- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Quantitative Evaluation of Competitive Nodulation among Different Sinorhizobium Strains
Published: Vol 5, Iss 15, Aug 5, 2015 DOI: 10.21769/BioProtoc.1555 Views: 7904
Reviewed by: Zhaohui LiuAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
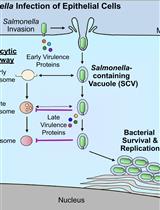
Bacterial Pathogen-mediated Suppression of Host Trafficking to Lysosomes: Fluorescence Microscopy-based DQ-Red BSA Analysis
Mădălina Mocăniță [...] Vanessa M. D'Costa
Mar 5, 2024 2932 Views
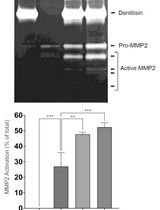
Purification of Native Dentilisin Complex from Treponema denticola by Preparative Continuous Polyacrylamide Gel Electrophoresis and Functional Analysis by Gelatin Zymography
Pachiyappan Kamarajan [...] Yvonne L. Kapila
Apr 5, 2024 2125 Views
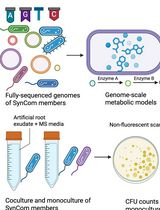
In Silico Prediction and In Vitro Validation of Bacterial Interactions in the Plant Rhizosphere Using a Synthetic Bacterial Community
Arijit Mukherjee [...] Sanjay Swarup
Nov 5, 2025 1710 Views
Abstract
Legumes play a vital role in global food supply because they are uniquely capable of fixing atmospheric nitrogen (N) through symbioses with root and stem nodule bacteria, collectively called the rhizobia. These commonly include bacteria in the genera Rhizobium, Mesorhizobium, Sinorhizobium (Ensifer), and Bradyrhizobium, although other genera of bacteria have now been shown to form root nodule symbioses with several legume species (Weir, 2012). The symbiotic interaction is important for agricultural productivity, especially in less developed countries where nitrogen fertilizer is expensive. However, nodulation ability and competitiveness have practical importance in agricultural production, because the inoculation of efficient rhizobia is often unsuccessful, due to large part to the presence of competitive populations of ineffective indigenous rhizobia in soils (Toro, 1996; Triplett and Sadowsky, 1992). This protocol allows one us to quantitatively evaluate the relative nodulation competitiveness of Sinorhizobium strains.
Materials and Reagents
- Two Sinorhizobium meliloti (S. medicae) strains (strain A and B) with different intrinsic antibiotic resistance (e.g. Strain A is neomycin sensitive and strain B is neomycin resistant at a certain concentration of neomycin)
Note: Alternately, strains can be differentiated by using gfp and rfp, antisera, or other marker genes (Triplett and Sadowsky, 1992).
- Medicago truncatula seeds
- Sodium hypochlorite
- Sodium chloride (Macron Chemical, catalog number: 7581-06 )
- Concentrated sulfuric acid (Thermo Fisher Scientific, catalog number: A300-212 )
- Ethanol (Decon Labs, catalog number: 2701 )
- Antibiotics
- Sunshine mix #5:Turface mixture at 1: 1 ratio (SunGro Horticulture)
- Turface MVP (Profile Product LLC)
- Tryptone (BD, catalog number: 211699 )
- Yeast extract (Thermo Fisher Scientific, catalog number: DF0127071 )
- CaCl2.2H2O (Thermo Fisher Scientific, catalog number: C79-3 )
- Agar (Sigma-Aldrich, catalog number: A1296 )
- KNO3 (Thermo Fisher Scientific, catalog number: P263-500 )
- Ca(NO3)2.4H2O (Acros, catalog number: 423535000 )
- Ca(H2PO4)2 (Spectrum, catalog number: C1145 )
- MgSO4.7H2O (J.T.Baker®, catalog number: 2504 )
- Fe-EDTA (Sigma-Aldrich, catalog number: E6760-5000 )
- MnCl2 (Thermo Fisher Scientific, catalog number: M33-500 )
- H3BO3 (Mallinckrodt, catalog number: 2549 )
- ZnSO4.7H2O (J.T.Baker®, catalog number: 4382 )
- NaMoO4 (Sigma-Aldrich, catalog number: S6646 )
- CuSO4.5H2O (Thermo Fisher Scientific, catalog number: C-493 )
- K2SO4 (EMD Millipore, catalog number: PX1595-1 )
- CaSO4.2H2O (Sigma-Aldrich, catalog number: C3771 )
- Paraffin (Thermo Fisher Scientific, catalog number: P-21 hard )
- TY medium (see Recipes)
- N2-free nutrient solution (see Recipes)
- Sterile paraffin coated sands (see Recipes)
Equipment
- MAGENTA® vessel GA-7 (Sigma-Aldrich, catalog number: V8505 ) and cotton rope (diameter ¼ inch) for Leonard jar assemblies
- Filter paper P5 (Thermo Fisher Scientific, catalog number: 09-801C )
- Plant growth chamber (TC2, Environmental Growth Chambers as an example but other plant growth chambers can be used)
- Autoclave
Procedure
- Medicago seed sterilization
- Place seeds in 15 ml plastic falcon tube. Add 3 ml of sulfuric acid to expose the seed for 5-8 min with occasional mixing in a fume hood with personal protective equipment.
