- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Competitive ELISA for Protein-lipopolysaccharide (LPS) Binding
Published: Vol 4, Iss 22, Nov 20, 2014 DOI: 10.21769/BioProtoc.1298 Views: 15685
Reviewed by: Anonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
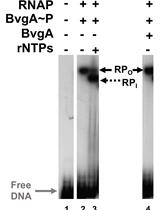
Combining Gel Retardation and Footprinting to Determine Protein-DNA Interactions of Specific and/or Less Stable Complexes
Meng-Lun Hsieh [...] Deborah M. Hinton
Dec 5, 2020 3620 Views
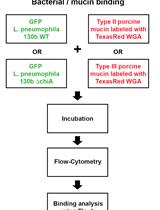
Assay for Assessing Mucin Binding to Bacteria and Bacterial Proteins
Lubov S. Grigoryeva [...] Nicholas P. Cianciotto
Mar 5, 2021 4805 Views
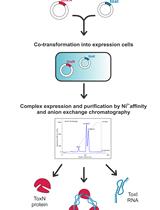
Large-scale Purification of Type III Toxin-antitoxin Ribonucleoprotein Complex and its Components from Escherichia coli for Biophysical Studies
Parthasarathy Manikandan [...] Mahavir Singh
Jul 5, 2023 2205 Views
Abstract
Lipopolysaccharide is the major constituent of the outer membrane of gram-negative bacteria and, once released from the bacterial surface into the bloodstream, is a potent activator of the host immune system, which can lead to septic shock. LPS has a hydrophilic region consisting of a repeating oligosaccharide that is strain-specific (O-antigen) and a core polysaccharide, which is covalently linked to a hydrophobic lipid moiety (lipid A). Lipid A is the most conserved part and is responsible for the toxicity of LPS. Therefore, finding molecules able to bind to this region and neutralize LPS toxicity is of relevant interest as it may provide new therapies to prevent septic shock (Chen et al., 2006). Several proteins and peptides were reported to bind LPS and alter its toxicity towards reduction and even enhancement (Brandenburg et al., 1998), such as serum albumin (Ohno and Morrison, 1989), lipopolysaccharide binding protein (LBP) (de Haas et al., 1999), casein (López-Expósito et al., 2008), lysozyme, the antibiotic polymyxin B and antimicrobial peptides (Chen et al., 2006). Although some of these proteins are neutral and even anionic/acidic (pI<7) (Jang et al., 2009), due to the amphipathic structure of LPS and the presence of negatively charged phosphate groups on the lipid A, the most important factors that are considered for optimal binding to LPS are a cationic/basic (pI>7) and amphipathic nature (Chen et al., 2006).
Here we describe a competitive ELISA that can be used to identify proteins or peptides that bind LPS, as a first approach before analyzing the possible activity in vitro and in vivo. In this ELISA, serial dilutions of the protein or peptide to be tested are preincubated with a fixed concentration of fluorescein isothiocyanate (FITC)-labeled LPS from Escherichia coli serotype O111:B4 and then added to wells of a microtitre plate which are blocked with a casein hydrolysate that binds LPS (Martínez-Sernández et al., 2014). Binding of the protein to LPS displaces LPS from binding to the casein, which is revealed using a horseradish peroxidase (HRP)-labeled anti-FITC polyclonal conjugate. This method allows simultaneous analysis of several proteins or peptides in a short period of time and no recognizing molecules (e.g., antibodies) to a specific protein or peptide are needed.
