- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Labeling of the Intestinal Lumen of Caenorhabditis elegans by Texas Red-dextran Feeding
Published: Vol 5, Iss 16, Aug 20, 2015 DOI: 10.21769/BioProtoc.1564 Views: 10581
Reviewed by: Peichuan ZhangJyotiska ChaudhuriAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Simultaneous Monitoring Cytoplasmic Calcium Ion and Cell Surface Phosphatidylserine in the Necrotic Touch Neurons of Caenorhabditis elegans
Yoshitaka Furuta and Zheng Zhou
Oct 20, 2021 2663 Views

Live-cell Imaging and Analysis of Germline Stem Cell Mitosis in Caenorhabditis elegans
Réda M. Zellag [...] Abigail R. Gerhold
Jan 5, 2022 4732 Views
SunTag-Based Single-Molecule Translation Imaging in Caenorhabditis elegans
Elise van der Salm [...] Suzan Ruijtenberg
Oct 20, 2025 2353 Views
Abstract
In this method, the intestinal lumen of Caenorhabditis elegans (C. elegans) is labeled with a fluorescent fluid-phase marker, Texas Red-dextran. Since dextran conjugates are membrane impermeable, animals fed with it show a red fluorescent signal in the lumen of the intestine. Texas Red-dextran in the lumen is not efficiently endocytosed by intestinal cells and is not effectively transported to the body cavity paracellularly. It is useful to determine whether round-shaped membrane structures are invaginations from the apical membrane or cytoplasmic vesicles. If the barrier function of the intestinal epithelium is impaired, Texas Red-dextran can leak from the intestinal lumen to the body cavity. Therefore, this method can be used to visualize apical membrane morphology in intestinal cells and to investigate the barrier properties of the intestinal epithelium.
Keywords: C. elegansMaterials and Reagents
- C. elegans strain
GK70 unc-119 (ed3) III; dkIs37[Pact-5-GFP-pgp-1;unc-119(+)]
- Escherichia coli OP50 strain (obtained from the Caenorhabditis Genetics Center)
- Texas Red-dextran (40,000 MW) (Life Technologies, Molecular Probes®, catalog number: D-1829 )
- Levamisole hydrochloride (Sigma-Aldrich, catalog number: L9756 )
- Agarose (Nacalai Tesque, catalog number: 01158-85 )
- Egg buffer (see Recipes)
- M9 buffer (see Recipes)
- Agarose pads (see Recipes)
Equipment
- Siliconized tubes (1.7 ml) (National Scientific Supply, catalog number: 20172-945 )
- Tabletop centrifuge (Hitachi Koki, model: CT15RE )
- Confocal laser scanning microscopy system (Olympus, model: FV1000 )
- Four clean glass slides and a coverslip
Procedure
- Place synchronized L1 larvae on an Escherichia coli (E. coli) OP50-seeded plate and incubate them for 3 days at 20 °C.
- Pick approximately 30-40 young adult animals from an OP50-seeded plate and transfer them into 500 μl of egg buffer in 1.7 ml siliconized tubes.
- Allow the animals to settle with gravity for a few minutes. Remove the supernatant and wash with 500 μl of egg buffer 3 times.
- Remove most of the supernatant, leaving approximately 10-20 μl of worm suspension and add 100 μl of egg buffer containing 1 mg/ml Texas Red-dextran.
- Incubate the animals for 90 min at room temperature with occasional gentle agitation every 30 min.
- Collect the animals by centrifuging at 400 x g for 2 min and discard the supernatant. Wash with 1 ml of egg buffer at least 3 times.
Note: Remove as much Texas Red-dextran in the worm suspension as possible.
- Add 1 M levamisole hydrochloride to the worm suspension to a final concentration 10~20 mM.
- Transfer the worm suspension to a 1.5% agarose pad on a glass slide and cover it with a coverslip.
- Observe the animals with confocal laser scanning microscopy (see Figure 1).

