- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Airbrush Infiltration Method for Pseudomonas syringae Infection Assays in Soybean
Published: Vol 5, Iss 6, Mar 20, 2015 DOI: 10.21769/BioProtoc.1427 Views: 10536
Reviewed by: Arsalan DaudiAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
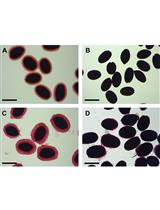
Analysis of Monosaccharides from Arabidopsis Seed Mucilage and Whole Seeds Using HPAEC-PAD
Gillian H. Dean [...] George W. Haughn
Dec 20, 2019 5777 Views
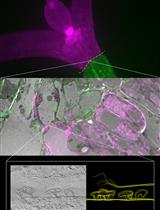
Targeting Ultrastructural Events at the Graft Interface of Arabidopsis thaliana by A Correlative Light Electron Microscopy Approach
Clément Chambaud [...] Lysiane Brocard
Jan 20, 2023 3010 Views
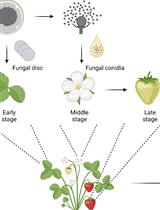
Botrytis cinerea in vivo Inoculation Assays for Early-, Middle- and Late-stage Strawberries
Piao Yang [...] Ye Xia
Oct 20, 2023 2829 Views
Abstract
We developed this protocol to assay the extent of proliferation of Pseudomonas syringae pv. glycinea in soybean leaves. This method specifically enables accurate pathogenesis assays of soybean plants at V2/V3 (2nd/3rd trifoliate) or higher stages of growth. The leaves of soybean plants at these growth stages are not amenable to bacterial infiltration using routine needleless syringe infiltration due to the high number of trichomes on these mature leaves. This method enables efficient infiltration of bacteria into the epidermal cells of mature leaves using a pressure pump.
Materials and Reagents
- Plant material
Soybean (Glycine max Merr.) plants of V2 (2nd trifoliate), V3 (3rd trifoliate), or higher stages of growth were used. Plants at VC (two leaf stage)/V1 (1st trifoliate) stages can also be used. Cultivars for various R loci: Rpg1-b (Harosoy), Rpg2 (Merit, Norchief), Rpg3 (Flambeau), Rpg4 (Flambeau), rpg (Essex). - Bacterial strains
- P. syringae pv. glycinea expressing avr gene of interest via the broad host range plasmids pDSK519 or pDSK600.
- P. syringae pv. glycinea expressing empty pDSK519/600 plasmids as control.
- P. syringae pv. glycinea expressing avr gene of interest via the broad host range plasmids pDSK519 or pDSK600.
- Media and buffers
- 10 mM MgCl2 (sterile)
- King’s B medium (plates and liquid media) (see Recipes)
- 10 mM MgCl2 (sterile)
- Other reagents
- Tryptone (Teknova, catalog number: T9012 )
- K2HPO4 (Fisher BioReagents, catalog number: BP363 )
- Glycerol (Affymetrix, catalog number: 16374 )
- Agar (Affymetrix, catalog number: 10654 )
- Silwet L-77 (Momentive, New Smyrna Beach)
- Antibiotics (Gold Biotechnology)
- Rifampicin (50 mg/ml) (R-120)
- Kanamycin (50 mg/ml) (K-120)
- Spectinomycin (100 mg/ml) (S-140)
- Sreptomycin (300 mg/ml) (S150)
- Rifampicin (50 mg/ml) (R-120)
- Tryptone (Teknova, catalog number: T9012 )
Equipment
- High-speed floor centrifuge (Thermo Fisher Scientific, Sorvall RC 6 plus)
- Spectrophotometer (Thermo Fisher Scientific, model: BioMate 5 )
- Pressure pump (Gast Manufacturing, model: DOA-P704-AA )
- Airbrush (Badger Air-brush, model: 250-2 )
- Test tubes (Thermo Fisher Scientific)
- Microcentrifuge tubes (Thermo Fisher Scientific)
- Glass rods (Thermo Fisher Scientific)
- Pellet pestles (Sigma-Aldrich)
- Cork borer (1 cm diameter, Thermo Fisher Scientific)
Procedure
- Bacterial growth and inoculum preparation
- Streak -80 °C stock of bacterial culture on King’s B agar with required antibiotics (Rifampacin and Kanamycin for pDSK519 plasmid; Rifampacin, Spectinomycin and Streptomycin for pDSK600 plasmid). Incubate the plate at 29 °C till bacterial colonies grow (24-48 h).