- Rapidly remove the acid with pipet, and wash 4 times with sterilized double-distilled water.
- Soak the seeds in 10% sodium hypochlorite for 90 sec.
- Rinse 8 times with sterile water.
- Place the seeds in a petri dish on a sterile paper filer wetted with 2 ml sterile water.
- Place seeds in the refrigerator at 4 °C (in the dark) for 3-5 days.
- Move seeds to room temperature (in the dark) and let sit for 1 day to pre-germinate.
- Place seeds in 15 ml plastic falcon tube. Add 3 ml of sulfuric acid to expose the seed for 5-8 min with occasional mixing in a fume hood with personal protective equipment.
- Sinorhizobium strains
- Sinorhizobium meliloti strains are grown in TY medium for 2 days, with shaking.
- The cultures are centrifuged at 8,000 x g for 10 min and washed twice with sterile 0.85% sodium chloride.
- The resultant cultures are diluted to107 cells/ml with sterile 0.85% sodium chloride.
- Prepare inoculant mixture using the diluted 107 cells/ml culture. The proportion of two strains can vary from 1:1,000 to 1,000:1.
- Sinorhizobium meliloti strains are grown in TY medium for 2 days, with shaking.
- Co-inoculation and plant growth condition
- Prepare sterile Leonard jar assemblies containing mixture of Sunshine mix #5 and Turface MVP at 1:1 ratio. Nitrogen-free plant nutrient solution was prepared for watering.
- Plant three germinated seeds in 1 cm deep hole and lightly cover the seeds with soil in each Leonard jar.
- Inoculate each seed with 1 ml of inoculant mixtures using a pipette, where the proportion of two strains varied from 1:1,000 to 1,000:1. We suggest using 5 different proportional inoculant mixtures, uninoculated controls, and control plants inoculated with a single strain.
- The jars are incubated in a plant growth chamber at 25 °C with a 16-h light condition and 21 °C for 8 h in the dark. Humidity should be between 50-70%. Light conditions should be 200-350 µmol/m/s.
- After one week, cover soil with sterilized paraffin coated sands and remove extra plants by cutting with a sterile forceps. Each pot only contains 1 plant.
- Prepare sterile Leonard jar assemblies containing mixture of Sunshine mix #5 and Turface MVP at 1:1 ratio. Nitrogen-free plant nutrient solution was prepared for watering.
- Recovery and identification of Sinorhizobium strains in nodules
- After 28 days of cultivation, 10-15 nodules are randomly collected from each plant root, and placed into 1.5 ml centrifuge tube by cutting the root about 0.5 cm on each side of the nodule.
- The surface of the nodules are sterilized by immersion in 95% ethanol for 10 sec and then in 2% (v/v) solution of sodium hypochlorite for 5 min.
- The nodules are rinsed with sterile water, 5 times, and transferred into a 96-well microtiter plate containing 100 μl of sterile 0.85% NaCl.
- The surface-sterilized nodule in each well is crushed by using a sterile toothpick.
- The suspension is streaked on TY plates.
- Three single colonies from each TY plate were transferred onto a TY plates containing antibiotic at specific concentrations that each strain is resistant to.
- Strains in nodules are determined by the antibiotic resistance.
- After 28 days of cultivation, 10-15 nodules are randomly collected from each plant root, and placed into 1.5 ml centrifuge tube by cutting the root about 0.5 cm on each side of the nodule.
- Quantitative evaluation of competitive nodulation
- The competitive ability for nodulation is quantified by nodule occupancy rates in seven inoculant mixtures, with the proposition of two strains ranging from 1:1,000 to 1,000:1.
- Nodule occupancy rate of strain A is calculated by dividing the number of nodules containing strain A (NA) by the total nodules (NA+ NB) tested.
- A strain which has statistically greater nodule occupancy rate when two strains co-inoculated at a 1:1 ratio (determined by microscopic count) has greater competitive ability for nodulation over the other strain.
- The competitiveness is quantitatively evaluated by the linear relationship between the logarithm of ratio of the numbers of nodules formed by each of two strains (NA/NB) and the logarithm of ratio of the cell numbers of two strains (IA/IB): log (NA/NB) = logCAB + k•log (IA/IB) (Amarger and Lobreau, 1982).
- The intercept of the regression line, the CAB value is an index representing the competitiveness of strain A in relation to that of strain B. CAB is close to 1 when two strains have equal competitiveness for nodulation. If CAB is greater than 1, stain A has greater nodulation competitiveness in relation to strain B. As an example, the nodulation competition between S. medicae WSM419 wild-type and its isogenic nolR mutant was presented in a paper by Sugawara and Sadowsky (2014).
