- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Quantification of Rae-1 Staining Intensity in Glomeruli
Published: Vol 6, Iss 5, Mar 5, 2016 DOI: 10.21769/BioProtoc.1754 Views: 7282
Reviewed by: Ivan ZanoniYang FuAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Protocol to Isolate Germinal Centers by Laser Microdissection
Farbod Bahreini [...] Kathrin Kalies
Jun 5, 2022 2534 Views
Immunohistochemistry of Immune Cells and Cells Bound to in vivo Administered Antibodies in Liver, Lung, Pancreas, and Colon of B6/lpr Mice
Kieran Adam and Adam Mor
Jul 20, 2022 3624 Views
Monitoring Group 2 Innate Lymphoid Cell Biology in Models of Lung Inflammation
Jana H. Badrani [...] Taylor A. Doherty
Jul 20, 2023 2711 Views
Abstract
NKG2D is expressed on all NK cells and on subsets of NKT, CD8, CD4 and γδ T cells. NKG2D is activated by NKG2D ligands, a stress-induced family of MHC-I-like proteins. These ligands are upregulated on stressed/infected cells and are not widely expressed on healthy adult tissue. NKG2D ligands have been widely studied as potential targets for immunotherapeutic approaches in cancer and auto-immune diseases such as Systemic Lupus Erythematosus. Here we describe a method for quantifying the expression levels of the Rae-1 NKG2D ligand in the glomeruli of healthy and diseased individuals via a novel algorithm. The MRL mouse was used as a positive control strain as it spontaneously generates a lupus-like phenotype, one of the main effects of which is severe glomerulonephritis. MRL/MpJ mice develop this phenotype spon¬taneously at ~12 months of age, whereas MRL/MpJlpr mice, which have the same genetic background but which generated a spontaneous homozygous mutation in the Fas allele, develop similar but more severe symptoms by ~3-4 months of age (Spada et al., 2015).
Keywords: NKG2D ligandsMaterials and Reagents
- Standard Cryomold (Electron Microscopy Sciences, Tissue-Tek, catalog number: 4557 )
- Female MRL/MpJ [3 months (healthy) and 12 months (diseased)] and MRL/MpJ-Faslpr/J [9 weeks (pre-disease) and 3 months (diseased)]
- Peroxidase Block (Dako, catalog number: K4009 )
- Goat anti-mouse Rae-1γ (R&D Systems, catalog number: AF1136 )
- Goat Histofine Simple Stain kit (NICHIREI CORPORATION, catalog number: 414161F )
- AEC + Substrate-Chromogen (Dako, catalog number: K3464 )
- PBS (Sigma-Aldrich, catalog number: P5493 )
- Aqueous mounting medium (Vector Labs, catalog number: H-5501 )
- Acetone (Sigma-Aldrich, catalog number: 320110 )
- Tissue-Freezing Medium (Leica Biosystems Nussloch GmbH, catalog number: 14020108926 )
- Haematoxylin (Sigma-Aldrich, catalog number: H9627 )
Equipment
- Leica Leitz DM RB Microscope (Leica Microsystems) with an adapted Olympus camera (Olympus Imaging America, model: DP70)
- Macintosh or Windows PC
Software
- Photoshop CS6 (Adobe)
- Matlab v. 2013a (MathWorks)
Procedure
- Sample preparation
- Kidneys were removed via a ventral midline incision into the abdomen of a sacrificed mouse. The intestines were retracted laterally to expose the animal’s kidneys. The kidneys are freed from the surrounding tissue and excised.
- Kidneys were briefly washed in 1x PBS, placed within a cryo-mold, covered with OCT tissue-freezing medium and snap frozen by placing the cryo-mold on top of liquid nitrogen.
- The frozen sections kidneys were then cut in 10 μm sections using a cryostat.
- For general guidelines on how to cut sections please refer to the work performed by Fischer et al., (2008).
- Samples to be processed were thawed at RT for 10 min and then fixed with ice-cold 100% acetone for 10 min at -20 degrees.
Notes:
- Isotype-incubated slides were used as negative controls.
- All wash steps were performed in coplin jars.
- Sections were washed for 2 x 5 min in 1x PBS.
- Endogenous peroxidase was blocked using DAKO’s endogenous peroxidase blocker for 7 min at RT as per manufacturer’s instructions.
- Sections were washed for 2 x 5 min in 1x PBS.
- Sections were incubated with Goat anti-mouse Rae-1γ antibody (1/100 dilution in 1x PBS with 10% FBS and 2.5% BSA) overnight at 4 °C.
