- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
An Assay to Test the Capacity of Arabidopsis Plant Defensin Type1 Protein to Induce Cellular Zinc (Zn) Tolerance in Yeast
Published: Vol 5, Iss 22, Nov 20, 2015 DOI: 10.21769/BioProtoc.1653 Views: 8748
Reviewed by: Tie LiuDe Michele Roberto

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Determination of Boron Content Using a Simple and Rapid Miniaturized Curcumin Assay
Thotegowdanapalya C. Mohan and Alexandra M. E. Jones
Jan 20, 2018 8728 Views
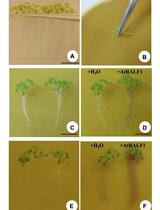

Evaluation of Root pH Change Through Gel Containing pH-sensitive Indicator Bromocresol Purple
Aparecida L. Silva [...] Daniel S. Moura
Apr 5, 2018 10042 Views

MAMP-triggered Medium Alkalinization of Plant Cell Cultures
Gabriel L. Fiorin [...] Paulo J.P.L. Teixeira
Apr 20, 2020 4995 Views
Abstract
Heterologous expression of genes in budding yeast Saccharomyces cerevisiae (S. cerevisiae) is especially suitable to functionally study the corresponding encoded protein at the cellular level (Bonneaud et al., 1991). This is mainly because many strains defective in specific activities are available and could be complemented by homologous genes existing across the eukaryotic kingdom (http://www.yeastgenome.org/). However, the protocol we describe here is not a complementation but a “gain-of-function” assay. It is based on a drop-test assay that we have set up to assess the cellular zinc tolerance conferred by the expression of heterologous genes in the wild-type S. cerevisiae. Different dilutions of a yeast culture expressing the heterologous gene of interest are grown on a range of zinc-enriched plates, and are then compared to the control yeast expressing the empty vector. Working with different concentrations of both yeast and zinc are essential to succeed in describing zinc tolerance phenotype upon yeast transformation (Mirouze et al., 2006). This test has also proven to be valuable to differentiate among related members of gene families as exemplified for Arabidopsis Plant Defensin type1 (Shahzad et al., 2013).
Materials and Reagents
- Plastic foil (Saran Wrap)
- Sterile square 12 cm x 12 cm Petri dishes (CML HealthCare, catalog number: BP 120SJ )
- Sterile-R Filtropur S (0.2 µm) (SARSTEDT AG, catalog number: 83.1826.001 )
- 20 ml sterile syringe without needle (Medline Scientific, catalog number: BS-20ES )
- 50 ml sterile tubes (SARSTEDT AG, catalog number: 62.547.254 )
- Autoclaved 1.5 ml Eppendorf tubes (SARSTEDT AG, catalog number: 72.690 )
- Sterile inoculating loops (Greiner Bio One International GmbH, catalog number: 731175 )
- Sterile pipet tips
- Parental BY4741 S. cerevisiae strain (Winston et al., 1995; Brachmann et al., 1998) having the genotypes MATa, his3∆1, leu2∆0, met15∆0, ura3∆0) expressing: the pFL38H vector (Bonneaud et al., 1991), complementing the his3∆1 auxotrophy
- Transformed BY4741 yeast harbouring both: the pFL38H vector, complementing the his3∆1 auxotrophy and the pYX212 vector (Ingenius, R&D systems) complementing the ura3∆0 mutation and expressing the entire coding sequences of Arabidopsis PDF1s, i.e., including the signal peptide (Shahzad et al., 2013)
- Yeast nitrogen base (YNB) without amino acids and without ammonium sulfate (BD, Difco, catalog number: 233520 )
- Ammonium nitrate (NH4NO3) (Sigma-Aldrich, catalog number: A9642 )
- D(+)Glucose anhydrous (C6H12O6) (EUROMEDEX, catalog number: UG3050 )
- Succinic acid (C4H6O4) (Sigma-Aldrich, catalog number: S7501 )
- Potassium hydroxide (KOH) (Bio Basic Canada Inc., catalog number: PB0441 )
- Agarose type D-5 (EUROMEDEX, catalog number: D5 )
- Zinc sulphate heptahydrate (ZnSO4.7H2O) (Sigma-Aldrich, catalog number: Z4750 )
- L-leucine (C6H13NO2) (Sigma-Aldrich, catalog number: L8912 )
- L-methionine (C5H11NO2S) (Sigma-Aldrich, catalog number: M9625 )
- Sterile ultra-pure water
- 0.5 M zinc sulfate
- 10 N KOH (see Recipes)
- 0.075 M L-leucine (see Recipes)
- 0.1 M L-methionine (see Recipes)
- Sterile liquid and solid selective yeast nitrogen base (YNB) media (see Recipes)
Equipment
- Sterile 250 ml borosilicate glass Erlen flask
- 1,000 ml bottle
- Typical 20 µl pipet
- Autoclave
- Heating oven (Universal Memmert, model: 400 )
- New BrunswickTM Agitator (Eppendorf, model: Innova 44 )
- A normal cell density meter (Thermo Fisher Scientific, model: 40 )
- Laminar flow hood (ADS, model: VP120 )
- Centrifuge (Eppendorf, model: 5804R )
- Integral water purification system (Merck Millipore Corporation, model: Milli-Q+ )
- Scanner (Epson, model: 124OU )
Procedure
Note: For each yeast clone to be analyzed.