- Inoculate a single colony of P. syringae in 10 ml King’s B medium with required antibiotics (Rifampacin and Kanamycin for pDSK519 plasmid; Rifampacin, Spectinomycin and Streptomycin for pDSK600 plasmid).
- Grow 12-16 h at 29 °C on rotary shaker at 200 rpm until the culture reaches OD (A600) 0.8-1.2. Do not use culture at OD600 <0.8, or >1.2.
- Centrifuge culture at 3000 rpm for 10 min. Discard the supernatant and resuspend the bacterial pellet in 10 ml, 10 mM MgCl2 by gentle pipetting (do not vortex).
- Measure OD (A600) of the bacterial suspension (this should be the same OD as your culture from King’s B medium, 0.8-1.2) and dilute to 105 CFU (colony forming units)/ml using 10 mM MgCl2 (1.0 OD measurement approximately corresponds to 2 x 108 CFU/ml).
- Add Silwett L-77 to a final concentration of 0.01% to the bacterial suspension and mix gently. Use bacterial suspensions within 1 h after preparation.
- Streak -80 °C stock of bacterial culture on King’s B agar with required antibiotics (Rifampacin and Kanamycin for pDSK519 plasmid; Rifampacin, Spectinomycin and Streptomycin for pDSK600 plasmid). Incubate the plate at 29 °C till bacterial colonies grow (24-48 h).
- Soybean infiltration
- Attach appropriate ports of airbrush (Figure 1) to the pressure pump (40 psi or less, Figure 2) and beaker containing bacterial suspension using rubber tubing.
- Infiltrate soybean plants at V2 stage on the abaxial surface of the trifoliate leaves using the airbrush, while holding the leaf against a flat surface (such as a Petri plate) ensuring that the pressure does not damage the leaf during infiltration (see Video 1).
- Inoculate one trifoliate per plant, and at least 5 plants per bacterial strain. 5-8 ml of bacterial inoculum is sufficient to completely infiltrate one trifoliate.
- Mock inoculations should be done similarly 10 mM MgCl2 + 0.01% Silwett L-77 instead of the bacterial suspension.
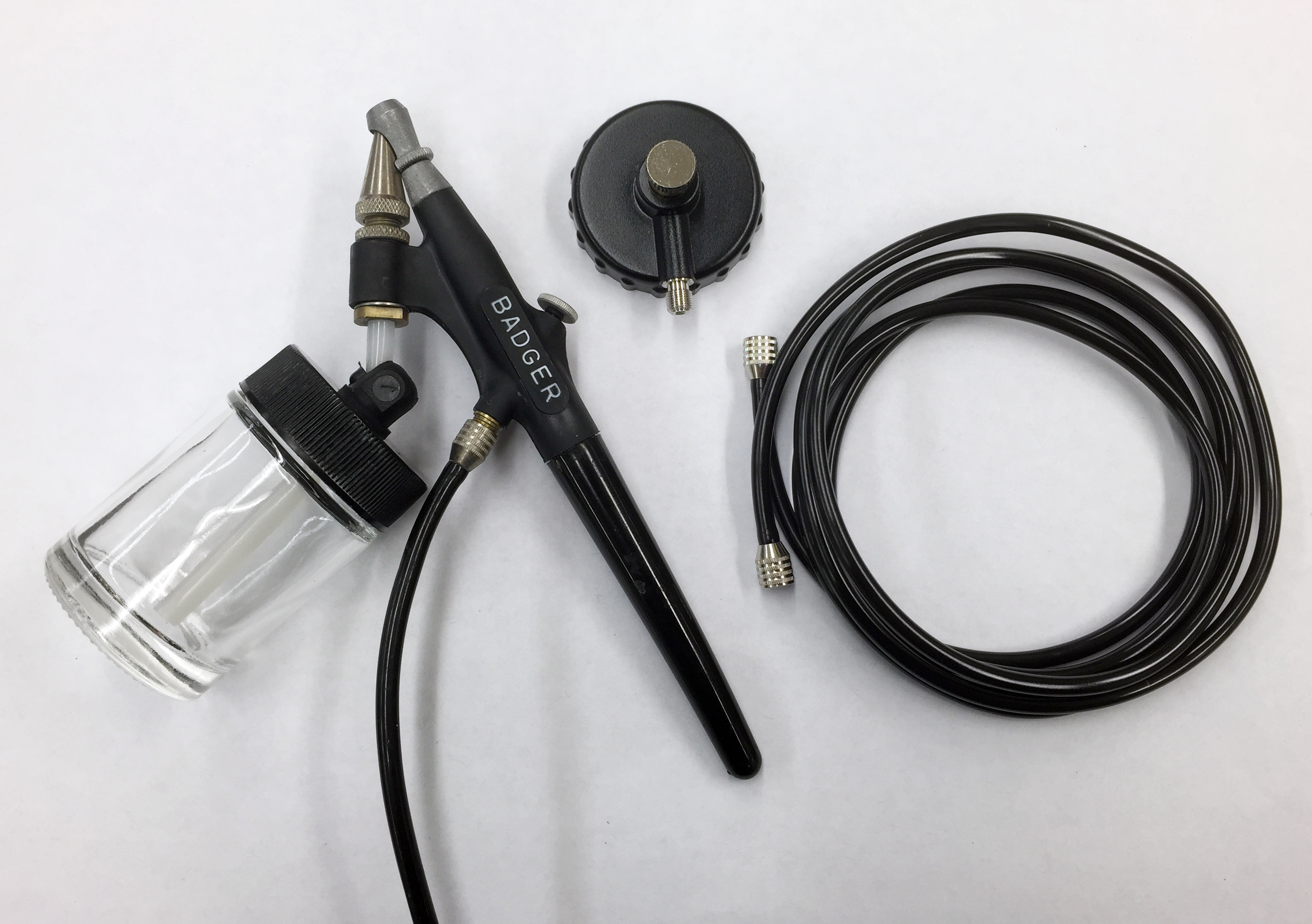

