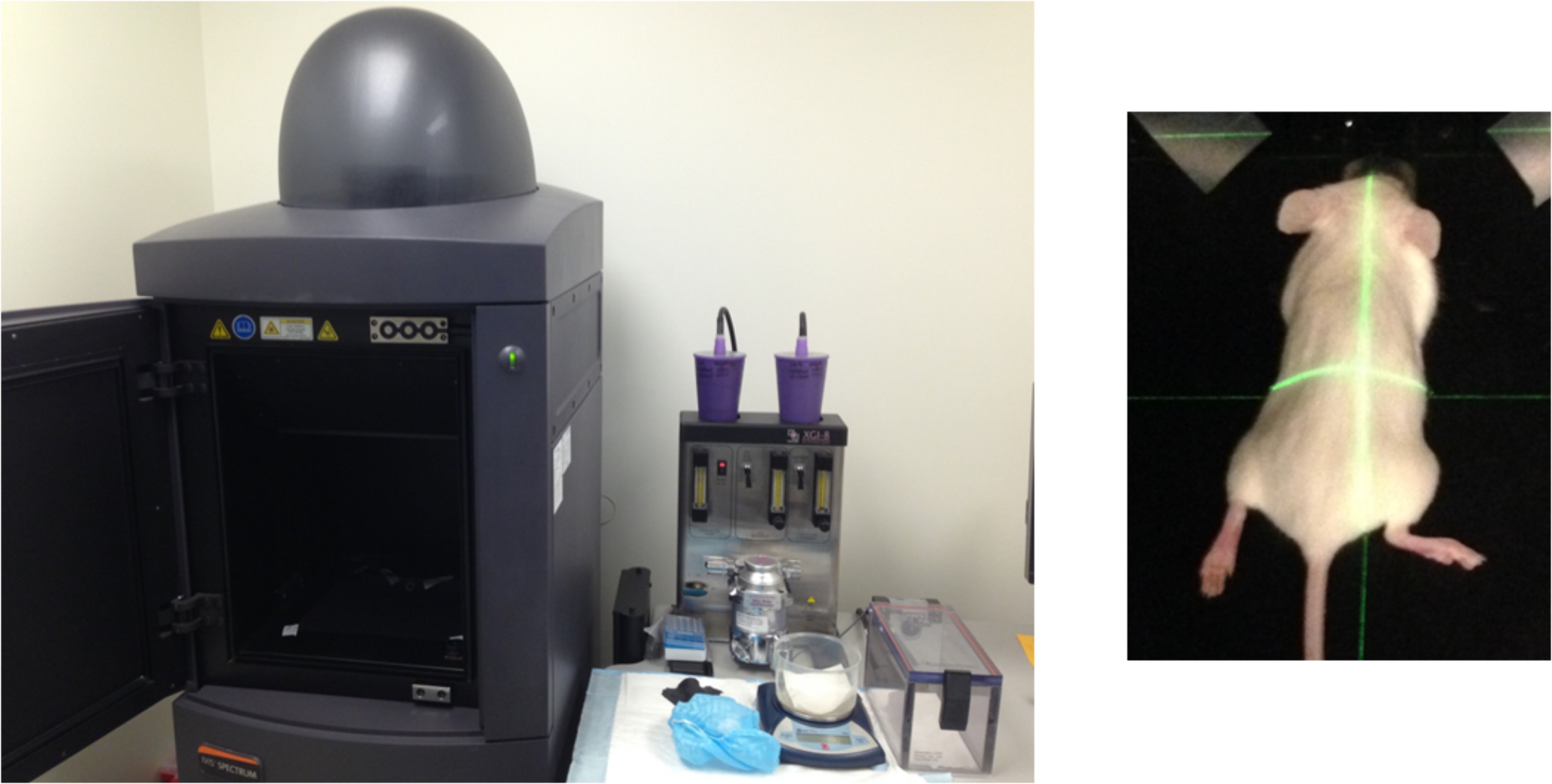

Figure 2. Image illustrates the IVIS spectrum imager and a mouse placed inside the camera box
- Prepare a fresh stock solution of D-Luciferin at 15 mg/ml in 1x DPBS.
