- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Accelerated Storage Stability Testing of a Potential Anti-Anthrax Therapeutic, EnvD
Published: Vol 4, Iss 21, Nov 5, 2014 DOI: 10.21769/BioProtoc.1281 Views: 8602
Reviewed by: Fanglian HeKanika Gera

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
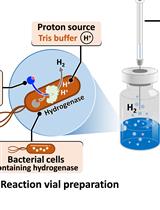
H2 Production from Methyl Viologen–Dependent Hydrogenase Activity Monitored by Gas Chromatography
Nuttavut Kosem
Dec 5, 2023 1796 Views
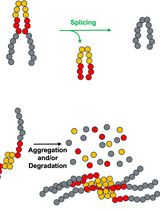
Monitoring Protein Stability In Vivo Using an Intein-Based Biosensor
John S. Smetana [...] Christopher W. Lennon
Apr 20, 2025 1618 Views
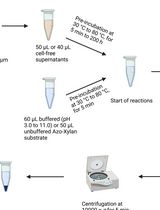
Endo-1,4-β-D-xylanase Assay Using Azo-Xylan and Variants Thereof
Luca Bombardi [...] Salvatore Fusco
Apr 20, 2025 1961 Views
Abstract
The purpose of stability testing is to determine how the properties of a particular therapeutic vary with time under the influence of specific environmental factors. Information regarding the long-term stability of therapeutics can be extrapolated by performing an accelerated storage stability study. Here, we describe an accelerated storage stability study for the potential anti-anthrax therapeutic, EnvD, a poly-γ-D-glutamic acid (PDGA) depolymerase. Storage conditions were based on those recommended by the International Conference on Harmonisation of Technical Requirements for Registration of Pharmaceuticals for Human Use (ICH). Owing to the high-molecular-weight and associated viscosity of PDGA in solution, loss of enzyme activity on storage can be determined as a reduction in the capacity of the depolymerase to reduce the viscosity of the polymer. This work supported by a Medical Research Council Capacity Building Studentship award.
Materials and Reagents
- Poly-γ-D-glutamic acid [prepared in-house; protocol for production can be found in Negus and Taylor (2014)]
- Recombinant EnvD enzyme (prepared in-house)
- PBS (Thermo Fisher Scientific, catalog number: 12821680 )
- Distilled water (prepared in-house)
- Compressed air (prepared in-house)
- 70% ethanol (see Recipes)
Equipment
- 1.5 ml microcentrifuge tubes (Eppendorf, catalog number: 0030120086 )
- Parafilm (Pechiney Plastic Packaging, catalog number: PM996 )
- 5 ml sterile plastic syringes (Terumo Medical Corporation, catalog number: SS-05S )
- Static incubator (Unitemps, discontinued) (B&T)
- Micro-viscometer (Anton Paar GmbH, model: AMVn, catalog number: 1569 )
- Glass capillary for micro-viscometer (1.6 mm internal diameter) (Anton Paar GmbH, catalog number: 67605 )
- Steel balls (1.5 mm diameter) (Anton Paar GmbH, catalog number: 73109 )
- Capillary filling adapater (Anton Paar GmbH, catalog number: 63390 )
Procedure
- Aliquots of freshly isolated EnvD (35 µg total protein) were adjusted to a final volume of 50 µl with sterile PBS in 1.5 ml microcentrifuge tubes and sealed with Parafilm.
- Microcentrifuge tubes were maintained at 37 °C over a period of 30 days to simulate long-term storage. The temperature of the incubator was periodically checked to ensure it did not fluctuate by more than ± 2 °C.
- Samples of EnvD were periodically (0 days, 1 day, 3 days, 7 days, 15 days, 30 days) removed from the incubator and rapidly combined with 400 µg of lyophilised PDGA substrate suspended in 1 ml of PBS.
- Reactions were allowed to proceed for 1 h at 37 °C in a static incubator and were terminated by heating at 95 °C for 10 min. Samples were stored at -20 °C before viscometeric analysis.
- Viscosity of PDGA following incubation with stored enzyme preparations was determined using an Anton Paar falling ball microviscometer. Frozen samples were thawed at room temperature and transferred to a glass viscometry capillary containing a solid steel ball using a 5 ml syringe and filling adapter.
- Viscosity was determined as the time taken for the ball to fall 25 cm through the sample at an angle of 15° to the horizontal; each automated, timed determination was performed four times.
- After each reading the capillary was cleaned with distilled water followed by 70% ethanol. The capillary was then dried with compressed air.
Representative data
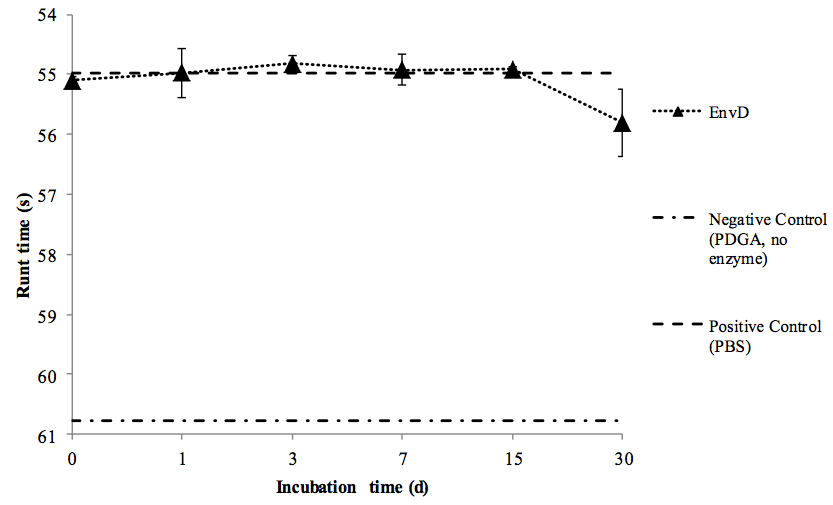
Figure 1. Stability of rEnvD during accelerated storage at elevated temperature. Samples of rEnvD were maintained at 37 °C for 30 d. At the time points indicated, aliquots were combined with PDGA (400 μg/ml), incubated for 1 h at 37 °C and the rune time determined. Ball run time (s) was measured in an Anton-Paar AMVn viscometer; the time taken for the ball to fall 25 cm at an angle of 15° to the horizontal was determined. Error bars represent ± 1 SD (n=8).
Recipes
- 70% ethanol
700 ml ethanol
300 ml distilled water
Acknowledgments
This work supported by a Medical Research Council Capacity Building Studentship award.
References
- Kimura, K. and Itoh, Y. (2003). Characterization of poly-gamma-glutamate hydrolase encoded by a bacteriophage genome: possible role in phage infection of Bacillus subtilis encapsulated with poly-gamma-glutamate. Appl Environ Microbiol 69(5): 2491-2497.
- Negus, D. and Taylor, P. W. (2014). A poly-gamma-(D)-glutamic acid depolymerase that degrades the protective capsule of Bacillus anthracis. Mol Microbiol 91(6): 1136-1147.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Negus, D. and Taylor, P. W. (2014). Accelerated Storage Stability Testing of a Potential Anti-Anthrax Therapeutic, EnvD. Bio-protocol 4(21): e1281. DOI: 10.21769/BioProtoc.1281.
Category
Microbiology > Microbial biochemistry > Protein > Activity
Biochemistry > Protein > Activity
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










