- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Determination of Protein-DNA (ZMYND11-DNA) Interaction by a Label-Free Biolayer Interferometry Assay
Published: Vol 5, Iss 4, Feb 20, 2015 DOI: 10.21769/BioProtoc.1402 Views: 13569
Reviewed by: Anonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
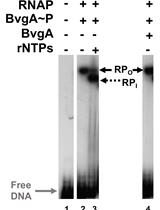
Combining Gel Retardation and Footprinting to Determine Protein-DNA Interactions of Specific and/or Less Stable Complexes
Meng-Lun Hsieh [...] Deborah M. Hinton
Dec 5, 2020 3585 Views
A Gel-Based Assay for Probing Protein Translocation on dsDNA
Christiane Brugger and Alexandra M. Deaconescu
Jul 20, 2021 4337 Views
Abstract
This protocol describes a robust technique for the measurement of ZMYND11-DNA interaction by a label-free Biolayer Interferometry (BLI). ZMYND11 is a novel histone reader protein that specifically recognizes H3.3K36me3 via its tandem Bromodomain, zinc-finger and PWWP domain (BP). ZMYND11 links the histone-variant-mediated transcription elongation control to tumour suppression and may therefore represent a novel class of drug targets. Like other PWWP domains, ZMYND11 PWWP domain shows highly positively charged surface and interacts with DNA. Previously reported methods include NMR, FP or EMSA. Biolayer interferometry (BLI) is an emerging technology for analyzing all kinds of biomolecular interactions, such as protein-protein and protein-DNA binding. BLI allows for the real time monitoring of the interactions between biomolecules without the need for reagents with enzymatic, fluorescent, or radioactive labels. The technology is based upon the changes in interference pattern of light reflected from the surface of an optical fiber when materials bind to the tip of the fiber. The technique represents an alternative to technologies such as surface plasmon resonance, providing a simple platform that enables label-free monitoring of biomolecular interactions without the use of flow cells. Label-free biosensor methods provide information on binding, kinetics, concentration, and the affinity of an interaction.
Keywords: Biolayer interferometryMaterials and Reagents
- Purified proteins of interest untagged or with tags such as GST, Flag, Biotin or His tag, depending on which kind of sensor to use (refer to manufacturer’s manual)
Note: Here GST-tagged ZMYND11-BP protein is used (Wen et al., 2014).
- Two complementary single stranded DNA (Life Technologies, InvitrogenTM)
- MilliQ water
- 10 mM glycine (pH 1.0~2.0)
- BLI binding buffer (see Recipes)
- DNA solutions (see Recipes)
Equipment
- Anti-GST antibody-coated biosensor tray (Pall Corporation, catalog number: 18-5096 )
- 96-well black microtiter plate (Greiner Bio-One, catalog number: 655209 )
- 1.5 ml Eppendorf microfuge tubes (Axygen, catalog number: MCT150C )
- Octet RED96 (fortebio) (Pall Corporation, catalog number: 30-5048) (Wen et al., 2014)
Software
- Octet Data analysis 7.0
Procedure
- Getting started
- Turning on the system:
Turn on the computer.
Next, make sure the sensor and sample positions are clear inside the Octet and turn on the power supply.
Finally, launch the Octet Data Acquisition software. Octet will initialize and home the optics box.
Warming up the Octet.
The Octet should be turned an hour prior to an experiment being run to allow for the lamp to warm up.
- Setting up experiment.
Use 200 µl samples in each well.
Pre-wet the sensors in BLI binding buffer for 30 min (time may vary depending on sensor – consult product insert):
Place 200 µl BLI binding buffer in each of eight wells of a 96-well black microtiter plate Insert the biosensors tray onto the 96-well black microtiter plate (Figure 1).

Figure 1. Flowchart to show how to pre-wet the BLT biosensors
- Turning on the system:
- Assay
- Start by defining the number of conditions you need to test, including appropriate controls such as protein sample in buffer only, or DNA sample in buffer only. This will determine the total volume of protein and DNA needed to conduct the entire experiment. Each well is conducted in a total volume of 200 μl.
