- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Detection of Phospho-KRAS by Electrophoretic Mobility Change in Human Cell Lines and in Tumor Samples from Nude Mice Grafts
(*contributed equally to this work) Published: Vol 5, Iss 6, Mar 20, 2015 DOI: 10.21769/BioProtoc.1421 Views: 10126
Reviewed by: HongLok LungThomas J. BartoshShalini Low-Nam

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
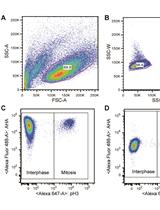
Measuring Protein Synthesis during Cell Cycle by Azidohomoalanine (AHA) Labeling and Flow Cytometric Analysis
Koshi Imami and Tomoharu Yasuda
Apr 20, 2019 9785 Views
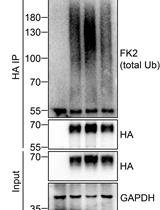
Analysis of the Ubiquitination and Phosphorylation of Vangl Proteins
Di Feng [...] Bo Gao
Oct 20, 2022 3464 Views
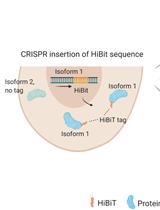
Isoform-specific, Semi-quantitative Determination of Highly Homologous Protein Levels via CRISPR-Cas9-mediated HiBiT Tagging
Kristina Seiler [...] Mario P. Tschan
Jul 20, 2023 2550 Views
Abstract
KRAS is the oncogene most frequently mutated in human solid tumors especially in pancreas, colon, small intestine, biliary tract and lung. We have recently demonstrated that oncogenic KRAS needs S181 phosphorylation to fully display its oncogenic features suggesting its inhibition as a therapeutic treatment against KRAS-driven tumors. Due to the importance to detect KRAS phosphorylation in human tumors and the absence of specific antibodies against phosphorylated KRAS, we developed a new protocol based on the Phos-tag SDS methodology to detect this post-translational modification for KRAS. Phos-tag is a molecule that binds specifically to phosphorylated proteins, decreasing their migration speed in SDS-PAGE and allowing its separation from the non-phosphorylated forms.
Materials and Reagents
- Human tumor cell lines
- Human tumor samples grown in immunodeficient mice from living cells or tissue origins
- Phos-tagTM acrylamide (Wako Chemicals GmbH, catalog number: 304-93521 , #AAL-107)
- λ Protein phosphatase (≥400,000 units/ml) (Calbiochem, catalog number: 539514-20KV )
- Anti- c- KRAS (clone Ab-1) mouse mAb (Calbiochem, catalog number: OP24 ) or any antibody against a putative tag-KRAS expressed (HA tag has been tested successfully)
- Protease inhibitors final concentrations
- Aprotinin: 150 nM (1 μg/ml) (Sigma-Aldrich, catalog number: A1153 ) (recommended stock solution 1 mg/ml)
- Leupeptin: 20 μM (10 μg/ml) (Sigma-Aldrich, catalog number: L2884 ) (recommended stock solution 1 mg/ml)
- Phenylmethanesulfonyl fluoride (PMSF): 1 mM (Sigma-Aldrich, catalog number: S6508 ) (recommended stock solution 100 mM)
- Aprotinin: 150 nM (1 μg/ml) (Sigma-Aldrich, catalog number: A1153 ) (recommended stock solution 1 mg/ml)
- Phosphatase inhibitors
- Sodium orthovanadate: 0.2 mM (Sigma-Aldrich, catalog number: S6508) (recommended stock solution 200 mM)
- Sodium fluoride: 5 mM (Sigma-Aldrich, catalog number: S7920 ) (recommended stock solution 1 M)
- Sodium orthovanadate: 0.2 mM (Sigma-Aldrich, catalog number: S6508) (recommended stock solution 200 mM)
- IGEPAL CA-630 (Sigma-Aldrich, catalog number: I8896 )
- Dynabeads Protein G for immunoprecipitation (Novex by Life Technologies, catalog number: 10003D )
- Temed
- 13% ammonium persulfate (APS)
- MnCl2 stock solutions 10 mM and 25 mM
- λ phosphatase lysis buffer (see Recipes)
- 3x SDS sample buffer (see Recipes)
- Electrophoresis running buffer (see Recipes)
- Transfer buffer (see Recipes)
- 5 mM Phos-tag stock solution (see Recipes)
- 20x PBS (see Recipes)
- Solution 1 for SDS-PAGE (see Recipes)
- Solution 2 for SDS-PAGE (see Recipes)
- Solution 3 for SDS-PAGE (see Recipes)
- Resolving composition of a 12% SDS-PAGE mini gel (1.5 mm thickness) for Phos-tag (see Recipes)
- Stacking composition of a 12% SDS-PAGE mini gel (1.5 mm thickness) (see Recipes)
Equipment
- Tissue grinder (Dounce) (1 ml) (Wheaton, catalog number: 357538 )
- Magnet DynaMagTM-2 (Life Technologies, catalog number: 12321D )
- Tube Rotator (Bibby Scientific Ltd., Stuart SB2)
Procedure
- Sample lysis and λ Phosphatase treatment
For all samples
- For each sample analysed (sample A), a negative control (sample B) treated with λ Phosphatase will be assayed concurrently. Prepare two tubes with ice-cold λ Phosphatase Lysis Buffer, freshly prepared, containing protease and phosphatase inhibitors (A) or only protease inhibitors (B).