Figure 1. Image of airbrush used for bacterial infiltration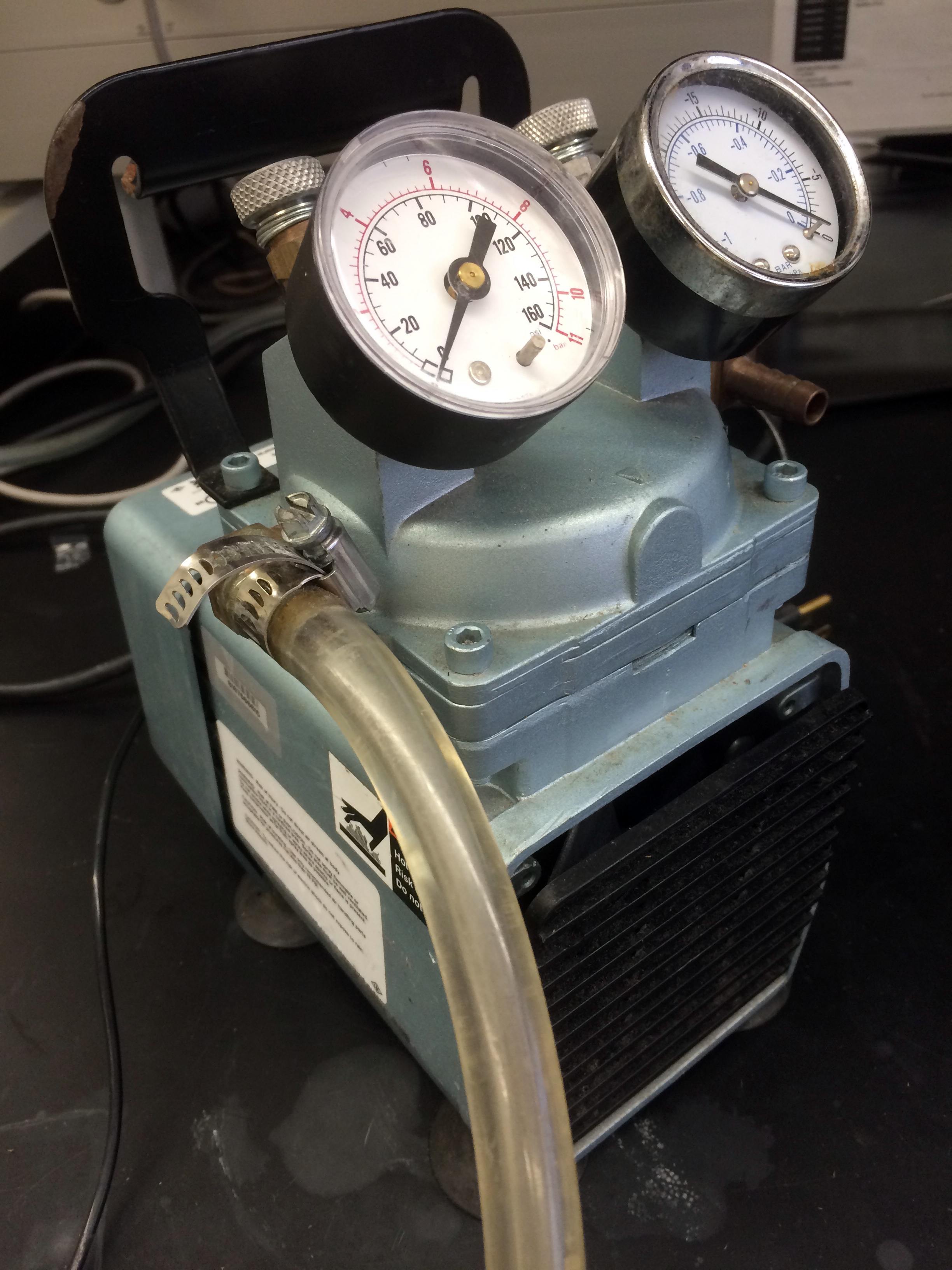
Figure 2. Image of pressure pump used for bacterial infiltration
- Attach appropriate ports of airbrush (Figure 1) to the pressure pump (40 psi or less, Figure 2) and beaker containing bacterial suspension using rubber tubing.
- Monitor bacterial proliferation
- Collect 3 leaf discs from inoculated leaves at 1 h post infiltration. The 1 h wait prevents erroneous bacterial numbers from any excess inoculum on the leaf surface. By 1 h post infiltration, the infiltrated leaves should no longer appear wet. Collect leaf discs using 1 cm cork borer from each plant. This is your 0 dpi sample.
- Homogenize leaf discs in 0.3 ml, 10 mM MgCl2 in a microcentrifuge tube manually using pellet pestle (Figure 3). For reproducibility and accurate bacterial count homogenize until no leaf pieces are visible. (Figure 4). Increase final volume to 1 ml after complete homogenization.
