- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Agrobacterium tumefaciens-mediated Transformation of Walnut (Juglans regia)
Published: Vol 4, Iss 19, Oct 5, 2014 DOI: 10.21769/BioProtoc.1258 Views: 14126

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Live Imaging of the Shoot Apical Meristem of Intact, Soil-Grown, Flowering Arabidopsis Plants
Gabriele Bradamante
Jun 20, 2024 2319 Views
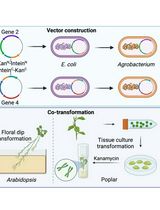
A Novel Gene Stacking Method in Plant Transformation Utilizing Split Selectable Markers
Guoliang Yuan [...] Xiaohan Yang
Feb 20, 2025 2019 Views

Micrografting Technique of Hevea brasiliensis In Vitro Plantlets
Florence Dessailly [...] Julie Leclercq
Feb 20, 2025 1523 Views
Abstract
Like many woody plant species, walnut (Juglans regia) can be difficult to genetically transform and regenerate. However, somatic embryos have been used successfully for over two decades as a target tissue for transformation and regeneration of transgenic walnut plants. Walnut somatic embryos, initiated originally from developing zygotic embryos or anther tissue, will proliferate numerous secondary embryos from single cells in the epidermal layer. These single cells in intact somatic embryos can be efficiently transformed by Agrobacterium tumefaciens (A. tumefaciens). This gene transfer system is most efficient when Agrobacterium binary vector plasmids contain a scorable maker gene (e.g. uidA) and a selectable marker gene (e.g. nptII). This system should be applicable to any crop that undergoes repetitive embryogenesis from single Agrobacterium-susceptible cells. Here we describe the method of transforming somatic embryos in detail so that this technique can be applied to walnut and other woody plant species.
Materials and Reagents
- Biological materials
- Juglans regia somatic embryo cultures
- Disarmed A. tumefaciens strain (e.g. EHA101) harboring binary vector of interest
For this protocol it is assumed that the vector harbors the nptII selectable marker gene and the uidA scorable marker gene (e.g. pDE00.0201 from Escobar et al., 2002).
- Juglans regia somatic embryo cultures
- Chemicals
- Driver Kuniyuki walnut (DKW) medium with vitamins (PhytoTechnology Laboratories®, catalog number: D2470 )
- Kanamycin sulfate (PhytoTechnology Laboratories®, catalog number: K378 )
- Timentin® (PhytoTechnology Laboratories®, catalog number: T869 )
- 4-Hydroxy-3',5'-dimethoxyacetophenone (Acetosyringone) (PhytoTechnology Laboratories®, catalog number: A104 )
- GelzanTM CM (Sigma-Aldrich, catalog number: G1910 )
- SeaKem® LE agarose (Lonza, catalog number: 50004 )
- Bacto® Agar/SuperPureTM agar (BTS, catalog number: A01PD )
- Proline (PhytoTechnology Laboratories®, catalog number: D689 )
- 5-Bromo-4-chloro-3-indolyl β-D-glucuronide (X-gluc) (Sigma-Aldrich, catalog number: B5285 )
- 1 M sodium phosphate buffer (pH 7.0)
- 0.5 M EDTA (pH 8.0) (PhytoTechnology Laboratories®, catalog number: E 582 )
- 0.005 M potassium ferricyanide (Sigma-Aldrich, catalog number: 702587 )
- 0.005 M potassium ferrocyanide (Sigma-Aldrich, catalog number: P3289 )
- Triton X-100 (Sigma-Aldrich, catalog number: 234729 )
- Indole-3-butyric acid (IBA) (Sigma-Aldrich, catalog number: B-5386 )
- 6-Benzylaminopurine (BAP) (Sigma-Aldrich, catalog number: B-3408 )
- Tryptone (BTS, catalog number: T01PD )
- Yeast extract (BTS, catalog number: Y02MG )
- NaCl (Thermo Fisher Scientific, catalog number: S271-3 )
- Saturated ZnSO4 or NH4NO3 solution
- 10x PCR buffer (Applied Biosystems)
- PCR primers
- dNTPs (Applied Biosystems)
- Taq DNA polymerase (Applied Biosystems)
- DNeasy Plant Mini Kit (QIAGEN)
- Driver Kuniyuki walnut (DKW) medium with vitamins (PhytoTechnology Laboratories®, catalog number: D2470 )
