- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Measurement of Cellular Redox in Pollen with Redox-Sensitive GFP (roGFP) Using Live Cell Imaging
Published: Vol 5, Iss 5, Mar 5, 2015 DOI: 10.21769/BioProtoc.1414 Views: 10869
Reviewed by: Ru ZhangElias BassilAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
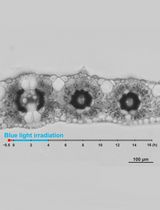
Live Leaf-Section Imaging for Visualizing Intracellular Chloroplast Movement and Analyzing Cell–Cell Interactions
Yuta Kato [...] Mitsutaka Taniguchi
Aug 5, 2025 2377 Views

Live-Cell Monitoring of Piecemeal Chloroplast Autophagy
Masanori Izumi [...] Shinya Hagihara
Nov 5, 2025 1723 Views
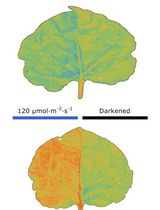
Chloroplast Movement Imaging Under Different Light Regimes With a Hyperspectral Camera
Paweł Hermanowicz [...] Justyna Łabuz
Dec 20, 2025 792 Views
Abstract
Redox homeostasis is a fundamental property of living cells and responds actively to both cellular metabolism and external stimulus. The development of redox-sensitive GFP (roGFP) enables dynamic monitoring of changes in cellular redox poise (Hanson et al., 2014). When excited alternatively at 405 nm and 488 nm, these probes exhibit significant opposing shifts at the emission spectra (505-530 nm), which enables ratiometric measurement of relative redox values. A more oxidized environment results in a higher 405/488 ratio. Previously, successful application of roGFPs in leaf epidermis or root cells has been reported. Here we provide a protocol describing the application of roGFP1 imaging in growing pollen tubes by confocal laser scanning microscopy.
Keywords: Pollen tubeMaterials and Reagents
- Transgenic tomato or tobacco pollen expressing roGFP1 under the control of the pollen-specific promoter LAT52
- 40% polyethylene glycol (molecular weight 4,000) (Sigma-Aldrich, catalog number: 81242 )
- Pollen germination medium (see Recipes)
Equipment
- Biovortexer (Bio Spec Products, catalog number: 1083 )
- Orbital shaker (Kylin-Bell Lab Instruments, model: TS-2 )
- 6 or 24 well cell culture plate
- Microscope slides and cover slips
- Confocal microscope (OLYMPUS, model: FV1000) or other microscopes equipped with both 405 nm and 488 nm laser lines
Software
- Olympus Fluoview version 3.0a
Note: ImageJ (version 1.49 g, Rasband, NIH, USA) can also be used to measure the intensities of confocal images.
Procedure
The method consists of harvesting pollen grains, incubating them in liquid germination medium, performing live cell imaging and making ratiometric analysis. Pollen will start to germinate at about 30 min. Then ratiometric imaging using a confocal microscope is performed. This can be achieved in a few minutes. Ratiometric image analysis can be performed at a later time point.
- In vitro pollen tube germination
- 2-10 mg mature pollen of tomato or tobacco is obtained by vibrating anthers of open flowers with a biovortexer for about 5 seconds each (Figure 1 A-C).
Note: Plant growth conditions affect pollen quality, especially for tomato. For example, appropriate temperature and enough sunshine are important factors required for proper pollen development. Moreover, infection with pathogens usually results in poor pollen yield and quality. Generally speaking, newly opened flowers contain pollen with greater vitality. - Dilute the pollen with pollen germination medium to a final concentration of 0.5-2 mg/ml. Transfer them to 6 or 24 well cell culture plates and incubate at 25-28 °C on an orbital shaker. Tomato and tobacco pollen are rotated horizontally at 60 and 140 rpm, respectively (Figure 1D). Pollen grains hydrate immediately and germinate at around 30 min after incubation. It is recommended to observe pollen tubes within 8 hours while they maintain normal growth.