- Dilute 10x using 10 mM MgCl2 and plate 100 μl on King’s B medium (one sample per plate) using a glass spreader. Incubate plates at 29 °C till colonies grow (usually 24-48 h). Plate minimum 3 technical replicates per bacterial strain for every genotype. Technical repeats indicate independent leaf extracts from the same set of plants infected with the same bacterial culture.
- Count the number of bacterial colonies on entire plate, and adjust for dilution factor to determine total bacterial number. Plot bacterial counts as LOG10 values of CFU/unit leaf disc. It is important to adjust the dilution factor to obtain colonies in a countable range (10-300) depending upon the genotype of the plant. It is recommended to repeat the experiment using lower dilutions for plating if colony numbers are less than 10 or using higher dilutions when colony numbers are more than 300 per plate.
- Repeat steps 1-4 at 3, 4, 6 dpi (or other desired time points). Dilute homogenized tissue 5,000-10,000 times dilution for 3 dpi and later samples. Use minimum 4-5 technical replicates for these time points. Use 3-4 biological replicates (indicates independent infections using independent bacterial culture and set of plants) per bacterial strain and per plant genotype.
- Hypersensitive reaction related symptoms are visually detectable by 6-7 dpi (Figure 5).

Figure 3. Image of pellet pestle used for homogenization of leaf material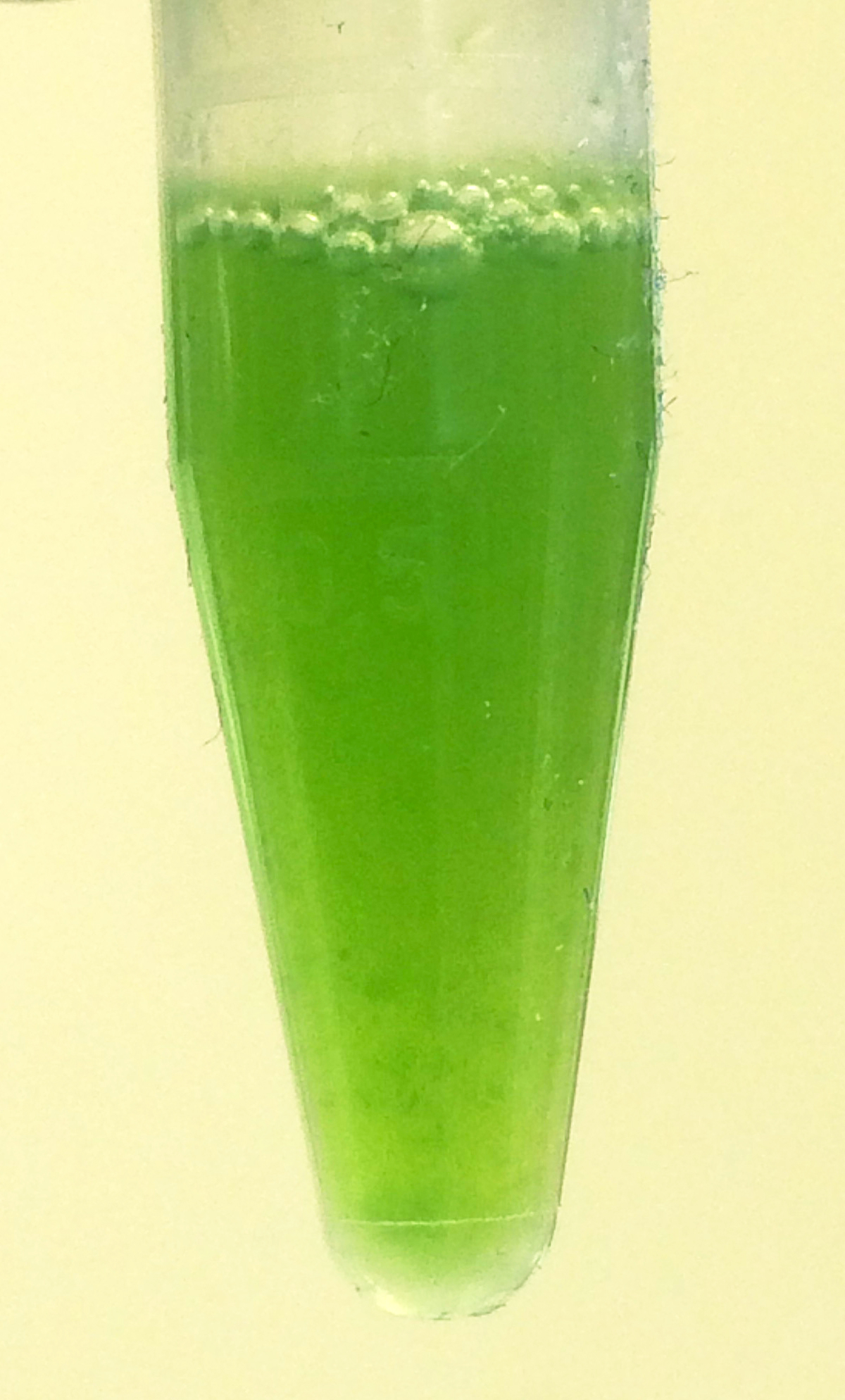
Figure 4. Image of homogenate of infected leaf material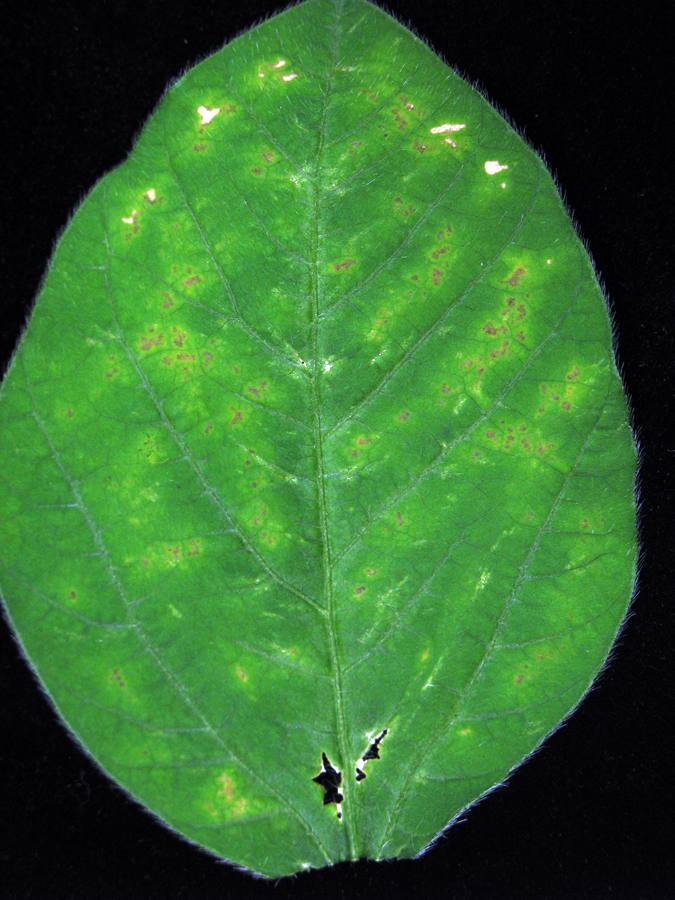
Figure 5. Morphological symptoms associated with the hypersensitive response of P. syringae pv. glycinea infected soybean leaves
- Collect 3 leaf discs from inoculated leaves at 1 h post infiltration. The 1 h wait prevents erroneous bacterial numbers from any excess inoculum on the leaf surface. By 1 h post infiltration, the infiltrated leaves should no longer appear wet. Collect leaf discs using 1 cm cork borer from each plant. This is your 0 dpi sample.