- Pre-wet the biosensors in BLI binding buffer for at least 30 min. Biolayer interferometry experiments were conducted using an Octet Red, manufactured by ForteBio (Figure 2). The instrument can measure the signal from eight sensors simultaneously.

Figure 2. The ForteBio Octet Red96 system (Left), biosensor tray (Middle) and resulting signal curves shown (Right)
- Sample plate definition
AB=Assay buffer, S=sample; S1, 2…7 are different concentrations of different DNA samples from low to high, RE=regeneration buffer
1
2
3
4
5
6
7
8
9
10
11
12
A
AB
GST-ZMYND11
AB
S1
AB
RE
AB
B
AB
GST-ZMYND11
AB
S2
AB
RE
AB
C
AB
GST-ZMYND11
AB
S3
AB
RE
AB
D
AB
GST-ZMYND11
AB
S4
AB
RE
AB
E
AB
GST-ZMYND11
AB
S5
AB
RE
AB
F
AB
GST-ZMYND11
AB
S6
AB
RE
AB
G
AB
GST-ZMYND11
AB
S7
AB
RE
AB
H
AB
GST-ZMYND11
AB
AB
AB
RE
AB
- Assign the sample type to the sample plate by left click the lane numbers. Here each lane of the plate is defined as ‘buffer (B), loading (L), buffer (B), sample (colored pink), buffer (B),…regeneration (R), buffer (B)’ (Figure 3).

Figure 3. Sample plate definition
- Prepare 30 µg/ml GST-ZMYND11 proteins in BLI binding buffer in a total volume of 200 μl/well multiplied by the number of conditions you need to test.
- Prepare a dilution series (six here) of your DNA sample in binding buffer ranging from 3.125 to 100 μM final concentrations in a total volume of 500 μl in case you want duplicate.
- Dispense 200 μl of protein/buffer in each well. Prepare samples in duplicate.
- Sensor assignment. Choose the right positions where the sensors are assigned.
- Insert the Sensor Tray into the Sensor Tray Holder and the Sample Plate into the Sample Plate Holder (refer to Figure 4 and manufacturer’s manual).

Figure 4. The internal structure of Octet Red96 system
Notes:
- Sample & concentration settings are quite important. Typical final protein concentration is around 20-50 µg/ml (determined by UV280 absorbance). According to the binding affinity (KD) to test, it is recommended that DNA dilution series are set in such a concentration range that cover 0.5x KD ~ 20x KD (the DNA concentration is determined by UV260 absorbance) and pre-test to determine the sample concentration and KD are necessary. The system used here has 8 sensors and uses a 96-well sample plate. Thus, interaction between a sensor-immobilized biomolecule and its binding partner in solution can be tested under different conditions with multiple sensors in parallel. For example, interactions were measured with six different concentrations of the binding partner in parallel.
Pre-test 1:
- Relative high concentration (10x KD/100x KD).
- Positive control.
- Blank control.
- Yes/No binding. Yes samples go next round detection.
- Yes samples go this round or KD test directly according to the researcher.
- Find the proper concentration range (using a large dilution factor such as 10 or 5
to determine the highest and lowest concentration) if pre-test 2 performed.
- No less than 4 concentrations should be included in sample dilution series with
a dilution factor 2 or 3.
- Positive/negative control (not necessary).
- Blank control.
- Relative high concentration (10x KD/100x KD).
- Once the sensors were prepared, the assays were quite easy to conduct because the instrument can perform all of the assay functions without attention from the analyst. For example, the 96-well plate does not need to be removed and washed with a separate instrument at various stages in the analysis. Also, substrate does not need to be prepared and added, etc. These are potential advantages of the biolayer interferometry technique (refer to manufacturer’s manual).
- Sample & concentration settings are quite important. Typical final protein concentration is around 20-50 µg/ml (determined by UV280 absorbance). According to the binding affinity (KD) to test, it is recommended that DNA dilution series are set in such a concentration range that cover 0.5x KD ~ 20x KD (the DNA concentration is determined by UV260 absorbance) and pre-test to determine the sample concentration and KD are necessary. The system used here has 8 sensors and uses a 96-well sample plate. Thus, interaction between a sensor-immobilized biomolecule and its binding partner in solution can be tested under different conditions with multiple sensors in parallel. For example, interactions were measured with six different concentrations of the binding partner in parallel.