- Media and supplements
Microbiological media- Agrobacterium liquid growth medium (LB liquid medium) (see Recipes)
- Agrobacterium growth plates (LB solid medium) (see Recipes)
Plant media- Driver Kuniyuki walnut (DKW) basal medium (see Recipes)
- Virulence induction medium (IM) (see Recipes)
- Acetosyringone medium (AS) plates (see Recipes)
- KAN/Timentin® selection medium (see Recipes)
- KAN only selection medium (see Recipes)
- DKW shoot medium (DKW basal medium with 1 mg/L BAP and 0.01 mg/L IBA) (see Recipes)
- Agrobacterium liquid growth medium (LB liquid medium) (see Recipes)
- Glass and Plasticware
- Sterile empty 100 x 15 mm Petri plates (VWR International, catalog number: 25384-342 )
- Sterile empty 35 x 10 mm Petri plates (BD, FalconTM, catalog number: 25373-041 )
- Sterile disposable 50 ml screw-cap centrifuge tubes (BD, FalconTM, catalog number: 352070 )
- Sterile disposable cotton-plugged 10 ml pipettes
- Plastic pipette tips (20, 200, and 1,000 µl) (Rainin, model: LTS )
- Sterile disposable 6-well multiwell plates (BD Biosciences, Falcon®, catalog number: 08-772-1G )
- Sterile disposable 96-well multiwell plates (BD Biosciences, Falcon®, catalog number: 08-772-2C )
- Filter paper cut to fit in 100 x 15 mm Petri plates and autoclaved
- Filter paper discs cut to the diameter of the wells of a 6-well multiwall plate and autoclaved
- 150 mm diameter desiccator (e.g. Nalgene, catalog number: 5313-0150 )
- Magenta GA-7 vessels (Magenta Corp.)
- Sterile empty 100 x 15 mm Petri plates (VWR International, catalog number: 25384-342 )
Equipment
- Spectrophotometer (e.g. Bio-Rad Laboratories, model: SmartSpec 3000 )
- Laboratory centrifuge (e.g. Sigma-Aldrich, Laborzentrifugen Laboratory Centrifuge 3K 10 )
- Micro-centrifuge (e.g. Denville, model: 260D )
- Continuous power supply units for electrophoresis (e.g. Thermo EC, model: EC 105 and Hoefer, model: HE 33 Mini Submarine unit)
- Electrophoresis units for agarose gels (e.g. Fisher Scientific, models: 03-500-124 , 03-500-132 , and 03-500-134 )
- Laminar flow hood (e.g. EdgeGard, Baker, model: EG3252 )
- Pipetman® 20, 200, and 1,000 µl (Rainin, model: LTS )
- Analytical balance (e.g. Mettler Toledo, model: AT261 DeltaRange® )
- Top loading electronic balance (e.g. Mettler, model: PM 2000 )
- pH meter (e.g. Corning Pinnacle, model: 540 )
- Constant temperature incubator (e.g. Napco, model: 301 )
- Water baths (e.g.Thermo Scientific, model: 2870 )
- Vortex mixer (e.g.: Scientific Industries, model: G 560 )
- Freezer (- 80 °C) (e.g. New Brunswick, model: U 700 Premium)
Procedure
- Obtain actively multiplying walnut somatic embryo cultures or initiate new cultures from zygotic embryos or anther tissue (Dandekar et al., 1989; Mendum and McGranahan, 1995; Leslie et al., 2006).
Several days before initiation of transformation:- Streak Agrobacterium strains out on a LB plates with appropriate antibiotics. Incubate at 28 °C for 48 h.
- Culture sufficient numbers of embryos of the desired genotypes.
- Prepare appropriate antibiotics.
- Make LB liquid medium.
- Make co-cultivation liquid medium.
- Make DKW basal, AS, and selection media.
Select embryos: Pick out 60-70 or more rapidly growing, small, white, uniform embryos (Figure 1A) and place them onto fresh DKW basal medium.
Day 1 (prepare liquid cultures)- Inoculate each strain of Agrobacterium to be used into a 50 ml conical centrifuge tube containing 20 ml of LB medium.
- Place the capped tubes on a rotary shaker at ~200 rpm at 25 °C.
- After 2 h add the appropriate selective antibiotics for the vector used and return to shaker.
- After shaking overnight, the bacterial cultures should be turbid. Determine the Abs600nm of a 10-1 dilution of the culture.