Materials and Reagents
- Casein (Hammarsten grade) (BDH Prolabo, VWR International, catalog number: 440203H )
- Lipopolysaccharides from Escherichia coli O111: B4-FITC conjugate (Sigma-Aldrich, catalog number: F3665 )
- Polymyxin B sulfate salt (Sigma-Aldrich, catalog number: P4119 )
- Proteins that were used for the example below (see Representative data) but which are not necessary for the assay:
- Salts
- Sheep anti-FITC: HRP (Bio-Rad Laboratories, AbD Serotec, catalog number: 640005 )
- SIGMAFASTTM OPD (Sigma-Aldrich, catalog number: P9187 )
- Tween® 20 (Merck KGaA, Merck Millipore, catalog number: 822184 )
- 0.3 M NaOH (prepared from 50% w/w NaOH solution) (18.94 M) (Sigma-Aldrich, catalog number: 415413 )
- 7.4% HCl (37% commercial solution diluted 1/5 with distilled water) (Merck KGaA, Merck Millipore, catalog number: 100317 )
- Phosphate buffered saline (PBS) containing 0.05% Tween® 20 (PBS-T)
- 3 N H2SO4 (1.5 M)
- Casein hydrolysate solution (see Recipes)
- Concentrated PB (see Recipes)
- PBS (see Recipes)
- PBS-EDTA (see Recipes)
Equipment
- 96-well microplate (polystyrene with clear flat bottom wells) (Greiner Bio-One GmbH, catalog number: 762071 )
- Automatic (mono- and multichannel) pipettes and pipette tips
- Calibrated beakers, graduated cylinders and conical flasks
- Magnetic stir bars and magnetic stirrer
- Manual or automatic microplate washer (e.g., DAS plate washer)
- Microcentrifuge tubes (e.g., Eppendorf; Kartell, catalog numbers: 297, for 1.5 ml , and 1298 , for 0.5 ml tubes)
- Microplate adhesive sealing films (e.g., AlumaSeal® II film; Sigma-Aldrich, catalog number: A2350 )
- Microplate reader (equipped for measuring absorbance at 492 nm) (e.g., Beckman Coulter, model: Biomek® plate reader )
- Microtitre plate shaker with a small orbit diameter (preferably 1.5 mm) set to 750 rpm (e.g., Stuart, model: SSM5 )
- pH meter
- Reagent reservoir for multichannel pipettes
- Scales
- Shaking incubator set to 37 °C and 150 rpm (e.g., Panasonic Corporation, SANYO Electric, model: Orbi-Safe TM)
Procedure
- Determine the number of wells needed for the assay. The test should be performed at least in duplicate. Following the ELISA worksheet example below, if you test in duplicate 8 serial dilutions of an unknown protein/peptide plus the same dilutions of polymyxin B (positive inhibition control) and a known protein/peptide with no binding ability to LPS (negative inhibition control) you need 52 wells: 3 x 8 x 2 = 48 wells for protein/peptide dilutions, two wells with LPS-FITC (reference control) and two wells without LPS-FITC (negative control).
- Add 200 µl of casein hydrolysate solution to the wells of a 96-well microplate. Seal the microplate with the adhesive film and incubate overnight at 4 °C without shaking.
- Separately, prepare a 2x solution of LPS O111: B4-FITC conjugate (5 µg/ml) and 2x serial dilutions of the protein/peptide to be tested (starting dilution 80 µg/ml) in PBS-EDTA. Prepare the same dilutions of polymyxin B (positive inhibition control) and a negative inhibition control (e.g., ovalbumin). Eight serial ¼ dilutions are optimal for most proteins. For the ELISA worksheet example prepare:
- 2x solution of LPS-FITC: 3.5 ml of LPS-FITC at 5 µg/ml in PBS-EDTA.
- 2x serial dilutions of the proteins/peptides in PBS-EDTA:
Table 1. Preparation of serial dilutions
Dilution (dil) number
Volume and source of proteins/peptides
Volume of PBS-EDTA
Protein concentration
1
3.2 μl from a 5 mg/ml stock
196.8 μl
80 µg/ml
2
50 μl of dil 1
150 μl
20 µg/ml
3
50 μl of dil 2
150 μl
5 µg/ml
4
50 μl of dil 3
150 μl
1.25 µg/ml
5
50 μl of dil 4
150 μl
0.312 µg/ml
6
50 μl of dil 5
150 μl
0.078 µg/ml
7
50 μl of dil 6
150 μl
0.020 µg/ml
8
50 μl of dil 7 150 μl
0.005 µg/ml
- 2x solution of LPS-FITC: 3.5 ml of LPS-FITC at 5 µg/ml in PBS-EDTA.
- Mix 120 µl of LPS-FITC solution (2x) with 120 µl of each dilution (2x) or PBS-EDTA (reference control) in individual microcentrifuge tubes, mix well, and incubate the samples for 1 h at RT.
- Aspirate the content of the wells coated with the casein hydrolysate and wash 3 times with 200 µl of PBS (no soak) at RT without shaking.
- Immediately, add 100 µl of the preincubated solutions or PBS-EDTA (negative control) to each well in duplicate. Seal the microplate and incubate for 30 min at RT under shaking at 750 rpm.
- Wash the plate 5 times with 200 µl of PBS-T (no soak) at RT without shaking.
- Add 100 µl of sheep anti-FITC: HRP diluted 1/4,000 in PBS-T. Seal the microplate and incubate for 30 min at RT under shaking at 750 rpm.
- Wash the plate as described in step 7.
- Prepare the chromogen [o-phenylenediamine dihydrochloride (OPD)] solution following the manufacturer’s instructions.
- Add 100 µl of the OPD solution to each well and incubate for 20 min at RT, without shaking and in the dark.