- Start by defining the number of conditions you need to test, including appropriate controls such as protein sample in buffer only, or DNA sample in buffer only. This will determine the total volume of protein and DNA needed to conduct the entire experiment. Each well is conducted in a total volume of 200 μl.
- Program
All experiments consisted of repeated cycles, each having six steps:
- Incubation with the BLI binding buffer to establish a baseline signal (baseline, 2 min).
- Incubation with protein sample with putative tags to be immobilized onto the biosensors (loading, 5 min).
- Incubation with the BLI binding buffer again to remove unbound proteins to a baseline signal (baseline, 2 min).
- Incubation with the DNA sample (association, 5 min).
- Incubation with the BLI binding buffer to remove the DNA sample (dissociation, 5 min) followed by a 90 sec regeneration step using glycine.
This is can be performed by assay definition. Here the assay includes the following steps: baseline-loading-baseline-association-dissociation-regeneration-neutralization.
Connect each step to corresponding lane and review the assay before click Run button.
Each assay cycle takes about 15 min. All experiments were conducted at 25 °C, and the total volume of the test solution in all cases was 0.2 ml. The samples are stirred by orbital shaking throughout the assay (1,000 rpm. vibration here).
- Incubation with the BLI binding buffer to establish a baseline signal (baseline, 2 min).
- Data analysis
- Save data as response upon binding (nm) vs time. Export file in a format suitable for import into Microsoft Excel or Origin for analysis.
Collected experimental data is analyzed according to the fitting algorism supplied by vendor of the Octet RED96 system. Usually, steady-state analysis shows great agreement with binding kinetics analysis and 1:1 fitting model is employed here.
Note: Check the chi-square value and R2 value supplied by the software. Chi-square/DOF (degrees of freedom): if using 1:1 fitting model, the DOF=1, and chi-square value should be equal to 3.5. Thus, chi-square<3.5 and 1>R2>0.8 indicate a reasonable data.
- Below is an example of a typical BLI experiment.

Figure 5. Experimental setup of BLI assay includes plate definition, assay definition and sensor assignment
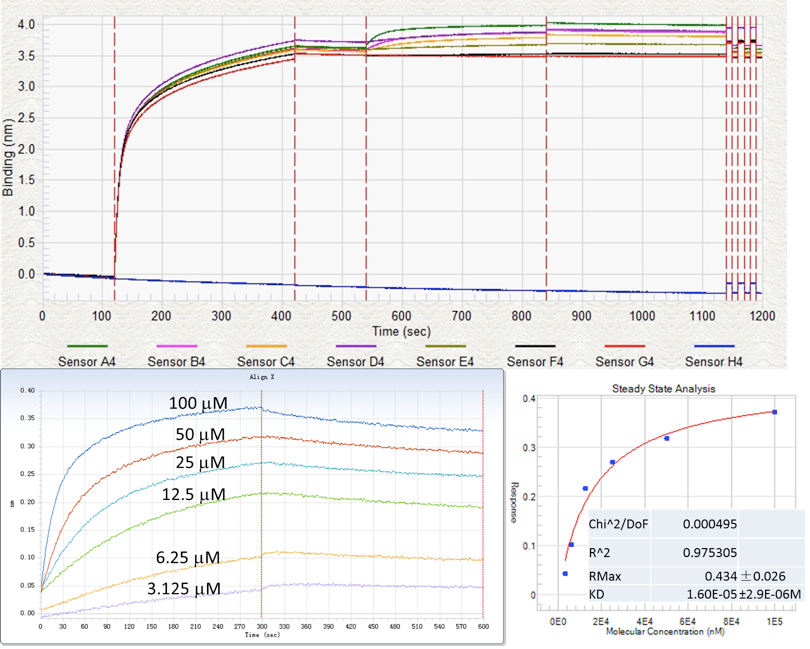
Figure 6. Runtime binding chart and data analysis of a typical BLI. GST-tagged BP protein (30 µg/ml) was immobilized onto anti-GST sensor and a series of dilution sample of methylated DNA (100 µM, 50 µM, 25 µM, 12.5 µM, 6.25 µM, 3.125 µM, from top to bottom) was used as the binding partner. Steady state analysis and 1:1 fitting model was employed here.
