- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Analysis of Protein Stability by the Cycloheximide Chase Assay
Published: Vol 5, Iss 1, Jan 5, 2015 DOI: 10.21769/BioProtoc.1374 Views: 63390
Reviewed by: Hsin-Yi ChangGuillermo GomezAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
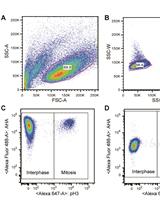
Measuring Protein Synthesis during Cell Cycle by Azidohomoalanine (AHA) Labeling and Flow Cytometric Analysis
Koshi Imami and Tomoharu Yasuda
Apr 20, 2019 9790 Views
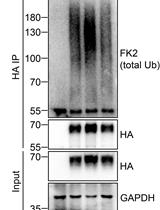
Analysis of the Ubiquitination and Phosphorylation of Vangl Proteins
Di Feng [...] Bo Gao
Oct 20, 2022 3466 Views
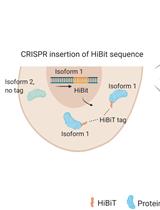
Isoform-specific, Semi-quantitative Determination of Highly Homologous Protein Levels via CRISPR-Cas9-mediated HiBiT Tagging
Kristina Seiler [...] Mario P. Tschan
Jul 20, 2023 2552 Views
Abstract
Comparison of protein stability in eukaryotic cells has been achieved by cycloheximide, which is an inhibitor of protein biosynthesis due to its prevention in translational elongation. It is broadly used in cell biology in terms of determining the half-life of a given protein and has gained much popularity in cancer research. Here we present a full cycloheximide chase assay in our laboratory using a lung adenocarcinoma cell line, CL1-5, as a model.
Materials and Reagents
- Cell lines
- CL1-5/Flag-wtSlug stable cells
- CL1-5/Flag-4SA stable cells
- CL1-5/Flag-wtSlug stable cells
- Antibodies
- Flag M2 antibodies (Sigma-Aldrich, catalog number: F3165 )
- β–actin antibodies (Sigma-Aldrich, catalog number: A5441 )
- Flag M2 antibodies (Sigma-Aldrich, catalog number: F3165 )
- Buffer and media
- 100 mg/ml cycloheximide (CHX) stock solution (Sigma-Aldrich, catalog number: C7698 )
- RPMI-1640 medium (Life Technologies, Gibco®, catalog number: 11875-176 )
- Fetal bovine serum (FBS) (Life Technologies, Gibco®, catalog number: 10082147 )
- HEPES (Sigma-Aldrich, catalog number: H3784 )
- Streptomycin/Penicillin (5,000 U/ml) (Life Technologies, Gibco®, catalog number: 15070063 )
- Sodium pyruvate (Sigma-Aldrich, catalog number: P5280 )
- Sodium bicarbonate (Sigma-Aldrich, catalog number: S5761 )
- G418 (Sigma-Aldrich, catalog number: A1720 )
- Sodium chloride (Sigma-Aldrich, catalog number: S7653 )
- IGEPAL CA-630 (Sigma-Aldrich, catalog number: I8896 )
- Sodium orthovanadate (Sigma-Aldrich, catalog number: S6508 )
- Sodium fluoride (Sigma-Aldrich, catalog number: S7920 )
- Sodium pyrophosphate (Sigma-Aldrich, catalog number: 221368 )
- Protease inhibitor cocktail with EDTA (Roche, Catalog number: 04693116001 )
- BCA protein assay kit (Pierce, catalog number: 23225 )
- DTT (Sigma-Aldrich, catalog number: 43815 )
- SDS (Sigma-Aldrich, catalog number: L3771 )
- Glycerol (Sigma-Aldrich, catalog number: G5516 )
- Bromophenoblue (Sigma-Aldrich, catalog number: B5525 )
- Complete medium (see Recipes)
- Protein lysis buffer (see Recipes)
- 5x sample buffer (see Recipes)
- 100 mg/ml cycloheximide (CHX) stock solution (Sigma-Aldrich, catalog number: C7698 )
Equipment
- CO2 incubator (Thermo Fisher Scientific, Forma series II)
- 35-mm culture dishes (Corning, catalog number: 430165 )
- 1.5 ml tubes
- Standard sonicator
Software
- MetaMorph software (Molecular Devices)
Procedure
Determine the time course of cycloheximide chase beforehand. There can be great variation among proteins. For an unfamiliar protein, it is recommended to start with a 4-hour interval and chase till 24 h. Cycloheximide may cause cytotoxicity to certain cells if cells are exposed to it over 20-24 h. It has been known that Slug is a labile protein with a T1/2 of about 40 min (Wang et al., 2009). CL1-5 cells stably overexpressing Flag-tagged wtSlug and non-phosphorylatable Slug-4SA were prepared using lentiviral transduction (Kao et al., 2014). Cells were grown in complete medium with 400 μg/ml G418. To compare the turnover of wtSlug and Slug-4SA, these stable cells were subjected to the cycloheximide chase assay.