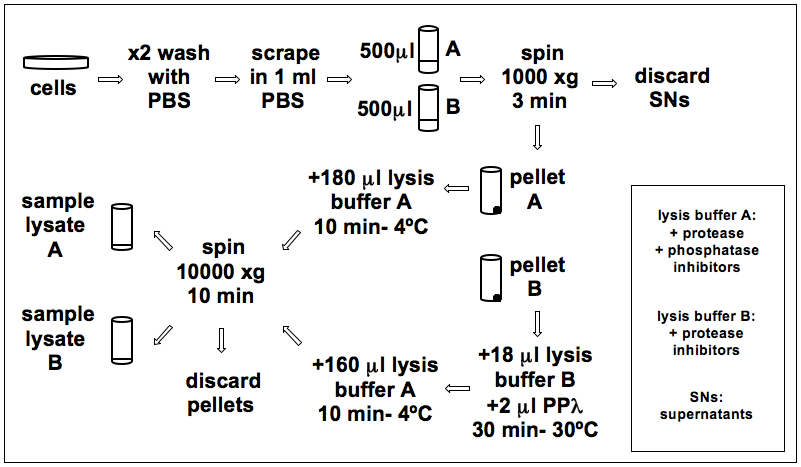
Figure 1. The flowchart of sample lysis and λ Phosphatase treatment
For cell lines
- Wash cells from a 10 cm dish (80% confluence) with 10 ml ice-cold phosphate buffered saline (PBS) twice.
- Scrape cells with 1 ml ice-cold PBS, collect in two 1.5 ml microfuge tubes A and B (500 μl each) and spin for 3 min at 1,000 x g at 4 °C. Discard supernatant.
- Add 18 μl ice-cold λ phosphatase lysis buffer containing only protease inhibitors to the pellet B and next 2 μl of λ Protein Phosphatase (40 units/ μl final concentration). Incubate at 30 °C for 30 min in a dry block.
- Add 180 μl ice-cold λ Phosphatase Lysis Buffer containing protease and phosphatase inhibitors to the pellet A. Resuspend the pellet by pipetting and after 10 min on ice, spin for 10 min at 10,000 x g at 4 °C. Transfer the supernatant to a new 1.5 ml tube (sample lysate A).
- Add 160 μl cold λ phosphatase lysis buffer containing protease and phosphatase inhibitors to the pellet B. Resuspend the pellet and after 10 min on ice, spin for 10 min at 10,000 x g at 4 °C.Transfer the supernatant to a new 1.5 ml tube (sample lysate B).
- Analyse protein content of the supernatant obtained from steps 4-5 by Lowry method.
For tumor samples
- Take two fragments of the same tumor of approximately 0.1 g each. Leave each fragment of tumor on a 35 mm dish on ice in the presence of 0.4 ml of cold λ phosphatase lysis buffer containing protease and phosphatase inhibitors (dish sample A) or only protease inhibitors (dish sample B), but not DTT and MnCl2 in any case (will be added in step 5). Mince the tumors in the smallest possible pieces with a scalpel and scissors and collect all volume, containing minced tissue and lysis buffer, in 1.5 ml microfuge tubes.
Note: If the sample is a frozen tumor, mincing of the tumor in the presence of the buffer has to be done while the tumor is thawing.
- Homogenize the samples with a tissue grinder on ice until no clumps remain in the suspension.
- Spin the tubes at 10,000 x g at 4 °C for 5 min to discard unlysed tissue and collect supernatants in new tubes. The volume of samples obtained is about 300 μl.
- If the tumors are mice xenografts, it is convenient to clarify the samples with Dynabeads protein G in order to eliminate putative mouse Igs that might be infiltrating the tumor thus interfering with phospho-KRAS detection:
- Transfer 30 μl of Dynabeads (previously vortexed) to a tube.