- Save data as response upon binding (nm) vs time. Export file in a format suitable for import into Microsoft Excel or Origin for analysis.
Notes
- The instrument can measure the response from eight sensors simultaneously. From preliminary experiments it was clear that substantial variability existed in the output from the individual sensors. The variability in response between sensors suggested that using certain sensors for generating the calibration curve and other sensors for measuring the responses from unknown (or spiked) samples would likely lead to poor quantitation of interaction.
- The optimization of BLI binding buffer aims to get your protein and the binding most stable. Theoretically, BLI is compatible with any kind of buffer component. Of noted is that if using biotinylated molecules, Tris should not be used. The optimal BLI binding has to be determined before BLI binding buffer.
Recipes
- BLI binding buffer
The composition of the binding buffer used can be optimized according to the stability and activity of each protein of interest.
The following recipe was used to study ZMYND11
25 mM NaH2PO4 (pH 7.0)
150 mM KCl
2 mM MgCl2
- DNA solutions
Two complementary single stranded DNA were chemically synthesized. Double stranded DNA was obtained by annealing following the protocol below:
- Mix equal volumes of both complementary oligos (at equimolar concentration of 100 mM) in a 1.5 ml microfuge tube.
- Place tube in a standard heatblock at 90-95 °C for 3-5 min.
- Remove the heat block from the apparatus and allow to cool to room temperature (or at least below 30 °C) on the workbench. Slow cooling to room temperature should take 45-60 min.
- Stored on ice or at 4 °C until ready to use.
50 mM stock prepared in annealing buffer: 20 mM Tris (pH 7.0), 50 mM NaCl and 1 mM EDTA
- Mix equal volumes of both complementary oligos (at equimolar concentration of 100 mM) in a 1.5 ml microfuge tube.
Acknowledgments
We thank W. Chu at Tsinghua the China National Center for Protein Sciences Beijing for providing facility support and technical assistance. This work was supported by grants to X. S. (CPRIT RP110471, Welch G1719, American Cancer Society RSG-13-290-01-TBE, and National institutes of Health (NIH)/MDACC CCSG CA016672), H. L. (The Major State Basic Research Development Program in China, 2011CB965300 and Program for New Century Excellent Talents in University), W. L. (CPRIT RP110471, NIH R01HG007538), B. L. (NIH R01GM090077, Welch I1713), Y. L. (China Postdoctoral Science Foundation, 2012M510413) and H. W. (MD Anderson IRG, Center for Cancer Epigenetics pilot grant). Y. L. is a Tsinghua-Peking Center for Life Sciences postdoctoral fellow. W.L. is a recipient of a Duncan Scholar Award and X.S. is a recipient of a Kimmel Scholar Award.
References
- Abdiche, Y., Malashock, D., Pinkerton, A. and Pons, J. (2008). Determining kinetics and affinities of protein interactions using a parallel real-time label-free biosensor, the Octet. Anal Biochem 377(2): 209-217.
- Abdiche, Y., Malashock, D., Pinkerton, A. and Pons, J. (2009). Exploring blocking assays using Octet, ProteOn, and Biacore biosensors. Anal Biochem 386(2): 172-180.
- Do, T., Ho, F., Heidecker, B., Witte, K., Chang, L. and Lerner, L. (2008). A rapid method for determining dynamic binding capacity of resins for the purification of proteins. Protein Expr Purif 60(2): 147-150.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Li, Y., Wen, H., Shi, X. and Li, H. (2015). Determination of Protein-DNA (ZMYND11-DNA) Interaction by a Label-Free Biolayer Interferometry Assay. Bio-protocol 5(4): e1402. DOI: 10.21769/BioProtoc.1402.
Category
Biochemistry > Protein > Interaction > Protein-DNA interaction
Molecular Biology > DNA > DNA-protein interaction
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










