- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
In Gel Detection of Lipase Activity in Crude Plant Extracts (Olea europaea)
Published: Vol 5, Iss 8, Apr 20, 2015 DOI: 10.21769/BioProtoc.1444 Views: 8511
Reviewed by: Renate WeizbauerAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
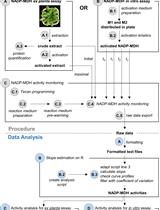
A Semi-throughput Procedure for Assaying Plant NADP-malate Dehydrogenase Activity Using a Plate Reader
Kevin Baudry and Emmanuelle Issakidis-Bourguet
Aug 20, 2023 1509 Views
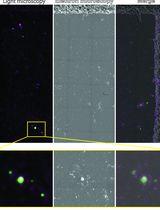
Sorghum bicolor Extracellular Vesicle Isolation, Labeling, and Correlative Light and Electron Microscopy
Deji Adekanye [...] Jeffrey L. Caplan
Oct 5, 2024 2149 Views

PhosphoLIMBO: An Easy and Efficient Protocol to Separate and Analyze Phospholipids by HPTLC From Plant Material
Louise Fougère [...] Yohann Boutté
Sep 5, 2025 1371 Views
Abstract
Here, we provide a detailed protocol describing a SDS-PAGE based procedure to assay in gel neutral lipase activity. Total protein extracts are separated by SDS-PAGE and gels are treated with lipase substrate-α-naphthyl palmitate. This long-chain fatty acid ester is hydrolysed by lipases present in the gel. The product resulting from this reaction can be then visualized in the gel as yellow-brownish activity bands. This relatively simple and effective method of lipase assay detection can be used for crude protein extracts from different plant tissues.
Keywords: LipaseMaterials and Reagents
- Homogenized plant tissues
- Liquid nitrogen
- 30% acrylamide stock (29: 1 acrylamide: bisacrylamide) (Bio-Rad Laboratories, catalog number: 161-0156 )
- TEMED (Bio-Rad Laboratories, catalog number: 161-0801 )
- Ammonium persulfate (Sigma-Aldrich, catalog number: A3678 )
- 2D Quant Kit (GE Healthcare, catalog number: 80-6483-56 )
- Trizma® base (Sigma-Aldrich, catalog number: T1503 )
- Bromphenol blue (Sigma-Aldrich, catalog number: B0126 )
- Glycerol
- Triton X-100 (Sigma-Aldrich, catalog number: T8532 )
- Na2HPO4 (Sigma-Aldrich, catalog number: S3264 )
- NaH2PO4 (Sigma-Aldrich, catalog number: 71505 )
- N,N-Dimethylformamide (Fluka, catalog number: 72438 )
- Fast blue B salt (Sigma-Aldrich, catalog number: D9805 )
- α-naphthyl palmitate (Sigma-Aldrich, catalog number: N9875 )
Note: α-naphthyl palmitate catalog number: N9875 is no longer available in Sigma-Aldrich, but is available in Santa Cruz Biotechnology, catalog number: CAS 15806-43-6. - Extraction buffer (see Recipes)
- 12% separating gel (see Recipes)
- 4% stacking gel (see Recipes)
- 2 x SDS sample buffer (see Recipes)
- 1 M sodium phosphate buffer (see Recipes)
- Developing solution (see Recipes)
Equipment
- Mortar and pestle
- Refrigerated centrifuge 5810R (Eppendorf, catalog number: 5810000.017 ) or similar, equipped with a rotor for 2 ml microcentrifuge tubes
- Magnetic stirrer
- 2 ml microcentrifuge tube
- Tubular glass vials
- Mini-PROTEAN® Tetra Cell (Bio-Rad Laboratories, catalog number: 165-8000 )
Software
- Quantity One software (Bio-Rad Laboratories)
Procedure
- Tissue homogenization and protein extraction
Note: Do not denature the sample at any time. Proteins must remain in their native, folded state.- Homogenize tissue (0.2 g) to a very fine powder in liquid nitrogen using a precooled mortar and pestle.
- Transfer the powder to a 2 ml microcentrifuge tube and add 1.5 ml of extraction buffer and mix well using vortex (see Note 3).