- Separate the Dynabeads from the solution with Magnet DynaMagTM, discard the supernatant, remove the tube from the magnet and wash dynabeads with λ phosphatase lysis buffer (1 ml, inhibitors free).
- Separate the Dynabeads from the buffer with a magnet, discard the buffer, and remove the tube from the magnet.
- Incubate the Dynabeads with the samples for 20 min in a tube rotator to allow gentle mixing at 4 °C. The volume of samples is about 300 μl (volume of supernatant obtained in step 3).
- Separate the Dynabeads from the samples with the magnet and transfer the clarified samples A and B to a new tube. Measure the volume of clarified samples.
- Transfer 30 μl of Dynabeads (previously vortexed) to a tube.
- Add 5 mM DTT (from stock 1 M DTT) and 2 mM MnCl2 (from stock 25 mM MnCl2) to clarified samples A and B. Mix gently.
- Take 40 μl of sample B to a new tube, add 4 μl of λ Protein phosphatase and incubate at 30 °C for 30 min. The rest of sample B is discarded.
- Add phosphatase inhibitors to sample B: 0.2 μl of 1 M sodium fluoride and 0.4 μl of 20 mM sodium orthovanadate (or 0.04 μl of 200 mM sodium orthovanadate) should be added to the 40 μl of sample B to achieve the 5 mM sodium fluoride and 0.2 mM sodium orthovanadate final concentrations that are stated in the Materials section.
- Analyse protein content of sample A from step 5 and sample B from step A6 by the Lowry method.
- For each sample analysed (sample A), a negative control (sample B) treated with λ Phosphatase will be assayed concurrently. Prepare two tubes with ice-cold λ Phosphatase Lysis Buffer, freshly prepared, containing protease and phosphatase inhibitors (A) or only protease inhibitors (B).
- Sample preparation and Phos-tag SDS and transfer
- Transfer 20 μg of each sample in 10 μl of final volume with λ phosphatase lysis buffer and add 5 μl of sample buffer 3x. Boil the samples for 1 min in a 1.5 ml tube.
- Load the samples into a 12% SDS-PAGE mini gel supplementing the resolving gel with 100 μM Phos-tag and 100 μM MnCl2.
- Run the gel 12 h to 15 h at 5 mA/gel at 4 °C.
- Soak the gel twice with general transfer buffer containing 1 mM EDTA for 20 min and once with EDTA-free general transfer buffer for 10 min.
- Transfer proteins from the gel onto PVDF for 3 h at 70 V.
- Perform Western blot with the appropriate antibodies: anti- K-RAS antibodies or any antibody against a putative tag-KRAS expressed.
- Transfer 20 μg of each sample in 10 μl of final volume with λ phosphatase lysis buffer and add 5 μl of sample buffer 3x. Boil the samples for 1 min in a 1.5 ml tube.
Representative data
A representative example of Phos-tag SDS-PAGE followed by Western blot using anti-KRAS antibodies, of samples from tumor development in mice after subcutaneously injecting NIH3T3 cells expressing phosphorylatable oncogenic KRAS (KRASG12V S181) is shown in Figure 2 below.

Figure 2. Representative data. 1. K-RasG12V S181 tumor. Non incubated sample; 2. K-RasG12V S181 tumor. Non incubated sample; 3. K-RasG12V S181 tumor. Incubated at 4 °C without phosphatase λ; 4. K-RasG12V S181 tumor. Incubated at 30 °C without phosphatase λ; 5. K-RasG12V S181 tumor. Incubated at 30 °C with phosphatase λ.
Notes
- It is important to start from samples with high protein concentration (at least 3 μg/μl) in order to obtain clear results.
- When loading the gel avoid using protein molecular weight markers because mobility is not proportional to molecular weight in the presence of Phos-tag reagent.
- When loading the gel, leave first and last lanes sample-free to avoid gel distortion.
- KRAS phosphorylated band is always retarded compared with unphosphorylated band, but the distance between bands may vary depending on the sample (endogenous KRAS, tag-KRAS expressed in cells or in tumors, etc.). In order to explain correctly the results a control sample treated with λ Phosphatase must be assayed in each experiment at the same time.
- Phostag-PAGE gels are quite brittle compared to the normal PAGE gels. Be extra careful when manipulating them.
- Load freshly-made samples whenever possible avoiding long-term reloading of the sample. Phospho-KRAS-Phostag complex seems to be sensitive to long-term storage as P-KRAS band would eventually disappear after some time.
- Both gel-running conditions and Phos-tag concentration may need to be optimized for each cell line, tumor type, treatment, etc.
- It is advisable to run the samples in a usual SDS-PAGE gel (no Phos-tag added) in the same electrophoresis chamber concurrently with the Phos-tag gel since the dye front is difficult to see and follow in the Phos-tag gels.