- Seed about 6 x 105 cells in 35-mm dishes and have them incubated in a CO2 incubator overnight.
- After 12-h incubation, remove the medium and add complete medium with 300 μg/ml* cycloheximide (dissolved in DMSO) into each dish.
*The concentration of cycloheximide depends on the cell line used. For CL1-5 cells, 300 μg/ml cyclcoheximide does not cause apoptosis within the time course of the chase assay. It is recommended to start a cell line test with different concentrations of cycloheximide (ranging from 50-300 μg/ml) for a least 8-h chase. For transfected cells, to avoid the sub-efficient cycloheximide concentrations, the expression of tagged proteins should be determined in the absence or presence of different concentrations of cycloheximide by Western blot at 24 h after transfection. - Prepare the protein lysis buffer with freshly-added protease inhibitor. About 200 μl of lysis buffer is needed for one 35-mm dish.
- Lyse t = 0 h cells with the protein lysis buffer and store the lysates in a -80 °C freezer.
- Collect lysates at other time points (t = 1, 2, 3, 6 h) according to the experimental design.
- After all the lysates have been collected, sonicate the protein lysates with a sonicator for 10 times (1 sec/time) on ice.
- Centrifuge lysates at 12,000 rpm at 4 °C for 30 min.
- Collect and transfer the supernatants to new tubes with corresponding labels.
- Determine protein concentrations by the BCA protein assay kit.
- Take 50 μg of proteins, add 5x sample buffer, and incubate samples at 100 °C for 10 min.
- Analyze the results by the SDS-PAGE assay and Western blotting (anti-Flag M2 antibodies with 1:5,000 dilution; β-actin antibodies with 1:10,000 dilution).
- Quantify the Western bands of Flag and β-actin in triplicates using MetaMorph Software or other equivalent softwares.
Quantification
Quantification was achieved by the MetaMorph software in this case. An analysis guide should be referred to for a first-time user. Basically, regions of measurement are selected and threshold is defined. Draw region of interest on top of the western blot bands and quantitate its total grey signal (this can be done in Image J and any other image analysis software). The values of total thresholded areas are measured; a ratio of Flag-Slug to its β-actin is then calculated. The final Slug protein turnover rate at each time point is the percentage of Slug/β-actin at t = 0 of each experimental group. A plot can be created afterwards.
Representative data
Western blots and quantification images have been published in Oncogene (Kao et al., 2014). For details, please refer to Figure 2f at Kao et al. (2014).
Recipes
- Complete medium
RPMI-1640
10% FBS
15 mM HEPES (pH 7.01)
100 U/ml streptomycin/penicillin
1 mM sodium pyruvate
44 mM sodium bicarbonate - Protein lysis buffer
20 mM Tris (pH 7.5)
100 mM NaCl
1% IGEPAL CA-630
100 mM Na3VO4
50 mM NaF
30 mM sodium pyrophosphate - 5x sample buffer
250 mM Tris-HCl (pH 6.8)
500 mM DTT
10% SDS
50% glycerol
0.1% bromophenoblue
Acknowledgments
This work was supported by grants from the National Science Council, Taiwan (NSC99-2628-B-006-031-MY3 NSC101-2325-B-006-018, NSC100-2321-B-002-071 and NSC101-2321-B002-068), National Taiwan University, Taiwan (10R71601-2), and National Institute of Health, USA (R01-GM-094231, to AIN). SP Wang is supported by a Human Frontier Science Program long-term fellowship. This protocol was adapted from previous work published in Oncogene (Kao et al., 2014).
References
- Kao, S. H., Wang, W. L., Chen, C. Y., Chang, Y. L., Wu, Y. Y., Wang, Y. T., Wang, S. P., Nesvizhskii, A. I., Chen, Y. J., Hong, T. M. and Yang, P. C. (2014). GSK3beta controls epithelial-mesenchymal transition and tumor metastasis by CHIP-mediated degradation of Slug. Oncogene 33(24): 3172-3182.
- Wang, S. P., Wang, W. L., Chang, Y. L., Wu, C. T., Chao, Y. C., Kao, S. H., Yuan, A., Lin, C. W., Yang, S. C., Chan, W. K., Li, K. C., Hong, T. M. and Yang, P. C. (2009). p53 controls cancer cell invasion by inducing the MDM2-mediated degradation of Slug. Nat Cell Biol 11(6): 694-704.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Kao, S., Wang, W., Chen, C., Chang, Y., Wu, Y., Wang, Y., Wang, S., Nesvizhskii, A. I., Chen, Y., Hong, T. and Yang, P. (2015). Analysis of Protein Stability by the Cycloheximide Chase Assay. Bio-protocol 5(1): e1374. DOI: 10.21769/BioProtoc.1374.
Category
Cancer Biology > General technique > Biochemical assays > Protein analysis
Biochemistry > Protein > Synthesis
Molecular Biology > Protein > Detection
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