- For 60-70 embryos, approximately 25 ml of co-cultivation suspension will be utilized. The desired Abs600nm of the co-cultivation suspension is ~0.5 (an Abs600nm reading of 0.5 is approximately equivalent to 2.5 x 108 bacteria/ml). To calculate the needed volume of overnight culture, use the following equation:
(Co-cultivation volume needed) x (desired Abs600nm of the co-cultivation suspension)
(Abs600nm of the Agrobacterium culture) x (dilution factor)
For example: If you need 25 ml of co-cultivation suspension at a concentration of 2.5 x 108 bacteria/ml and an aliquot of the Agrobacterium culture diluted 1: 10 gives an A600 reading of 0.371, how much of the Agrobacterium culture do you need to dilute for use?
(25) x (0.5) = 3.36 ml
(0.371) x (10) - Using a sterile pipette, place the calculated volume of Agrobacterium culture into a sterile plastic 50 ml centrifuge tube and centrifuge for 10 min at 4,000 x g and ambient temperature to pellet the bacteria.
- Pour or pipette the supernatant into a waste container and resuspend the pellet in the co-cultivation medium. The pellet is easier to resuspend in a small volume (0.5 ml first) by carefully pipetting up and down, followed by bringing the solution to required final volume.
- Return the tubes to the rotary shaker at ~200 rpm at 25 °C for 1-2 h.
- Streak Agrobacterium strains out on a LB plates with appropriate antibiotics. Incubate at 28 °C for 48 h.
- Co-cultivation
- Place desired number of embryos into a well of a sterile six-well multiwell plate. The well should be no more than half filled with embryos (about 20-23 embryos). Use more than one well for each transformation in case of contamination problems. Transformations of multiple genotypes can be performed in different wells of the same plate but if using multiple bacterial vectors it is advisable to use separate plates to avoid cross-contamination.
- Dispense the appropriate volume of Agrobacterium co-cultivation suspension (about 8 ml, or enough to cover the embryos) into each well using sterile 10 ml pipettes with cotton-plugged ends.
- Incubate at ambient temperature for at least 10-15 min (Figure 1B and Note 1).
- Place a sterile filter paper into a number of empty sterile Petri plates equal to the number of wells used.
- Pipet as much excess co-cultivation liquid as possible from each well into a waste container.
- Transfer the embryos from each well onto the filter paper in the Petri plates using sterile forceps. This will wick excess co-cultivation liquid from the embryos.
- Transfer the embryos to labeled plates of AS medium (about 10 per plate) and place the plates in the dark for 48 h at 20-22 °C (see Video 1).
Video 1. Walnut transformation
Day 4: Selection
After co-cultivating for 48 h, transfer the embryos to plates of KAN/Timentin medium containing 200 mg/L kanamycin and 200 mg/L Timentin (see Note 2). Other selection medium may be utilized depending on the selectable marker gene (see Note 3). Incubate the culture plates in the dark at ambient temperature (20-24 °C).
Day 6 and onward:- Transfer embryos to fresh KAN/Timentin medium after two days and again after an additional five days. This helps to reduce bacterial overgrowth. Thereafter transfer the embryos to fresh KAN/Timentin medium weekly for 8-12 weeks.
- As new somatic embryos begin to develop from the surface of the original (E0) embryos, separate them from the parent embryos. Label these new embryos as E1 embryos. Repeat this process for one more generation (E2 embryos; Figure 1C).
- After 8-12 weeks of selection, embryos can be moved to selection medium containing only kanamycin. Observe the embryo cultures carefully for the next several weeks to ensure that no residual Agrobacterium has survived.
- Place desired number of embryos into a well of a sterile six-well multiwell plate. The well should be no more than half filled with embryos (about 20-23 embryos). Use more than one well for each transformation in case of contamination problems. Transformations of multiple genotypes can be performed in different wells of the same plate but if using multiple bacterial vectors it is advisable to use separate plates to avoid cross-contamination.
- Scoring for GUS expression
- As E2 embryos emerge, test them for GUS (uidA) activity (Jefferson, 1987).
- Pipette 40 µl of X-gluc staining solution (see recipes) into wells of a sterile 96-well multiwell plate.
- Using a fine point scalpel remove a small piece of tissue (cotyledon tips work well) from each well-formed and healthy E2 embryo of interest. Put the tissue piece in the X-gluc solution and label and mark the location of the embryo from which it was excised.
- Incubate at 37 °C and monitor the development of blue color. Color change should be apparent in 10 min to 2 h (Figure 1D).