Representative data
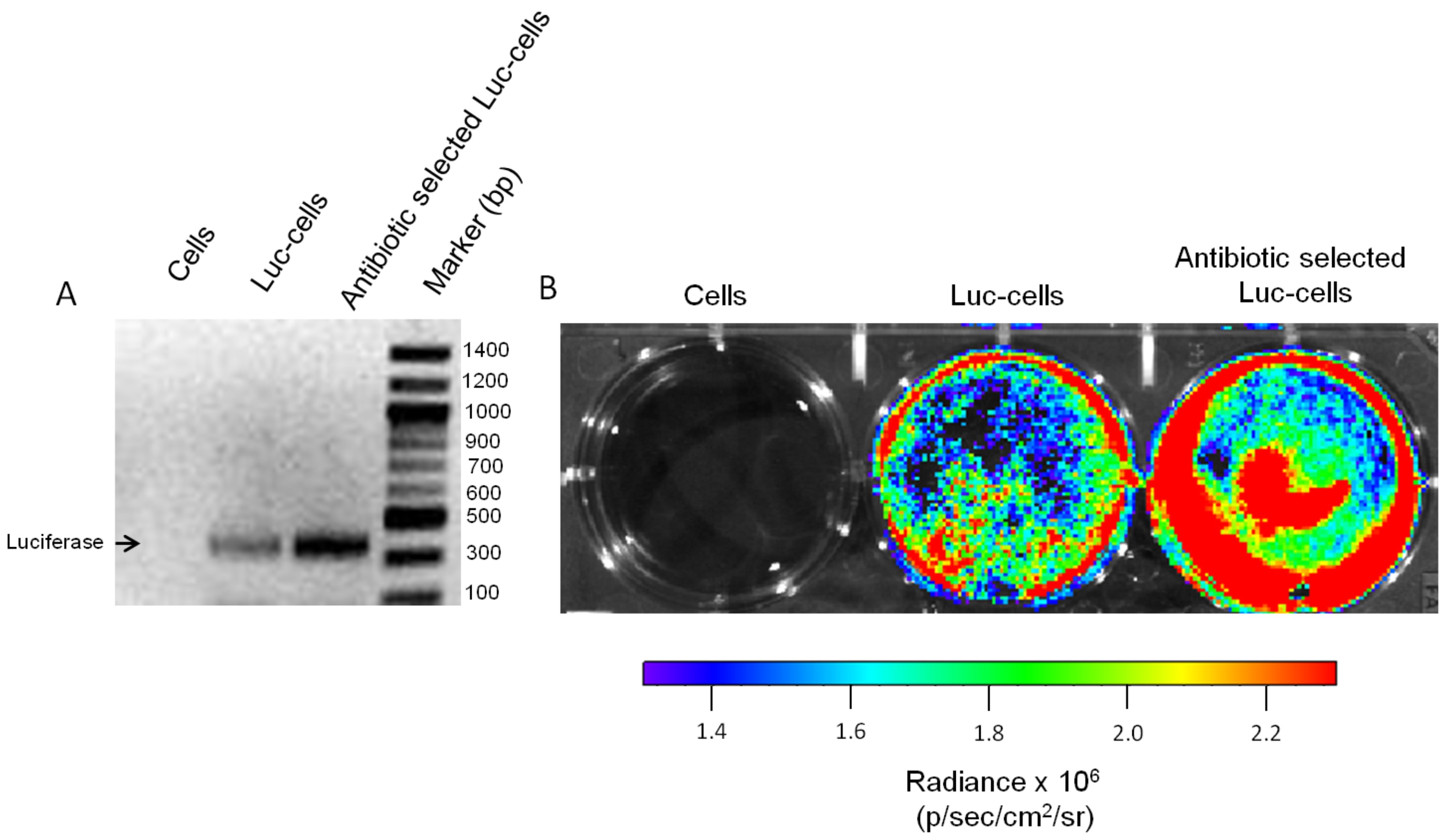
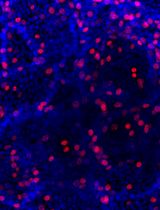
Figure 3. Luciferase detection in cells. Multiple methods can be used for testing the success of cell infection, which include but are not limited to: A. Genetic detection of luciferase gene by PCR and/or B. Testing luciferase activity.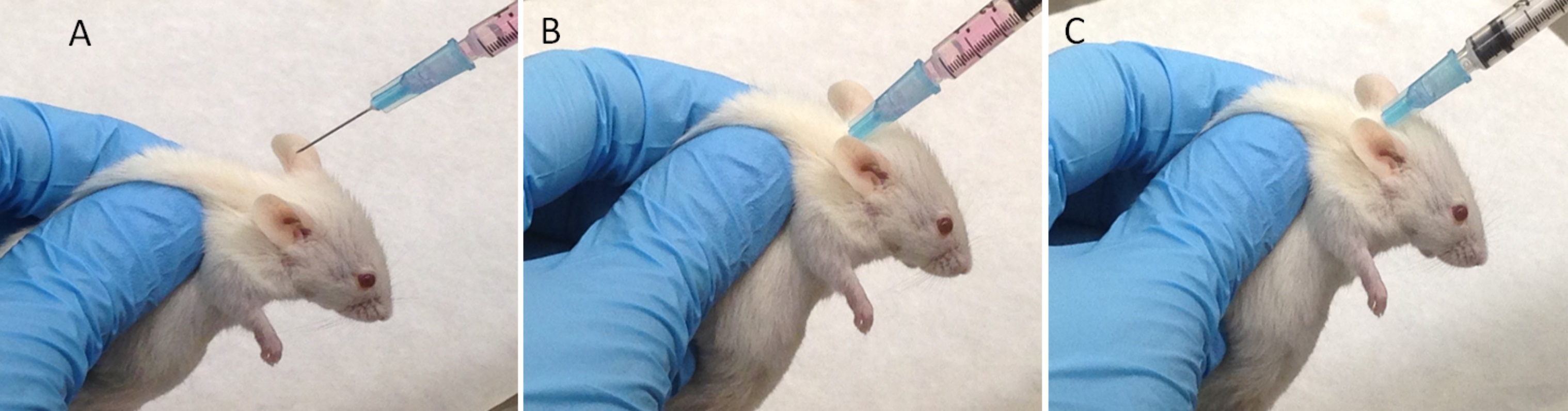
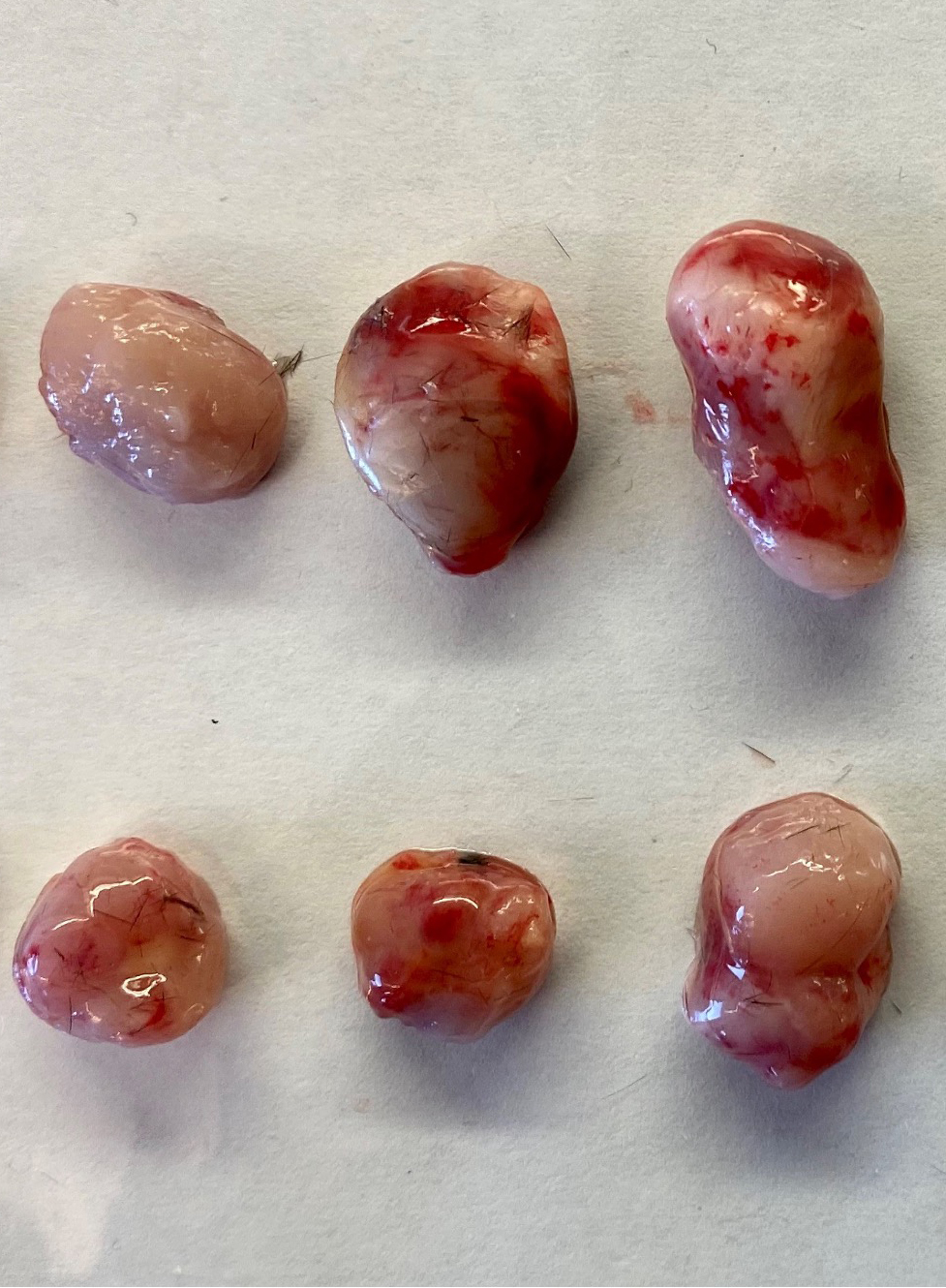
Figure 4. Subcutaneous injection of cells in mice. A. Lift the skin over the back of the neck to make a tent. B. Insert the needle at the tent base, holding it parallel to the animal’s body to avoid puncturing underlying structures. C. Aspirate to create a light vacuum and ensure that the needle has not entered a blood vessel. Slowly inject the cell and extracellular matrix suspension. 