Note: Temperature is critical for in vitro pollen tube growth and has a nonnegligible impact on the redox potential of growing pollen tubes. It is important to keep the pollen medium at a constant temperature throughout the experiment.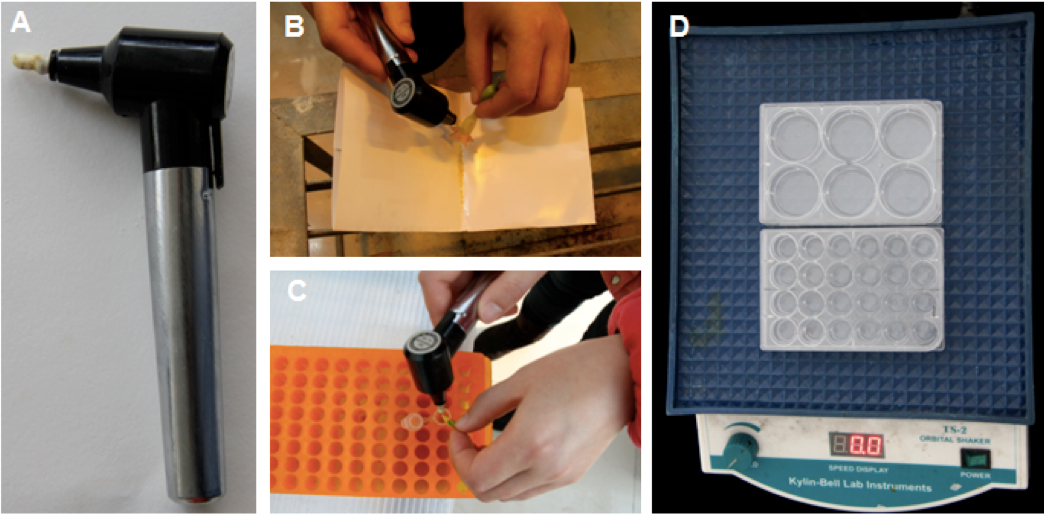
Figure 1. Pollen collection and liquid germination assay. A. An image of a biovortexer. B-C. Mature pollen of tobacco (B) or tomato (C) was collected by vibrating using a biovortexer. D. Pollen in germination medium was transferred to a 6 well culture plate (~2 ml/well) or a 24 well culture plate (0.5 ml/well) and germinated on an orbital shaker under fixed room temperature.
- 2-10 mg mature pollen of tomato or tobacco is obtained by vibrating anthers of open flowers with a biovortexer for about 5 seconds each (Figure 1 A-C).
- Microscope settings
Detailed parameters for an Olympus FV1000 are listed below. Other instruments need to be set to similar parameters.- Chose the “multi-track mode”. In track 1, the excitation wavelength is 405 nm. In track 2, the excitation wavelength is 488 nm. Emission for both tracks is collected with a band-pass filter of 505-530 nm. A transmission image can be collected synchronously in either track 1 or track 2.
- Setting options.
Scan mode XY Scan direction One way Image size 800 * 800 Bits/pixel 12 Bits Sampling speed 4.0 μs/Pixel C. A. 80-120 μm Integration type Line Kalman Integration count 4
Note: The images should be taken in line mode to guarantee correct ratiometric analysis (Meyer and Brach, 2009). Other settings, including laser power, gain setting, sampling speed, pinhole size (C. A.), integration count, can be optimized empirically, but only images taken with identical settings are compared. In our study, Images were acquired with a 20x lens (UPLSAPO; NA0.75) with a bit depth of 212. Calibration of roGFP can be done to determine the absolute value of the redox potential and to see the redox dependent change dynamics. While we hadn’t attempt to calibrate the roGFP intensity in Huang et al. (2014), readers can refer to Meyer and Brach (2009).
- Chose the “multi-track mode”. In track 1, the excitation wavelength is 405 nm. In track 2, the excitation wavelength is 488 nm. Emission for both tracks is collected with a band-pass filter of 505-530 nm. A transmission image can be collected synchronously in either track 1 or track 2.
- Redox-Sensitive GFP Imaging
- Hydrating pollen grain or growing pollen tubes can readily be imaged with confocal microscope.
Note: The cellular redox states in germinating pollen or growing pollen tubes at different time points are highly dynamic. It is important to compare two successive samples within a short periods of time (i.e., within 0.5 h). - Add about 40 μl sample on a microscope slide, cover the sample softly with a cover slip and leave them for about 2 min.
- Place the slide on the microscope. Use the ‘fast’ scanning in both tracks to adjust the laser intensities and gain settings for roGFP-expressing pollen tubes. Avoid images that are too dim or overexposed. Do not change the laser intensity and gain setting once they are determined.
- Start collecting images.
- Hydrating pollen grain or growing pollen tubes can readily be imaged with confocal microscope.
- Image processing and ratiometric analysis
- Open the combined images with Olympus Fluoview version 3.0a.
- Click “Ratio/Concentration” tool. Set appropriate ratio range and dividing the 405 nm excitation image (Figure 2B) by the 488 excitation image (Figure 2C) on a pixel-by-pixel basis. A resultant image (12 bit) and the color bar are shown in Figure 1D.
- Select the “Hi-Lo” pseudocolor scheme for both 405 and 488 channels in the lookup-table (LUT settings). In this color scheme, dim pixels are displayed in blue (Figure 2I, ROI3) while overexposed pixels are displayed in red (Figure 2G, ROI2).
- Draw a region-of-interest (ROI) for ratiometric analysis using the ROI toolbar. Avoid areas that are too dim (ROI3 in Figure 2I) or are overexposed (ROI2 in Figure 2G).
- Select “Intensity profile” tool and measure the signal intensities of ROI1 in both 405 and 488 channels. Save the intensity information to an Excel file.
- Mask out pixels with low signal intensities [a threshold of 100 was used in Huang et al. (2014)] before calculating the average intensities for each channel. Final ratio for statistic analysis is obtained by dividing the average intensity of 405 channel by that of 488 channel.