- If the tissue piece developed the distinctive blue color, the E2 embryo from which it was cut should be separated and multiplied on selection medium until E3 embryos are available for DNA analysis.
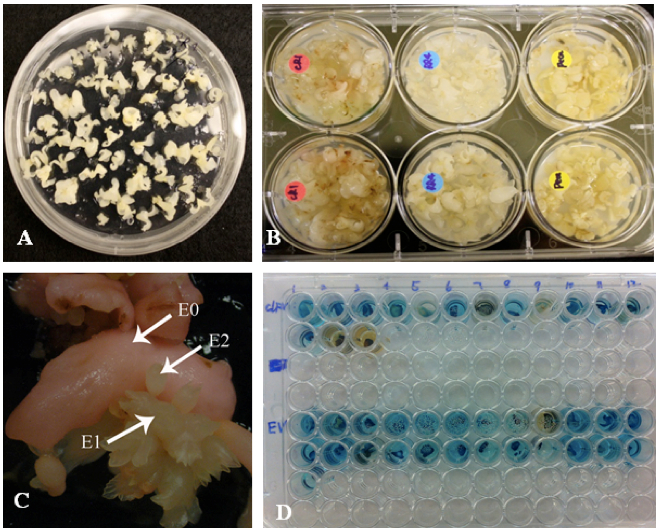

Figure 1. Walnut transformation. A. Embryos with well-develped cotyledens ready for transformation. B. Embyos in co-cultivation suspension. C. Primary embryo (E0) developing secondary embryos (E1 and E2). D. GUS test, transformed embryos turn blue in X-Gluc solution.
- As E2 embryos emerge, test them for GUS (uidA) activity (Jefferson, 1987).
- PCR based verification of transformation
- Select actively proliferating E3 embryos from different embryo lines and use for DNA isolation.
- Isolate total DNA using a DNeasy Plant Mini Kit according to the manufacturer’s protocols.
- Perform PCRs using appropriate primers to confirm the presence of the nptII gene (or other gene of interest). Set up the PCR reactions using 250 ng of genomic DNA in each 25 μl PCR reaction.
- After PCR is completed, 5 μl of PCR product is electrophoresed to verify amplification of an appropriately-sized band. DNA from untransformed embryos can be used as a negative control (Figure 2).
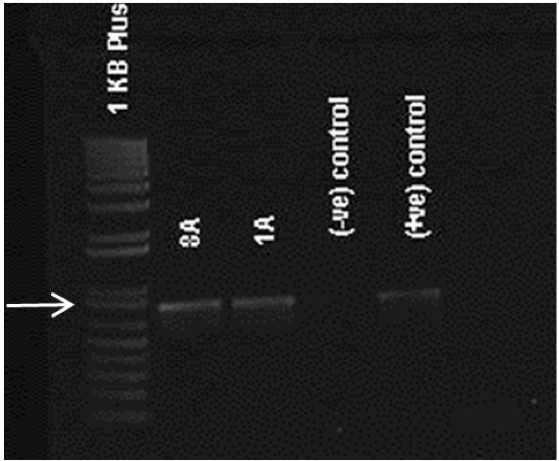
Figure 2. Confirmation of transgenic nature. Agarose gel of PCR products showing 790 bp bands in transformed lines 8A and 1A for APh3/Aph4 primers. Negative controls or non-transformed lines does not show any bands.
- Select actively proliferating E3 embryos from different embryo lines and use for DNA isolation.
- Somatic embryo germination and plant production
- After verification of transformation, some E3 embryos from the desired embryo lines can be desiccated to initiate germination. Choose well-formed somatic embryos and place them in 35 x 10 mm sterile Petri plates with no medium. Cover the plates but leave them unsealed (do not wrap with Parafilm®) and place them in the dark at ambient temperature (20-24 °C) on the rack of a well-sealed desiccator containing 10-15 ml of saturated ZnSO4 or NH4NO3 in the bottom (Figure 3A).
- After the embryos become an opaque white (typically 2-7 days) remove the embryos from the desiccator and place them on DKW shoot medium in Magenta GA-7 vessels or Petri plates.
- Culture at ambient temperature under cool white fluorescent lights (16 h day length, ~100 µE) for 2 to 8 weeks.
- Most embryos will produce roots, but typically fewer than 10% of embryos develop shoots. Roots will usually emerge from embryos in 7-10 days (Figure 3B).