- Sections were washed for 2 x 5 min in 1x PBS.
- Sections were incubated with Goat Histofine Simple Stain kit for 30 min at RT as per manufacturer’s instructions.
- Sections were washed for 2 x 5 min in 1x PBS.
- Sections were stained with AEC+ Substrate-Chromogen for 15-20 min at RT as per manufacturer’s instructions.
- Reaction was stopped by rinsing the slides gently with distilled water from a wash bottle (do not focus flow directly on tissue).
- Sections were hematoxylin counterstained for 3 min.
- Reaction was stopped by placing slides under running tap water for 10 min.
- 1 drop of aqueous mounting medium was placed on each slide and coverslip was carefully added as not to form bubbles.
- Slides were allowed to dry for 1 h at RT.
- Chromogen deposition was measured by quantitative immunohistochemistry by use of an established method (see section B "Image analysis").
- Images of glomeruli (100x magnification) were acquired under a Leica microscope (vertical Leitz DM RB) with an adapted Olympus (DP70) camera; image files were saved in a tagged image file format (antibody stained images are referred to as EXP and control images are referred to as CONTROL).
- Kidneys were removed via a ventral midline incision into the abdomen of a sacrificed mouse. The intestines were retracted laterally to expose the animal’s kidneys. The kidneys are freed from the surrounding tissue and excised.
- Image analysis
All images were then opened in Photoshop and processed as following [as per Matkowskyj et al., (2000)]:- Open Image in Photoshop.
- Define size parameters for Marquee tool.
- Double click Marquee icon.
- Define area in pixels (200 x 200 pixel region).
- Double click Marquee icon.
- Identify region to be analyzed with Marquee tool.
- Store in a new file.
- Function “Edit” select “Copy”.
- Function “Edit” select “New File”.
- In the new file select “Paste”.
- Function “Save” select tiff for file type.
- Function “Edit” select “Copy”.
- Store a new file in Matlab folder.
- Open Matlab.
At the cursor (») type
»a= double(imread(‘EXP’,‘tiff’));
»b=double(imread(‘CONTROL', ‘tiff’));
»ea=norm(a(:,:,1)) +norm(a(:,:,2)) +norm(a(:,:,3));
»eb=norm(b(:,:,1)) +norm(b(:,:,2)) +norm(b(:,:,3));
»et=abs(ea-eb);
»en=sqrt(et∘2/N) - Following this protocol the amount of chromogen/pixel was determined by selecting glomeruli (25 glomeruli/group) in a 200 x 200 pixel region and subtracting the mathematical energy (EM - calculated at step B6) of the control slide (CONTROL - not exposed to primary antibody) from that of a homologous glomerulus on the experimental slide (EXP - exposed to Rae-1 antibody).
Note: The 200 x 200 pixel region was used because it proved to be the region size which was capable of including the glomeruli of all samples in the most homologous way possible. This size must be adapted according to the samples at hand.
- Open Image in Photoshop.
Representative data
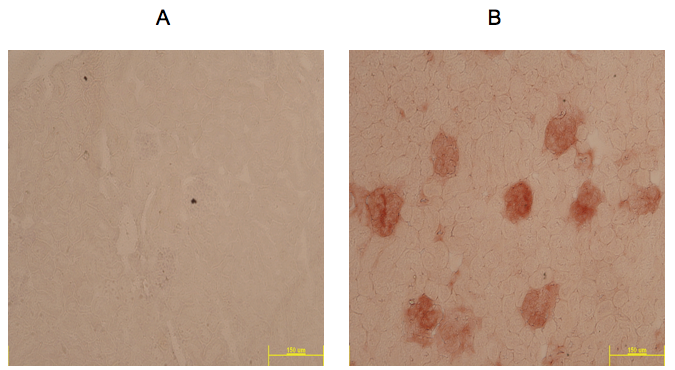
Figure 1. Example of IHC Rae-1 staining (red) in kidneys from 1 year old MRL/MpJ mouse model taken at 10x. A. Negative control slide; B. Slide incubated with Rae-1 antibody.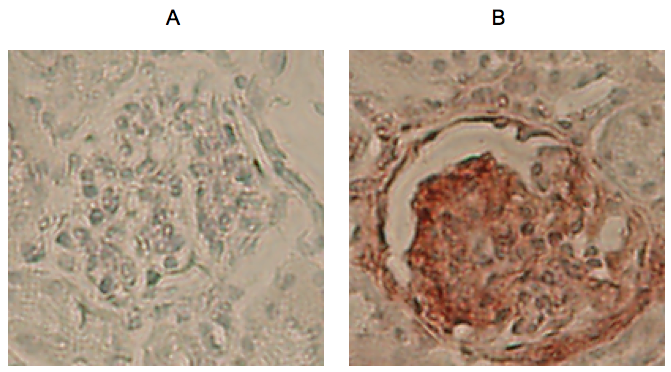
Figure 2. Example 200 x 200 pixel regions extracted from glomeruli at 100x magnification. A. Glomerulus from negative control slide; B. Glomerulus from Rae-1 incubated slide.