Note: If samples are used immediately or stored properly it is not necessary to use protease inhibitors. - Stir sample in tubular glass vials using a magnetic stirrer for 1 h at 4 °C.
- Centrifuge at 13,500 x g for 30 min at 4 °C.
- After centrifugation, collect the supernatant.
- Quantify extract and make aliquots.
Note: The protein concentration was measured using the 2D Quant Kit following the manufacturer’s instructions.
- Homogenize tissue (0.2 g) to a very fine powder in liquid nitrogen using a precooled mortar and pestle.
- SDS-PAGE
- Mix protein sample (approx. 25 μg per sample) with an equal volume of 2x SDS sample buffer. Do not heat at any time.
- Load protein sample and perform the SDS-PAGE on a 12% separation gel.
- Mix protein sample (approx. 25 μg per sample) with an equal volume of 2x SDS sample buffer. Do not heat at any time.
- In-gel detection of lipase activity
- Remove SDS from the polyacrylamide gels by washing them three times for 30 min each in a solution containing 0.05 M phosphate buffer (pH 7.0) and 2.5 % (v/v) Triton X-100.
- Incubate the gels for 30 min at 37 °C in a developing solution in the dark.
- Wash gels with distilled water three times for 10 min on a shaker.
- The gels can be stored in distilled water up to two months at 4 °C without a significant loss of staining intensity.
- To quantify the level of lipase activity, the Quantity One software can be used to measure the density of appearing brown bands.
- Remove SDS from the polyacrylamide gels by washing them three times for 30 min each in a solution containing 0.05 M phosphate buffer (pH 7.0) and 2.5 % (v/v) Triton X-100.
Recipes
- Extraction buffer
0.5 M sodium phosphate buffer (pH 7.0) - 12% separating gel
Add the following solutions (total volume: 10 ml)30% acrylamide/bisacrylamide 3 ml 1.5 M Tris-HCl (pH 8.8) 2.5 ml 10% SDS 100 μl dH2O 4.4 ml 10% ammonium persulfate 50 μl TEMED 10 μl - 4% stacking gel
Add the following solutions (total volume: 5 ml)30% acrylamide/bisacrylamide 0.5 ml 0.5 M Tris-HCl (pH 6.8) 1.25 ml 10% SDS 50 μl dH2O 3.2 ml 10% ammonium persulfate 25 μl TEMED 5 μl - 2 x SDS sample buffer
100 mM Tris-HCl (pH 6.8)
4% (w/v) SDS
0.2% (w/v) bromophenol blue
20% (v/v) glycerol - 1 M sodium phosphate buffer (pH 7.0)
57.7 ml 1 M Na2HPO4
42.3 ml 1 M NaH2PO4 - Developing solution
Mix 40 mg of α-naphthyl palmitate, prepared in 16 ml of N, N-Dimethylformamide with 80 mg of Fast blue BB salt in 144 ml of 0.1 M phosphate buffer (pH 7.0)
Acknowledgments
This work was supported by the Spanish Ministry of Science and Innovation (MICINN) (ERDF-cofinanced project AGL2008-00517) and the Junta de Andalucía (ERDF-cofinanced project P2010-CVI5767).
References
- Rejon, J. D., Zienkiewicz, A., Rodriguez-Garcia, M. I. and Castro, A. J. (2012). Profiling and functional classification of esterases in olive (Olea europaea) pollen during germination. Ann Bot 110(5): 1035-1045.
- Zienkiewicz, A., Zienkiewicz, K., Rejon, J. D., Alche Jde, D., Castro, A. J. and Rodriguez-Garcia, M. I. (2014). Olive seed protein bodies store degrading enzymes involved in mobilization of oil bodies. J Exp Bot 65(1): 103-115.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Zienkiewicz, A., Rejón, J. D., Zienkiewicz, K., Castro, A. J. and Rodríguez-García, M. I. (2015). In Gel Detection of Lipase Activity in Crude Plant Extracts (Olea europaea). Bio-protocol 5(8): e1444. DOI: 10.21769/BioProtoc.1444.
Category
Plant Science > Plant biochemistry > Lipid
Biochemistry > Lipid > Lipid-protein interaction
Biochemistry > Protein > Activity
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