- Add 25 µl of 3 N H2SO4 to each well to stop the reaction.
- Read the absorbance of the wells at 492 nm within 30 min.
- The percentage of inhibition for each protein dilution is calculated according to the formula: [(average OD reference control) - (average OD protein dil + LPS-FITC)]/ (average OD reference control) x 100. OD: Optical density.
*Alternatively, incubations in steps 6 and 8 can be performed in an incubator for 1 h at 37 °C without shaking, after sealing the plate with the adhesive film.
Table 2. ELISA worksheet example
1
2
3
4
5
6
7
A
100 µl LPS-FITC (reference control)
100 µlProtein dil 1 + LPS-FITC100 µlPolymyxin B dil 1 + LPS-FITC100 µlOvalbumin dil 1 + LPS-FITCB
100 µlProtein dil 2 + LPS-FITC100 µlPolymyxin B dil 2 + LPS-FITC100 µlOvalbumin dil 2 + LPS-FITCC
100 µlPBS-EDTA(negative control)100 µlProtein dil 3 + LPS-FITC100 µlPolymyxin B dil 3 + LPS-FITC100 µlOvalbumin dil 3 + LPS-FITCD
100 µlProtein dil 4 + LPS-FITC100 µlPol ymyxin B dil 4 + LPS-FITC100 µlOvalbumin dil 4 + LPS-FITCE
100 µlProtein dil 5 + LPS-FITC100 µlPolymyx in B dil 5 + LPS-FITC100 µlOvalbumin dil 5 + LPS-FITCF
100 µlProtein dil 6 + LPS-FITC100 µlPolymyxin B dil 6 + LPS-FITC100 µlOvalbumin dil 6 + LPS-FITCG
100 µlProtein dil 7 + LPS-FITC100 µlPolymyxin B dil 7 + LPS-FITC100 µlOvalbumin dil 7 + LPS-FITCH
100 µlProtein dil 8 + LPS-FITC100 µlPolymyxin B dil 8 + LPS-FITC100 µlOvalbumin dil 8 + LPS-FITC
Representative data

Figure 1. Example of inhibition curve obtained with several molecules using the reported method to analyze LPS binding. LPS-FITC (0.25 µg/well) was incubated with four-fold dilutions of polymyxin B (circles), LBP human recombinant (squares), bovine serum albumin (BSA) (inverted triangles) and myoglobin from equine skeletal muscle (triangles). It is noteworthy that an excess of protein/peptide might have a “zone effect” in the assay, as occurs with polymyxin B.
Notes
- The ideal OD range is 0.6-1.2 for the reference control, which was obtained using LPS-FITC at 0.25 µg/well. As the FITC content may vary between batches this concentration might need to be adjusted before performing the inhibition assay. The extent of labeling of the LPS-FITC used to develop this method was of 7.20 µg FITC/mg LPS.
- Binding of a protein to LPS does not imply that it is linked to a biological effect. Further experiments should be done to test the activity in vitro and in vivo.
- Depending on several factors (amino acid composition, type of interaction with LPS, number of binding sites, etc.) each protein or peptide needs a different optimal concentration to obtain the maximal inhibition. Hence, several dilutions of the proteins or peptides need to be assayed.
- The main advantage of using a competitive ELISA instead of an indirect ELISA is that in the latter, the protein/peptide to be tested is coated to the wells and blocking agents may interfere with LPS binding (e.g., binding LPS per se, causing steric hindrance). Casein hydrolysate is commonly used as a blocking agent in ELISA and also serves as LPS binding agent, enabling its use in the competitive system reported.
- Inhibition will occur if the protein or peptide to be tested binds to the same or a close LPS region as the casein hydrolysate. Although casein hydrolysate is composed of several derived peptides and therefore the mechanism of binding to LPS is poorly understood, as polymyxin B inhibits LPS binding to casein, this suggests that the casein hydrolysate binds at least to the lipid A region of LPS.
- LPS is incubated in PBS-EDTA in the absence of detergents as both LPS and detergents have an amphipathic nature and may give unexpected results difficult to interpret when incubated together.
Recipes
- Casein hydrolysate solution
Note: The preparation is based on the method described by Pearce-Pratt and Roser (2010) with some modifications.- Dissolve 2.5 g of casein in 80 ml of 0.3 M NaOH and incubate overnight at 37 °C under shaking at 150 rpm
- Adjust pH to 8 slowly with 7.4% HCl
Note: Aggregation of casein fragments occurs transiently while the pH is adjusted. Keep the pH meter electrode out of the solution when you add HCl. Read the pH or add more HCl dropwise always after the aggregates are dissolved.