- The competitive ability for nodulation is quantified by nodule occupancy rates in seven inoculant mixtures, with the proposition of two strains ranging from 1:1,000 to 1,000:1.
Representative data
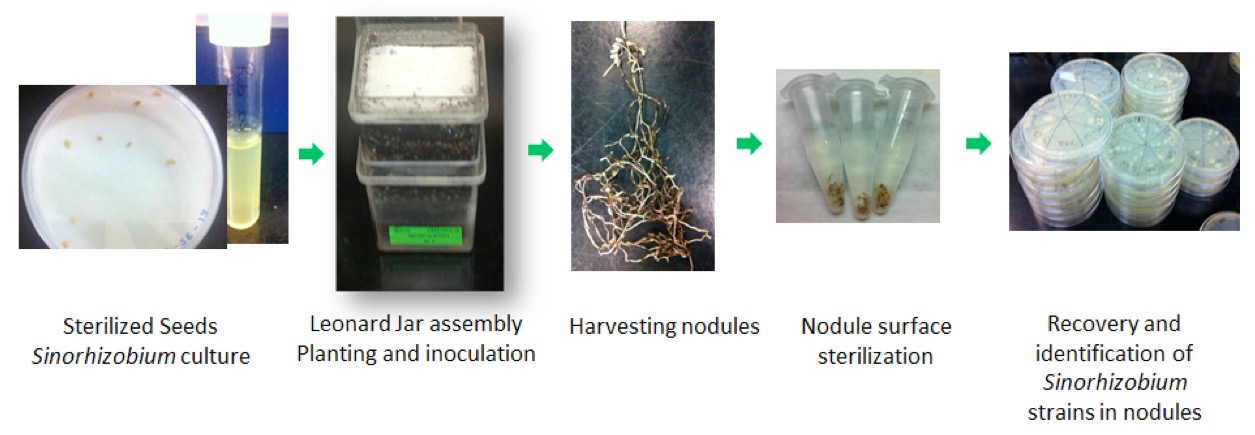

Figure 1. Experimental protocol for recovery and identification of Sinorhizobium meliloti strains in nodules of the host legume Medicago truncatula
Recipes
- TY medium
5 g tryptone
3 g yeast extract
1.3 g CaCl2.6H2O or 0.87g CaCl2.2H2O
Bring it to 1 L with distilled H2O and sterilize by autoclaving
For the plate, 1.5% agar is added
- N2-free nutrient solution (Bucciarelli et al., 2006)
KNO3
15 mM
Ca(NO3)2.4H2O
12.5 mM
Ca(H2PO4)2
1 mM
MgSO4.7H2O
1 mM
Fe-EDTA
0.01 mM
MnCl2
0.004 mM
H3BO3
0.02 mM
ZnSO4.7H2O
0.0004 mM
NaMoO4
0.0001 mM
CuSO4.5H2O
0.0001 mM
K2SO4
12.5 mM
CaSO4.2H2O
9 mM
- Sand
- Dissolve 1 g of paraffin into 100 ml chloroform (mix in the hood, takes time to dissolve).
- Pour dissolved paraffin on 2 kg white fine sand in a metal pan.
- Mix paraffin/chloroform thoroughly with sand and evaporate the chloroform overnight inside hood.
- Distribute paraffin coated sand into an autoclavable container and autoclave for one hour.
- Dissolve 1 g of paraffin into 100 ml chloroform (mix in the hood, takes time to dissolve).
Acknowledgments
This study was supported by grant 1237993 form The National Science Foundation.
References
- Amarger, N. and Lobreau, J. P. (1982). Quantitative study of nodulation competitiveness in Rhizobium strains. Appl Environ Microbiol 44(3): 583-588.
- Bucciarelli, B., Hanan, J., Palmquist, D. and Vance, C. P. (2006). A standardized method for analysis of Medicago truncatula phenotypic development. Plant Physiol 142(1): 207-219.
- Sugawara, M. and Sadowsky, M. J. (2014). Enhanced nodulation and nodule development by nolR mutants of Sinorhizobium medicae on specific Medicago host genotypes. Mol Plant Microbe Interact 27(4): 328-335.
- Toro, A. (1996). Nodulation competitiveness in the Rhizobium-legume symbiosis. World J Microbiol Biotechnol 12(2): 157-162.
- Triplett, E. W. and Sadowsky, M. J. (1992). Genetics of competition for nodulation of legumes. Annu Rev Microbiol 46: 399-428.
- Weir, B. S. (2012). The current taxonomy of rhizobia. NZ Rhizobia website.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Chun, C. L., Nelson, M. S. and Sadowksy, M. J. (2015). Quantitative Evaluation of Competitive Nodulation among Different Sinorhizobium Strains. Bio-protocol 5(15): e1555. DOI: 10.21769/BioProtoc.1555.
Category
Microbiology > Microbe-host interactions > Bacterium
Microbiology > Microbial cell biology > Cell isolation and culture
Plant Science > Plant metabolism > Nitrogen
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