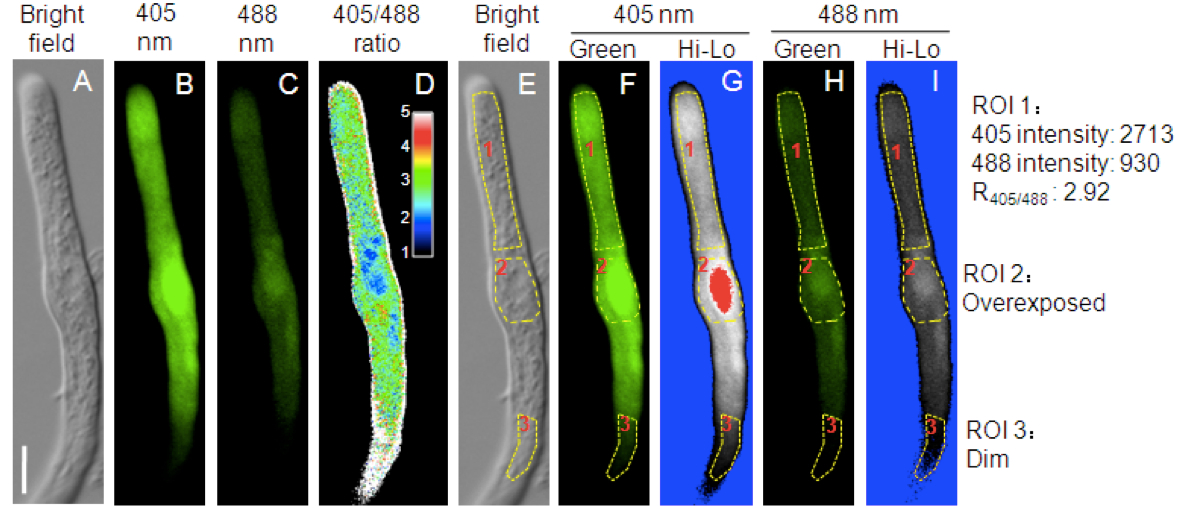
Figure 2. Redox-dependent fluorescence of roGFP1 in the cytosol of a representative tomato pollen tube. A. Bright field. B. The 505-530 nm emission signal after 405 nm excitation. C. The 505-530 nm emission signal after 488 nm excitation. D. The 405/488 ratio of roGFP1 fluorescence. The inset showed the color scale for the ratio values. E. Three ROIs (Region of Interest) on the bright field image. F-I. Fluorescent images pseudo colored in green F-H or in the “Hi-Lo” scheme G-I. Bar = 10 μm
- Open the combined images with Olympus Fluoview version 3.0a.
Recipes
- Pollen germination medium
Pollen germination medium should be prepared freshly every time from stock solutions listed below. All stock solutions can be autoclaved and stored at 4 °C for about half a year. It is important to let the germination medium to warm up to room temperature before using.Stock solution For 14 ml PGM Final concentration 40% polyethylene glycol 8.4 ml 24% (w/v) 50% sucrose 700 µl 2.5% (w/v) 1 M MES-KOH (PH 6.0) 280 µl 20 mM 1% boric acid 140 µl 1.6 mM 2% MgSO4.7H2O 140 µl 0.8 mM 1% KCl 140 µl 1.3 mM 1 M Ca(NO3)2 42 µl 3 mM H2O 4.2 ml —
Acknowledgments
The roGFP1 was kindly provided by Dr. James Remington. This work was supported by the Natural Science Foundation of China (Grant 30970266 to Dong Zhang and Grant 31170291 to W.-H.T.) and the Ministry of Science and Technology of China (Grant 2012AA10A302 to W. -H. T.). We thank Xiao-Shu Gao for help with roGFP imaging and Yu-Jie Li for taking photographs.
References
- Hanson, G. T., Aggeler, R., Oglesbee, D., Cannon, M., Capaldi, R. A., Tsien, R. Y. and Remington, S. J. (2004). Investigating mitochondrial redox potential with redox-sensitive green fluorescent protein indicators. J Biol Chem 279(13): 13044-13053.
- Huang, W. J., Liu, H. K., McCormick, S. and Tang, W. H. (2014). Tomato pistil factor STIG1 promotes in vivo pollen tube growth by binding to phosphatidylinositol 3-Phosphate and the extracellular domain of the pollen receptor kinase LePRK2. Plant Cell 26(6): 2505-2523.
- Meyer, A. J. and Brach, T. (2009). Dynamic redox measurements with redox-sensitive GFP in plants by confocal laser scanning microscopy. Methods Mol Biol 479: 93-107.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Huang, W. and Tang, W. (2015). Measurement of Cellular Redox in Pollen with Redox-Sensitive GFP (roGFP) Using Live Cell Imaging. Bio-protocol 5(5): e1414. DOI: 10.21769/BioProtoc.1414.
Category
Plant Science > Plant biochemistry > Other compound
Plant Science > Plant cell biology > Cell imaging
Cell Biology > Cell imaging > Fluorescence
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