On day -1
- Use a sterile inoculating loop to inoculate each yeast clone to be tested in 20 ml liquid selective YNB media in sterile 250 ml glass Erlen flask.
Note: At least three independent yeast clones are tested for each construct (harboring either recombinant or control plasmids). - Culture yeast cells at 30 °C under 200 rpm agitation Over Night (O/N).
On day 0
Prepare Petri dish for drop assay.
- Prepare sterile selective YNB solid media containing the appropriate amino acids and appropriate zinc sulfate concentration to be tested.
- Under laminar flux hood, pour 50 ml of sterile selective YNB solid media in each square 12 cm x 12 cm Petri dish.
Note: Tolerance to zinc excess is usually tested around the final concentrations of 23, 25, 27.5, 30 and 32.5 mM. The control condition is without zinc added (≈1.4 µM). The leucine and methionine auxotrophy are complemented by the addition of the corresponding amino acids to the final concentrations of 0.75 mM and 0.33 mM, respectively. - Let media solidify, cover opened, for 15 min. Close and store square Petri dish under the hood.
Prepare yeast for drop assay.
Note: All subsequent manipulations with yeast are performed under sterile conditions in laminar flow hood. - Transfer O/N yeast culture in sterile 50 ml Falcon tubes.
- Centrifuge for 5 min at 5,000 x g at room temperature.
- Discard supernatant.
- Suspend yeast culture with the same volume of sterile water (20 ml).
- Repeat steps 7 to 9.
- Measure optical density at 600 nm after suitable dilution (usually 1/10) to get an OD600nm of around 0.4.
- In a 1.5 ml sterile Eppendorf tube, make appropriate yeast culture dilution with sterile water in order to reach OD600nm of 1 in 500 µl volume and then make serial dilution in order to reach OD600nm of 10-1, 10-2, 10-3 in a maximum of 500 µl.
- Use a pipet and sterile filter tips to drop 10 µl of each yeast dilution onto selective YNB media supplemented with ZnSO4 at various final concentrations (see Figure 1).
Notes:- For steps 10 to 12: Yeast cells sediment particularly rapidly; hence make sure to fully agitate the yeast culture before taking a sample and / or before measuring optic density of a suspension and/or before drop assay.
- For drop assays, separate the center of each drop by ≈1 cm.
- For steps 10 to 12: Yeast cells sediment particularly rapidly; hence make sure to fully agitate the yeast culture before taking a sample and / or before measuring optic density of a suspension and/or before drop assay.
- Allow drops to dry under the laminar flux hood.
- Close Petri dish and wrap them plastic foil (type Saran Wrap).
- Incubate in 30 °C incubator upside down.
- Monitor yeast growth every day and record results by scanning the Petri dish under black screen at a 600 dots per inch (dpi) resolution (usually less than one minute per scan). For each zinc concentration, the yeast expressing the protein of interest is compared to the yeast harboring the empty pYX212.
Note: Control plates, i.e., without zinc added, are usually read 2 days after incubation. This is necessary to verify that the expression of the protein of interest does not disturb yeast growth at very low zinc concentration. Plates for zinc tolerance assay (upon zinc addition) are usually read between 4 to 7 days after incubation.
Representative data
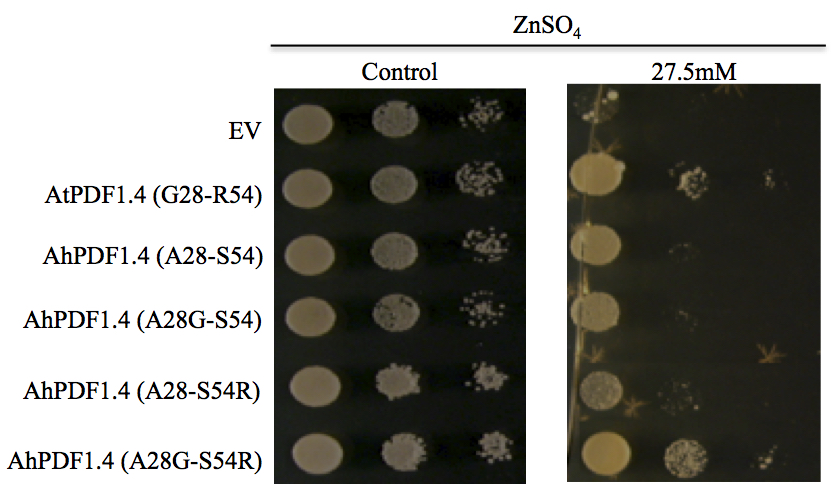

Figure 1. In vitro functional characterisation of amino-acid substitutions impacting zinc tolerance capability in AhPDF1.4 and AtPDF1.4. Serial dilutions of the S. cerevisiae BY4741 strain expressing the pYX212 empty vector (EV) or pYX212 harbouring AhPDF1.4, AtPDF1.4 or its mutated versions of were analysed. Replacement of either A28 to G (A28G) or S54 to R (S54R) or both (A28G- S54R) were generated. The various yeast transformants were spotted on medium supplemented with 1.4 µM (control), or 27.5 mM ZnSO4, as indicated above the panels. Each spot was made with 10 µl of a yeast culture diluted at the OD600nm mentioned below the drops. Pictures were taken at day 2 for control and day 11 for the Zn treatments; they are representative of the three experiments, which have been performed with three independent yeast transformants. Reproduced from (Shahzad et al., 2013; Supplementary Figure 4).