Recipes
- λ phosphatase lysis buffer
50 mM Tris-HCl (pH 8)
150 mM NaCl
2 mM EDTA
10% glycerol
1% IGEPAL CA-630
dH2O
Add fresh:
5 mM DTT from stock 1 M DTT
2 mM MnCl2 from stock 25 mM MnCl2
Protease and phosphatase inhibitors (for stocks See Materials and Reagents)
To prepare 5 ml λ phosphatase lysis buffer containing protease and phosphatase inhibitors:
5 μl aprotinin
50 μl leupeptin
50 μl PMSF
5 μl sodium orthovanadate
25 μl sodium fluoride
- 3x SDS sample buffer
0.5 M Tris-HCl (pH 6.8)
1.5 mg Bromophenol blue
0.60 g SDS
3 ml glycerol
3.9 ml mercaptoethanol
Add dH2O to 10 ml
- Electrophoresis running buffer
25 mM Tris-HCl (pH 8.3)
192 mM Glycine
0.1% SDS
- Transfer buffer
25 mM Tris-HCl (pH 8.3)
192 mM glycine
0.02 % SDS
20 % ethanol
- 5 mM Phos-tag stock solution
Resuspend the oily product, Phos-tagTM acrylamide (10 mg) placed in a small plastic tube in 0.10 ml methanol.
Dilute the solution with 3.2 ml distilled water by pipetting
Store the solution in the 2-ml microtubes at 4 °C in the dark
- 20x PBS (pH 7.2)
170 g NaCl
21.5 g Na2HPO4·2H2O
7.0 g NaH2PO4·H2O
Add dH2O to 1,000 ml
For PBS (1x)
250 ml PBS (20x)
4,750 ml dH2O
- Solution 1 for SDS-PAGE
Tris-HCl 0.75 M (pH 8.8)
SDS 0.2 %
- Solution 2 for SDS-PAGE
30% acrylamide
0.8 % bis acrylamide
- Solution 3 for SDS-PAGE
Tris-HCl 0.25 M (pH 6.8)
SDS 0.2%
- Resolving composition of a 12% SDS-PAGE mini gel (1.5 mm thickness) for Phos-tag SDS
5 ml solution 1
4 ml solution 2
1 ml dH2O
200 μl Phos-tag stock solution 5 mM
100 μl MnCl2 stock solution 10 mM
14 μl Temed
50 μl APS 13%
- Stacking composition of a 12% SDS-PAGE mini gel (1.5 mm thickness)
0.36 ml solution 2
1.5 ml solution 3
1.2 ml dH2O
7.5 μl Temed
30 μl APS 13%
Acknowledgments
We are grateful to José Lozano, Associate Professor at Universidad de Málaga (Spain) and to Maribel Geli, Senior Research Scientist at the Molecular Biology Institute of Barcelona (ibmb)-CSIC for providing useful comments during the setting up of the conditions. This technique is part of a study supported by MICINN-Spain (SAF2010-20712) and RTICC (MINECO-Spain; groups RD12/0036/0049, RD12/0036/0031, and RD12/ 0036/0034). C. Barceló and N. Paco are recipients of predoctoral fellowships from MEC-Spain and Generalitat de Catalunya, respectively.
References
- Barceló, C., Etchin, J., Mansour, M. R., Sanda, T., Ginesta, M. M., Sanchez-Arevalo Lobo, V. J., Real, F. X., Capella, G., Estanyol, J. M., Jaumot, M., Look, A. T. and Agell, N. (2014). Ribonucleoprotein HNRNPA2B1 interacts with and regulates oncogenic KRAS in pancreatic ductal adenocarcinoma cells. Gastroenterology 147(4): 882-892 e888.
- Barceló, C., Paco, N., Morell, M., Alvarez-Moya, B., Bota-Rabassedas, N., Jaumot, M., Vilardell, F., Capella, G. and Agell, N. (2014). Phosphorylation at Ser-181 of oncogenic KRAS is required for tumor growth. Cancer Res 74(4): 1190-1199.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Barceló, C., Paco, N., Cabot, D., Garrido, E., Agell, N. and Jaumot, M. (2015). Detection of Phospho-KRAS by Electrophoretic Mobility Change in Human Cell Lines and in Tumor Samples from Nude Mice Grafts. Bio-protocol 5(6): e1421. DOI: 10.21769/BioProtoc.1421.
Category
Cancer Biology > General technique > Biochemical assays > Protein analysis
Molecular Biology > Protein > Detection
Biochemistry > Protein > Activity
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