- Fully germinated embryos possessing both roots and shoots should be removed from the medium as soon as possible and planted in potting soil (for example UC Mix, 25%: 42%: 33% sand: fir bark: peat moss). Alternatively, epicotyls can be excised, micropropagated on DKW shoot medium, and then rooted to generate multiple plants.
- To acclimatize, keep plants on soil at 100% humidity for 2 weeks and then gradually reduce the humidity.
- Established plants can be repotted to larger containers as needed and maintained in a greenhouse or lath house.
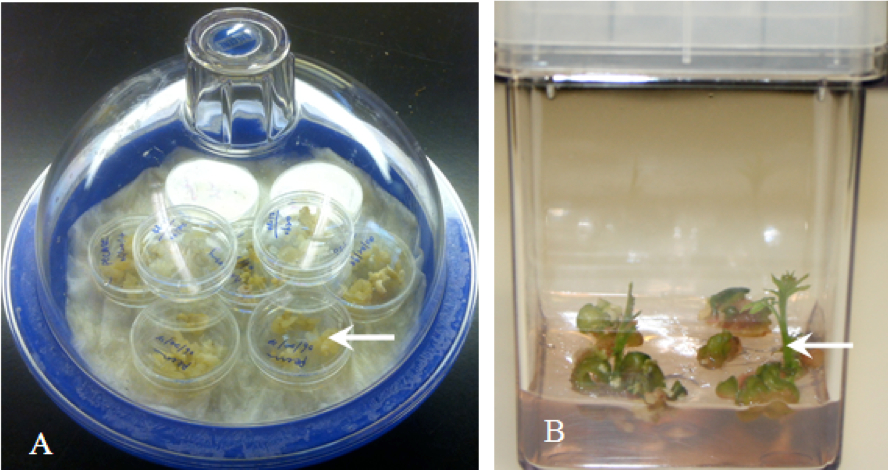

Figure 3. Plant production using trnasgenic somatic embryos. A. Drying embryos in dessicator B. New shoots emerging from transformed walnut somatic embryos.
- After verification of transformation, some E3 embryos from the desired embryo lines can be desiccated to initiate germination. Choose well-formed somatic embryos and place them in 35 x 10 mm sterile Petri plates with no medium. Cover the plates but leave them unsealed (do not wrap with Parafilm®) and place them in the dark at ambient temperature (20-24 °C) on the rack of a well-sealed desiccator containing 10-15 ml of saturated ZnSO4 or NH4NO3 in the bottom (Figure 3A).
Notes
- Physical wounding is not necessary when somatic embryos are used for transformation.
- Transfer embryos in a well-spaced pattern on each plate so that if Timentin-resistant bacteria begin to multiply they are not moved to all the embryos on the plate.
- Hygromycin B (25 mg/L) can serve as an alternative selectable marker in place of kanamycin or as a second selectable marker if performing co-transformation to insert two genes simultaneously.
- Kanamycin sulfate in solution has a very high pH. If used at a concentration greater than 100 mg/L for selection adjust the pH of the kanamycin stock solution to 5.5 with dilute HCl prior to filter sterilizing.
Recipes
- Kanamycin sulfate (50 mg/ml stock)
5 g dissolved in 100 ml dH2O
Filter-sterilize
Stored at -20 °C in 10-15 ml aliquots - Timentin (100 mg/ml stock)
6.2 g dissolved in 62 ml dH2O
Filter sterilize
Stored at -20 °C in 10-15 ml aliquots - 100 mM Acetosyringone
19.6 mg dissolved in 1 ml 95% EtOH
Use capped centrifuge tube and vortex
Parafilm
Stored at room temperature - X-Gluc staining solution
Dissolve X-Gluc to a 0.3% w/v solution in dimethylformamide
Dilute with 100 mM sodium phosphate buffer (pH 7.0) containing 0.006% Triton X-100 and 0.5 mM K+Fe cyanide to make a 1 mM X-gluc working solution
Filter-sterilize
Stored refrigerated - Indole-3- butyric acid (IBA) (0.1 mg/ml stock)
10 mg dissolved in 100 ml
Dissolve IBA powder in a few drops of 1 N KOH
Diluted with dH2O - 6-Benzylaminopurine (BAP) (1 mg/ml stock)
25 mg dissolved in 25 ml
Dissolve BAP powder in a few drops of 1 N KOH
Diluted with dH2O - Media
Driver Kuniyuki walnut (DKW) basal medium
To make 1 L:
5.32 g DKW basal medium with vitamin powder dissolved in dH2O
30 g sucrose
pH to 5.5 using 1 N KOH
2.2 g/L GelzanTM
Autoclave and pour into 40 100 x 15 mm Petri plates - LB liquid medium
To make 100 ml
1 g Tryptone
0.5 g yeast extract
1 g NaCl
pH 6.8-7.2
Autoclave - LB growth plates
To make 100 ml
1 g Tryptone
0.5 g yeast extract
1 g NaCl
0.8 g Bacto® agar
pH 6.8-7.2