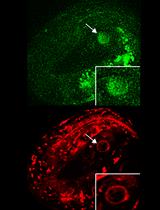
Figure 1. Animals expressing GFP-PGP-1 were fed Texas Red-dextran. GFP-PGP-1 and Texas Red-dextran labeled the intestinal apical membrane (large arrowheads) and the intestinal lumen (large arrows), respectively. Texas Red-dextran that is taken up by feeding is restricted to the intestinal lumen when the junctional seals of intestinal epithelium cells are intact (Carberry et al., 2012). A. RNAi-mediated knockdown of C. elegans chaperonin subunit (CCT-5) disorganized the microvillus actin filament and microtubules, leading to aberrant morphology of the intestinal apical membrane (Saegusa et al., 2014). B. In such animals, the apical membrane, labeled by GFP-PGP-1 (C. elegans P-glycoprotein homolog), formed bubble-like membrane structures invaginated from the apical membrane (B, small arrows). Such structures contain Texas Red-dextran (B, small arrows). By contrast, GFP-PGP-1-containing vesicles underneath the apical membrane (B, small arrowheads) did not contain Texas Red-dextran, indicating that these are exocytic vesicles. Images were obtained using an FV1000 confocal microscopy system. GFP-PGP-1; excitation: 473 nm, detection: 490-540 nm. Texas-Red dextran; excitation: 559 nm, detection: 575 to 675 nm. The scale bars indicate 10 μm.
Notes
- Although Texas Red-dextran is not efficiently taken up by the intestinal cells in this feeding method, some other apical endocytic markers such as rhodamine-dextran (Mr 40K) and Texas Red BSA are reported to be available to visualize endocytic vesicles (Grant et al., 2001).
- Depletion of tight junction components such as claudins (CLC-1-4 in C. elegans) causes the infiltration of dextran conjugates from the intestinal lumen into the body cavity (Asano et al., 2003).
Recipes
- Egg buffer
118 mM NaCl
48 mM KCl
2 mM CaCl2
2 mM MgCl2
25 mM HEPES
pH 7.3
Note: This solution is more physiologically isotonic than M9 for C. elegans tissues (Grant et al., 2001).
- M9 buffer
42 mM Na2HPO4
22 mM KH2PO4
86 mM NaCl
1 mM MgSO4
- Agarose pads (prepare beforehand)
- Melt 1.5% agarose in M9 buffer in a microwave oven and transfer it to a glass tube.
- Keep the agarose in a molten state by placing the tube in a heat block at 65 °C.
- Place a clean glass slide between two glass slides sealed with pieces of labeling tape over both ends.
- Drop the 1.5% agarose onto the center glass slide and place another clean slide on top, perpendicular to the other three slides.
- Gently remove the top perpendicular slide after the agarose solidifies.
- Use the agarose before it dries up.
- Melt 1.5% agarose in M9 buffer in a microwave oven and transfer it to a glass tube.
Acknowledgments
This work has been supported by the JSPS KAKENHI Grant Number 26291036, the Sumitomo Foundation, the Naito Foundation, and the Mochida Memorial Foundation for Medical and Pharmaceutical Research (to Ken Sato). The protocol has been adapted from Saegusa et al. (2014).
References
- Asano, A., Asano, K., Sasaki, H., Furuse, M. and Tsukita, S. (2003). Claudins in Caenorhabditis elegans: their distribution and barrier function in the epithelium. Curr Biol 13(12): 1042-1046.
- Carberry, K., Wiesenfahrt, T., Geisler, F., Stöcker, S., Gerhardus, H., Überbach, D., Davis, W., Jorgensen, E., Leube, R. E. and Bossinger, O. (2012). The novel intestinal filament organizer IFO-1 contributes to epithelial integrity in concert with ERM-1 and DLG-1. Development 139(10): 1851-1862.
- Grant, B., Zhang, Y., Paupard, M. C., Lin, S. X., Hall, D. H. and Hirsh, D. (2001). Evidence that RME-1, a conserved C. elegans EH-domain protein, functions in endocytic recycling. Nat. Cell Biol. 3(6): 573-579.
- Saegusa, K., Sato, M., Sato, K., Nakajima-Shimada, J. and Harada, A. and Sato, K. (2014). Caenorhabditis elegans chaperonin CCT/TRiC is required for actin and tubulin biogenesis and microvillus formation in intestinal epithelial cells. Mol. Biol. Cell 25(20): 3095-3104.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Saegusa, K. and Sato, K. (2015). Labeling of the Intestinal Lumen of Caenorhabditis elegans by Texas Red-dextran Feeding . Bio-protocol 5(16): e1564. DOI: 10.21769/BioProtoc.1564.
- Saegusa, K., Sato, M., Sato, K., Nakajima-Shimada, J. and Harada, A. and Sato, K. (2014). Caenorhabditis elegans chaperonin CCT/TRiC is required for actin and tubulin biogenesis and microvillus formation in intestinal epithelial cells. Mol. Biol. Cell 25(20): 3095-3104.
Category
Cell Biology > Cell imaging > Live-cell imaging
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link