Notes
The BY4741 S. cerevisiae strain has auxotrophic markers, in particular for the production of the amino acid histidine (his3∆1). Free histidine is a potent metal chelator and could, in particular, chelate zinc if this metal is present in the culture medium. The histidine (his3∆1) auxotrophy must thus be complemented by the expression of the corresponding HIS gene (here expressed from the pFL38 plasmid) and not by the addition of free histidine, which would bias the zinc tolerance test.
Recipes
- 10 N KOH
Filter sterilized using Sterile-R Filtropur S (0.2 µm)
Keep at +4 °C - 0.075 M L-leucine
Filter sterilized using Sterile-R Filtropur S (0.2 µm)
Keep at +4 °C - 0.1 M L-methionine
Filter sterilized using Sterile-R Filtropur S (0.2 µm)
Keep at +4 °C - Sterile liquid and solid selective YNB media (1 L)
- Mix in ultra-pure water
YNB without amino acids without ammonium sulfate: 1.7 g
NH4NO3: 6.4 g
D-Glucose: 20 g
Succinic acid: 5.9 g - Adjust pH to 4.5 with 10 N KOH
- Adjust volume and dispatch 500 ml in 1,000 ml Bottle
- For solid media add:
Agarose: 20 g/L - Autoclave at 120 °C for 20 min
- Upon cooling of solid media, add necessary sterile amino acid solutions complementing auxotrophy (here leucine and methionine) and sterile zinc in order to reach concentrations that will be tested.
Note: Care should be taken to add necessary sterile amino acid solutions and sterile zinc when solid media has cooled down to ~ 40 to 45 °C. This will prevent both the degradation of added amino acids and the precipitation of the metal.
- Mix in ultra-pure water
Acknowledgments
This work was supported by the University of Montpellier (LM), the French Ministère de l’Enseignement Supérieur et de la Recherche (PB), the Pakistani Higher Education Commission (ZS) and the Centre National de la Recherche Scientifique (FG).
References
- Bonneaud, N., Ozier-Kalogeropoulos, O., Li, G. Y., Labouesse, M., Minvielle-Sebastia, L. and Lacroute, F. (1991). A family of low and high copy replicative, integrative and single-stranded S. cerevisiae/E. coli shuttle vectors. Yeast 7(6): 609-615.
- Brachmann, C. B., Davies, A., Cost, G. J., Caputo, E., Li, J., Hieter, P. and Boeke, J. D. (1998). Designer deletion strains derived from Saccharomyces cerevisiae S288C: a useful set of strains and plasmids for PCR-mediated gene disruption and other applications. Yeast 14(2): 115-132.
- Mirouze, M., Sels, J., Richard, O., Czernic, P., Loubet, S., Jacquier, A., Francois, I. E., Cammue, B. P., Lebrun, M., Berthomieu, P. and Marquès, L. (2006). A putative novel role for plant defensins: a defensin from the zinc hyper-accumulating plant, Arabidopsis halleri, confers zinc tolerance. Plant J 47(3): 329-342.
- Shahzad, Z., Ranwez, V., Fizames, C., Marquès, L., Le Martret, B., Alassimone, J., Gode, C., Lacombe, E., Castillo, T., Saumitou-Laprade, P., Berthomieu, P. and Gosti, F. (2013). Plant Defensin type 1 (PDF1): protein promiscuity and expression variation within the Arabidopsis genus shed light on zinc tolerance acquisition in Arabidopsis halleri. New Phytol 200(3): 820-833.
- Winston, F., Dollard, C. and Ricupero-Hovasse, S. L. (1995). Construction of a set of convenient Saccharomyces cerevisiae strains that are isogenic to S288C. Yeast 11(1): 53-55.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Shahzad, Z., Marquès, L., Berthomieu, P. and Gosti, F. (2015). An Assay to Test the Capacity of Arabidopsis Plant Defensin Type1 Protein to Induce Cellular Zinc (Zn) Tolerance in Yeast. Bio-protocol 5(22): e1653. DOI: 10.21769/BioProtoc.1653.
Category
Microbiology > Heterologous expression system > Saccharomyces cerevisiae
Plant Science > Plant physiology > Ion analysis
Cell Biology > Cell-based analysis > Ion analysis > Zinc
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