- Add 2 ml of concentrated PB (the pH lowers to approximately 7.5)
- Adjust pH to 7.2 with 7.4% HCl
- Made up to 200 ml with distilled water and readjust the pH if necessary
- Dissolve 2.5 g of casein in 80 ml of 0.3 M NaOH and incubate overnight at 37 °C under shaking at 150 rpm
- Concentrated PB (0.75 M, pH 7.2)
For a 50 ml solution:
Dissolve 4.08 g of Na2HPO4 anhydrous (Mw: 141.96) and 1.22 g of NaH2PO4.H2O (Mw: 137.99) up to 50 ml with distilled water
- PBS (150 mM, pH 7.2)
For a 1 L solution:
Dissolve 1.63 g of Na2HPO4 anhydrous (Mw: 141.96), 0.49 g of NaH2PO4.H2O (Mw: 137.99) and 7.89 g of NaCl (Mw: 58.44) up to 1 L with distilled water
- PBS (150 mM, pH 7.2)-EDTA (1 mM)
Add EDTA disodium salt 2-hydrate at a concentration of 1 mM in PBS
As EDTA lowers the pH, readjust the pH to 7.2 with NaOH
Acknowledgments
This work was supported by grants AGL2011-30563-C03 (Ministerio de Ciencia e Innovación, Spain) and CN 2012/155 (Xunta de Galicia, Spain). Victoria Martínez-Sernández holds a predoctoral fellowship (Programa de Formación del Profesorado Universitario) from the Spanish Ministerio de Educación, Cultura y Deporte. This protocol was modified from Martínez-Sernández et al. (2014).
References
- Brandenburg, K., Koch, M. H. and Seydel, U. (1998). Biophysical characterisation of lysozyme binding to LPS Re and lipid A. Eur J Biochem 258(2): 686-695.
- Chen, X., Dings, R. P., Nesmelova, I., Debbert, S., Haseman, J. R., Maxwell, J., Hoye, T. R. and Mayo, K. H. (2006). Topomimetics of amphipathic beta-sheet and helix-forming bactericidal peptides neutralize lipopolysaccharide endotoxins. J Med Chem 49(26): 7754-7765.
- de Haas, C. J., van der Zee, R., Benaissa-Trouw, B., van Kessel, K. P., Verhoef, J. and van Strijp, J. A. (1999). Lipopolysaccharide (LPS)-binding synthetic peptides derived from serum amyloid P component neutralize LPS. Infect Immun 67(6): 2790-2796.
- Jang, H., Kim, H. S., Moon, S. C., Lee, Y. R., Yu, K. Y., Lee, B. K., Youn, H. Z., Jeong, Y. J., Kim, B. S., Lee, S. H. and Kim, J. S. (2009). Effects of protein concentration and detergent on endotoxin reduction by ultrafiltration. BMB Rep 42(7): 462-466.
- Lopez-Exposito, I., Amigo, L. and Recio, I. (2008). Identification of the initial binding sites of alphas2-casein f(183-207) and effect on bacterial membranes and cell morphology. Biochim Biophys Acta 1778(10): 2444-2449.
- Martinez-Sernandez, V., Mezo, M., Gonzalez-Warleta, M., Perteguer, M. J., Muino, L., Guitian, E., Garate, T. and Ubeira, F. M. (2014). The MF6p/FhHDM-1 major antigen secreted by the trematode parasite Fasciola hepatica is a heme-binding protein. J Biol Chem 289(3): 1441-1456.
- Ohno, N. and Morrison, D. C. (1989). Lipopolysaccharide interaction with lysozyme. Binding of lipopolysaccharide to lysozyme and inhibition of lysozyme enzymatic activity. J Biol Chem 264(8): 4434-4441.
- Pearce-Pratt, R. and Roser, B. (2010). Comparison of Blocking Agents for ELISA. Thermo Fisher Scientific Technical Bulletin: 07a.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Martínez-Sernández, V. and Ubeira, F. M. (2014). Competitive ELISA for Protein-lipopolysaccharide (LPS) Binding. Bio-protocol 4(22): e1298. DOI: 10.21769/BioProtoc.1298.
- Ohno, N. and Morrison, D. C. (1989). Lipopolysaccharide interaction with lysozyme. Binding of lipopolysaccharide to lysozyme and inhibition of lysozyme enzymatic activity. J Biol Chem 264(8): 4434-4441.
Category
Microbiology > Microbial biochemistry > Protein > Interaction
Biochemistry > Protein > Immunodetection > ELISA
Biochemistry > Lipid > Lipid-protein interaction
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