Representative data
- The protocol has high reproducibility (>90%) in our hands. We normally use at least four biological replicates per genotype for each experiment, and each infection experiment is conducted with a minimum three technical replicates. The standard error for colony count ranges between LOG10 value of 0.01-0.2.
- See Selote and Kachroo (2010); Selote et al. (2013) and (2014); Wang et al. (2014) for typical bacterial counts observed on various cultivars.
Notes
- Care should be taken to avoid damaging the leaves during the inoculation process.
- It is best to use fully expanded leaves for infection and to inoculate all leaves of one trifoliate per plant.
- Spraying the bacterial suspension over the leaf surface is generally sufficient. It is not necessary to saturate leaves with the bacterial suspension as excessive pressure during infiltration can result in wilting of the leaf.
Recipes
- King’s B medium (plates and liquid media)
20 g tryptone
1.5 g K2HPO4
10 ml glycerol
15 g agar, make up volume to 1 L with water
Autoclave, then add 5 ml of sterile MgSO4 (1 M)
Acknowledgments
This work was supported by funding from the United Soybean Board (project #1291) and the Kentucky Soybean Promotion Board. This protocol is modified from a previous method used by Keen et al. (1990).
References
- Fu, D. Q., Ghabrial, S. and Kachroo, A. (2009). GmRAR1 and GmSGT1 are required for basal, R gene-mediated and systemic acquired resistance in soybean. Mol Plant Microbe Interact 22(1): 86-95.
- Kachroo, A., Fu, D. Q., Havens, W., Navarre, D., Kachroo, P. and Ghabrial, S. A. (2008). An oleic acid-mediated pathway induces constitutive defense signaling and enhanced resistance to multiple pathogens in soybean. Mol Plant Microbe Interact 21(5): 564-575.
- Keen, N. T., Tamaki, S., Kobayashi, D., Gerhold, D., Stayton, M., Shen, H., Gold, S., Lorang, J., Thordal-Christensen, H., Dahlbeck, D. and Staskawicz, B. (1990). Bacteria expressing avirulence gene D produce a specific elicitor of the soybean hypersensitive reaction. Mol Plant Microbe Interact 3(2): 112–121.
- Selote, D. and Kachroo, A. (2010). RPG1-B-derived resistance to AvrB-expressing Pseudomonas syringae requires RIN4-like proteins in soybean. Plant Physiol 153(3): 1199-1211.
- Selote, D., Robin, G. P. and Kachroo, A. (2013). GmRIN4 protein family members function nonredundantly in soybean race-specific resistance against Pseudomonas syringae. New Phytol 197(4): 1225-1235.
- Selote, D., Shine, M. B., Robin, G. P. and Kachroo, A. (2014). Soybean NDR1-like proteins bind pathogen effectors and regulate resistance signaling. New Phytol 202(2): 485-498.
- Singh, A. K., Fu, D. Q., El-Habbak, M., Navarre, D., Ghabrial, S. and Kachroo, A. (2011). Silencing genes encoding omega-3 fatty acid desaturase alters seed size and accumulation of Bean pod mottle virus in soybean. Mol Plant Microbe Interact 24(4): 506-515.
- Wang, J., Shine, M. B., Gao, Q. M., Navarre, D., Jiang, W., Liu, C., Chen, Q., Hu, G. and Kachroo, A. (2014). Enhanced disease susceptibility1 mediates pathogen resistance and virulence function of a bacterial effector in soybean. Plant Physiol 165(3): 1269-1284.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Shine, M., Fu, D. and Kachroo, A. (2015). Airbrush Infiltration Method for Pseudomonas syringae Infection Assays in Soybean . Bio-protocol 5(6): e1427. DOI: 10.21769/BioProtoc.1427.
Category
Plant Science > Plant immunity > Disease bioassay
Plant Science > Plant immunity > Perception and signaling
Plant Science > Plant physiology > Tissue analysis
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link