Representative results are from the quantification of Rae-1 in the glomeruli of healthy, pre-disease and diseased models of MRL mice. Using this technique, we were capable of determining that Rae-1 expression levels were higher in diseased models of MRL mice compared to healthy and pre-diseased ones (Spada et al., 2015). 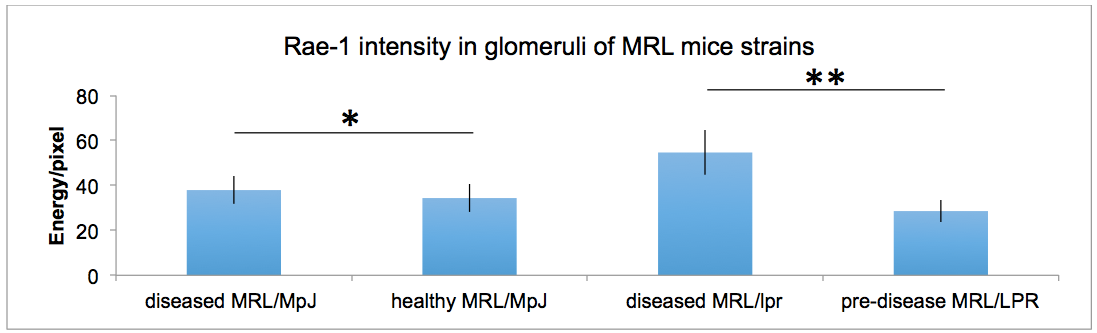
Figure 3. Quantification of Rae-1γ staining in MRL mouse glomeruli. Data were analyzed with a 2-tailed, unpaired Student’s t-test (mean 6 SD; n = 5/group; **P, 0.0001; *P, 0.0025).
Notes
- This technique allows for the quantification of the absolute amount of Rae-1 antibody chromogen per pixel in the glomeruli of healthy and diseased mice. However this technique is not limited to the glomeruli nor to the Rae-1 antibody, as it can be modified as needed for the determination of staining intensity of any antibody in any cellular region or structure.
- It is crucial that all samples (including negative controls) be processed and analyzed simultaneously in order to minimize variation in the staining results.
Acknowledgments
This work was supported, in part, by grants from the Spanish Ministry of Economy and Competitiveness (SAF-2008-00471 and SAF-2011-23639; to D. F. B.), the Madrid regional government (CCG08-CSIC/SAL-3451; to D. F. B.), and the Spanish Ministry for Health, Social Services and Equality, Carlos III Health Institute Cooperative Research Thematic Network program: Research Network in Inflammation and Rheumatic Diseases (RIER; RD08/ 0075/0015; to D. F. B.) and Spanish National Biobank Network (RD09/0076/00101 to the FJD). R. S. was supported by an FPI predoctoral fellowship from the Spanish Ministry of Economy and Competitiveness. We want to thank the work from Matkowskyj et al. (2000) which served as the basis for this protocol.
References
- Fischer, A. H., Jacobson, K. A., Rose, J. and Zeller, R. (2008). Cryosectioning tissues. CSH Protoc 2008: pdb prot4991.
- Matkowskyj, K. A., Schonfeld, D. and Benya, R. V. (2000). Quantitative immunohistochemistry by measuring cumulative signal strength using commercially available software photoshop and matlab. J Histochem Cytochem 48(2): 303-312.
- Spada, R., Rojas, J. M., Perez-Yague, S., Mulens, V., Cannata-Ortiz, P., Bragado, R. and Barber, D. F. (2015). NKG2D ligand overexpression in lupus nephritis correlates with increased NK cell activity and differentiation in kidneys but not in the periphery. J Leukoc Biol 97(3): 583-598.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Spada, R. and Barber, D. F. (2016). Quantification of Rae-1 Staining Intensity in Glomeruli. Bio-protocol 6(5): e1754. DOI: 10.21769/BioProtoc.1754.
Category
Cancer Biology > Invasion & metastasis > Biochemical assays
Immunology > Immune cell staining > Immunodetection
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