Figure 5. Bioluminescence in vivo imaging. A. Representative luciferase time course (every 2 min) imaging of mice bearing subcutaneous tumors injected with luciferin. B. Graph illustrates the bioluminescence saturation of the whole mouse (displayed in A.) obtained until time point 8 min, when the experiment was decided to be concluded.
Acknowledgments
We thank the TJ Martell Foundation and Agilent Technologies for their support.
References
- Contag, C. H., Spilman, S. D., Contag, P. R., Oshiro, M., Eames, B., Dennery, P., Stevenson, D. K. and Benaron, D. A. (1997). Visualizing gene expression in living mammals using a bioluminescent reporter. Photochem Photobiol 66(4): 523-531.
- Contag, P. R., Olomu, I. N., Stevenson, D. K. and Contag, C. H. (1998). Bioluminescent indicators in living mammals. Nat Med 4(2): 245-47.
- Davis, H. E., Rosinski, M., Morgan, J. R. and Yarmush, M. L. (2004). Charged polymers modulate retrovirus transduction via membrane charge neutralization and virus aggregation. Biophys J 86(2): 1234-1242.
- Edinger, M., Sweeney, T. J., Tucker, A. A., Olomu, A. B., Negrin, R. S. and Contag, C. H. (1999). Noninvasive assessment of tumor cell proliferation in animal models. Neoplasia 1(4): 303-10.
- Harvey, E. N. (1927). The oxidation-reduction potential of the luciferin-oxyluciferin system. J Gen Physiol 10(3): 385-393.
- Hidalgo, M., Amant, F., Biankin, A. V., Budinska, E., Byrne, A. T., Caldas, C., Clarke, R. B., de Jong, S., Jonkers, J., Maelandsmo, G. M., Roman-Roman, S., Seoane, J., Trusolino, L. and Villanueva, A. (2014). Patient-derived xenograft models: an emerging platform for translational cancer research. Cancer Discov 4(9): 998-1013.
- Siolas, D. and Hannon, G. J. (2013). Patient-derived tumor xenografts: transforming clinical samples into mouse models. Cancer Res 73(17): 5315-19.
- Sweeney, T. J., Mailander, V., Tucker, A. A., Olomu, A. B., Zhang, W., Cao, Y., Negrin, R. S. and Contag, C. H. (1999). Visualizing the kinetics of tumor-cell clearance in living animals. Proc Natl Acad Sci U S A 96(21): 12044-12049.
- Vidal, S. J., Rodriguez-Bravo, V., Quinn, S. A., Rodriguez-Barrueco, R., Lujambio, A., Williams, E., Sun, X., de la Iglesia-Vicente, J., Lee, A., Readhead, B., Chen, X., Galsky, M., Esteve, B., Petrylak, D. P., Dudley, J. T., Rabadan, R., Silva, J. M., Hoshida, Y., Lowe, S. W., Cordon-Cardo, C. and Domingo-Domenech, J. (2015). A targetable GATA2-IGF2 axis confers aggressiveness in lethal prostate cancer. Cancer Cell 27(2): 223-239.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Carceles-Cordon, M., Rodriguez-Fernandez, I., Rodriguez-Bravo, V., Cordon-Cardo, C. and Domingo-Domenech, J. (2016). In vivo Bioluminescence Imaging of Luciferase-labeled Cancer Cells. Bio-protocol 6(6): e1762. DOI: 10.21769/BioProtoc.1762.
Category
Cancer Biology > General technique > Tumor formation
Cancer Biology > Tumor immunology > Tumor formation > Cell isolation and detection
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link