Autoclave and pour in 100 x 15 mm Petri plates - Virulence induction medium (IM)
To make 500 ml
2.66 g DKW basal medium with vitamin powder
15 g sucrose
0.5 ml 100 µM acetosyringone (to make 1 µM final concentration)
575 mg proline (to make 1mM final concentration)
pH 5.2 using 1 N KOH
Filter sterilize
Stored refrigerated (4 °C) in 50 ml aliquots - Acetosyringone medium (AS) plates
To make 1 L
5.32 g DKW basal medium with vitamin powder
30 g sucrose
1 ml 100 µM acetosyringone (add this before autoclaving)
pH to 5.5 using 1 N KOH
2.2 g/L GelzanTM
Autoclave and pour in 100 x 15 mm Petri plates - KAN/Timentin selection medium
To make 1 L
5.32 g DKW basal medium with vitamin powder
30 g sucrose
pH 5.5 using 1 N KOH
Dispense into 1 L screw-cap bottles (500 ml per bottle)
1.1 g GelzanTM to each bottle
Autoclave, and cool to 60 °C in a water bath
Then add 200 mg/L pH adjusted filter sterilized kanamycin and 200 mg/L filter sterilized Timentin (see Note 4)
Mix thoroughly and pour into sterile 100 x 15 mm Petri plates.
When solidified, store refrigerated in the original plastic sleeves until ready for use - KAN only selection medium
The same procedure as KAN/Timentin selection medium but without the Timentin - DKW shoot medium
To make 1 L
5.32 g of DKW basal medium with vitamin powder
30 g sucrose
1 mg/L BAP and 0.01 mg/L IBA
pH to 5.5
2.1 g/L Gelzan®
Microwave until the medium boils
Mix thoroughly on a stir plate
Dispense into Magenta Corporation GA7 vessels (approximately 30 ml of medium each)
Autoclave
Acknowledgments
Note that a similar protocol has been described by Leslie et al. (2006) and Dandekar et al. (1989).
References
- Dandekar, A. M., McGranahan, G. H., Leslie, C. A. and Uratsu, S. L. (1989). Agrobacterium-mediated transformation of somatic embryos as a method for the production of transgenic plants. J Tissue Culture Methods 12(4): 145-150.
- Escobar, M. A., Leslie, C. A., McGranahan, G. H. and Dandekar, A. M. (2002). Silencing crown gall disease in walnut (Juglans regia L.). Plant Science 163(3): 591-597.
- Jefferson, R. A. (1987). Assaying chimeric genes in plants: the GUS gene fusion system. Plant Mol Biol Rep 5(4): 387-405.
- Leslie, C. A., Uratsu, S. L., McGranahan, A. M., and Dandekar, G. H. (2006). Methods Mol Biol. In: Wang K. (ed). Agrobacterium protocols 2nd ed, Vol. 2. Humana Press, Vol. 344: 297 -312.
- Mendum, M. L., and McGranahan, G. (1995). Somatic embryogenesis of clonal chandler. Walnut Res Rep, California Walnut Board 25-27.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Walawage, S. L., Leslie, C. A., Escobar, M. A. and Dandekar, A. M. (2014). Agrobacterium tumefaciens-mediated Transformation of Walnut (Juglans regia). Bio-protocol 4(19): e1258. DOI: 10.21769/BioProtoc.1258.
- Araji, S., Grammer, T. A., Gertzen, R., Anderson, S. D., Mikulic-Petkovsek, M., Veberic, R., Phu, M. L., Solar, A., Leslie, C. A., Dandekar, A. M. and Escobar, M. A. (2014). Novel roles for the polyphenol oxidase enzyme in secondary metabolism and the regulation of cell death in walnut. Plant Physiol 164(3): 1191-1203.
Category
Plant Science > Plant transformation > Agrobacterium
Plant Science > Plant developmental biology > General
Molecular Biology > DNA > Transformation
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










