- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Hanging Drop Aggregation Assay of Breast Cancer Cells
Published: Vol 5, Iss 3, Feb 5, 2015 DOI: 10.21769/BioProtoc.1393 Views: 21407
Reviewed by: HongLok LungAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

In vitro Flow Adhesion Assay for Analyzing Shear-resistant Adhesion of Metastatic Cancer Cells to Endothelial Cells
Shin-Ae Kang [...] Takemi Tanaka
Feb 20, 2016 14268 Views
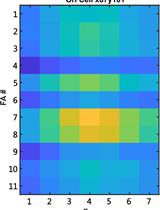
Studying Cellular Focal Adhesion Parameters with Imaging and MATLAB Analysis
Ling-Yea Yu [...] Feng-Chiao Tsai
Nov 5, 2023 1910 Views
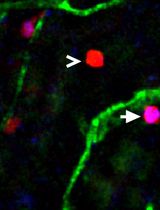
Utilizing EdU to Track Leukocyte Recruitment to the Brain
Zoie K. Lipfert [...] David P. Sullivan
Dec 5, 2025 1600 Views
Abstract
Hanging drop assay can be used to investigate cell-cell cohesion and cell-substratum adhesion through generation of 3D spheroids under physiological conditions. It also can be used to elucidate the role of cell-cell or cell-ECM interactions in specifying spatial relationships between two (or more) different cell populations. This simple method requires no specialized equipment and provides a means of generating tissue-like cellular aggregates for measurement of biomechanical properties for molecular and biochemical analysis in a physiologically relevant model.
Keywords: Hanging dropMaterials and Reagents
- Human breast cancer cell lines T-47D (ATCC, catalog number: ATCC® HTB-133TM ), MCF7 (ATCC, catalog number: ATCC® HTB-22TM )
- Trypsin-EDTA (0.05%) (phenol red) (Life Technologies, catalog number: 25300062 )
- Gibco® RPMI media 1640 (Life Technologies, catalog number: 11875-085 )
- Gibco® PBS (Life Technologies, catalog number: 10010023 )
- Fetal bovine serum (Life Technologies, catalog number: 16000-044 )
- Gibco® penicillin-streptomycin-glutamine (100x) (Life Technologies, catalog number: 10378-016 )
- Complete medium (see Recipes)
Equipment
- CorningTM culture dishes (Corning, catalog number: 430591 )
- Inverted microscope (ZEISS)
- Haemocytometer (Thermo Fisher Scientific, catalog number: 02-671-52 )
Procedure
- Plate T-47D and MCF7 cells in complete medium and let them grow to reach a confluency of 80%.
- Remove the complete medium from culture dish and add 0.05% trypsin for 3-5 min after PBS washing. As soon as cells have detached, collect cells by adding complete medium.
- After spinning down (1,000 x g for 5 min), count the cells using a haemocytometer and adjust cell concentration to 2.0 x 106 cells/ml# in complete culture medium.
- Carefully deposit cells in 30 μl drops on the inner side of a 100 mm dish lid. It is possible to place at least 30 drops per lid (Figure 1A).
- Turn the lid of the plate upside down and place on top of the plate filled with 10 ml PBS (to humidify the culture chamber after distribution of the drops) (Figure 1B). Cells under the force of gravity will be aggregated at the bottom of the hanging drop. Make sure avoid placing the drops close to the plate edges and drops are placed sufficiently apart so as to not touch.
- Monitor the drops daily and incubate until either cell sheets or aggregates have formed, and photograph cell clusters at the endpoint of experiment* by a microscopy within 20 min. Adjust the lighting and focus to provide the most contrast between the 3D structure and background**.
- Collect cell clusters by wide end of tips (100 µl pipette tip cut widely 5-6 mm from the end), and disburse them by pipetting up and down at least 7 times following with a wash of PBS supplemented with 10% fetal bovine serum.
- Count single cells released from the clusters using a haemocytometer.

Figure 1. A. 30 μl drops on the inner side of a 100 mm dish lid; B. lid with drops upside down and on top of the plate filled with 10 ml PBS.
Representative data

Figure 2. Photographs of representative spheroids from T-47D and MCF7 breast cancer cell lines. T-47D and MCF7 cells were cultured in hanging drops and formation of spheroids was observed at 2 and 4 days after culture.
Notes
- This protocol works equally well with other types of cancer cells or genetic modified cancer cells (Defresne et al., 2011; Yip and Cho, 2013; Kumar et al., 2014; Teng et al., 2013).
- #Cell concentration may need to be adjusted up- or down depending on cell size.
- *The assay may be conducted longer than 6 days if desired.
- **The use of fluorescence microscopy for cells that forced express fluorescent protein (such as GFP, RFP) or have a fluorescent label may improve contrast and subsequent analysis.
Recipes
- Complete medium
90% RPMI media 1640
10% fetal bovine serum
1% penicillin-streptomycin-glutamine
Acknowledgments
This study was supported, in part, by the National Institutes of Health Grant CA120510.
References
- Defresne, F., Bouzin, C., Grandjean, M., Dieu, M., Raes, M., Hatzopoulos, A. K., Kupatt, C. and Feron, O. (2011). Preconditioned endothelial progenitor cells reduce formation of melanoma metastases through SPARC-driven cell-cell interactions and endocytosis. Cancer Res 71(14): 4748-4757.
- Kumar, A., Fan, D., Dipette, D. J. and Singh, U. S. (2014). Sparstolonin B, a novel plant derived compound, arrests cell cycle and induces apoptosis in N-myc amplified and N-myc nonamplified neuroblastoma cells. PLoS One 9(5): e96343.
- Teng, Y., Mei, Y., Hawthorn, L. and Cowell, J. (2013). WASF3 regulates miR-200 inactivation by ZEB1 through suppression of KISS1 leading to increased invasiveness in breast cancer cells. Oncogene 33(2): 203-211.
- Yip, D. and Cho, C. H. (2013). A multicellular 3D heterospheroid model of liver tumor and stromal cells in collagen gel for anti-cancer drug testing. Biochem Biophys Res Commun 433(3): 327-332.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Teng, Y. (2015). Hanging Drop Aggregation Assay of Breast Cancer Cells. Bio-protocol 5(3): e1393. DOI: 10.21769/BioProtoc.1393.
Category
Cancer Biology > Invasion & metastasis > Cell biology assays > Cell adhesion
Cancer Biology > General technique > Tumor formation > Cell adhesion
Cell Biology > Cell movement > Cell migration
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











