- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Pulse Chase of Suspension Cells
Published: Vol 4, Iss 13, Jul 5, 2014 DOI: 10.21769/BioProtoc.1170 Views: 13348
Reviewed by: Anonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
An NMR Approach for Investigating Membrane Protein–Lipid Interactions Using Native Reverse Micelles
Sara H. Walters and Brian Fuglestad
Jul 20, 2024 2314 Views
A Protocol to Purify Human Mediator Complex From Freestyle 293-F Cells
Hui-Chi Tang [...] Ti-Chun Chao
Feb 20, 2025 2191 Views
Purification of the Active-State G Protein-Coupled Receptor ADGRL4 for Cryo-Electron Microscopy Using a Modular Tag System and a Tethered mini-Gq
David M. Favara and Christopher G. Tate
Mar 5, 2026 250 Views
Abstract
Pulse-chase method is a powerful technique used to follow the dynamics of proteins over a period of time. The expression level, processing, transport, secretion or half-life of proteins can be tracked by metabolically labeling the cells, such as with radiolabeled amino acids (pulse step). This protocol describes the condition used to study the folding and disulfide bond formation of immunoglobulin in suspension cells. With some minor modifications, this protocol can be adapted to study the degradation rate or the secretion of target proteins.
Keywords: Protein foldingMaterials and Reagents
- Pulse chase
- Cells growing in suspension
- HBSS (Life Technologies, InvitrogenTM, catalog number: 14175-095 )
- RPMI without methionine and cysteine (Sigma-Aldrich, catalog number: R7513 )
- Dialyzed FBS (Life Technologies, InvitrogenTM, catalog number: 26400044 )
- N-Ethylmaleimide (NEM) (Sigma-Aldrich, catalog number: R3876 )
- Cyclohexamide (CHX) (Sigma-Aldrich, catalog number: C7698 )
- Express 35S protein labelling mix (Perkin Elmer, catalog number: NEG072014MC )
- Methionine (Sigma-Aldrich, catalog number: M5308 )
- Cysteine (Sigma-Aldrich, catalog number: C7352 )
- Labeling medium (see Recipes)
- Chase medium (see Recipes)
- 2x stop buffer (see Recipes)
- Cells growing in suspension
- Cell lysis and immunoprecipitation
- Antibody against protein of interest
- Protein A/G beads (Thermo Fisher Scientific, catalog number: 20422 )
- Complete Protease Inhibitor Tablets (Roche Diagnostics, catalog number: 11836145001 )
- Lysis buffer (see Recipes)
- Antibody against protein of interest
- SDS-PAGE
- 4-12% Bis-Tris protein gel (Life Technologies, InvitrogenTM)
- MOPS running buffer (Life Technologies, InvitrogenTM, catalog number: NP0001 )
- Amplify solution (GE Healthcare, catalog number: NAMP100 )
- Gel drying solution (Life Technologies, InvitrogenTM, catalog number: LC4025 )
- Gel fixing solution (see Recipes)
- 4-12% Bis-Tris protein gel (Life Technologies, InvitrogenTM)
Equipment
- Incubator
- Eppendorf tube
- 26-gauge needle
- 1 ml syringe
- Heat block
- Gel dryer
- Phosphor imaging screen
Procedure
Cells are pulse-labeled and chased in a single tube and an aliquot of cells is removed from this tube for each time point of the chase.
After determining the numbers of chase time points (x), prepare enough cells for the experiment (x + 1, 2 x 106 per sample). See Note 1.
- Pulse chase
- Wash cells (2 x 106 per sample) with 2 ml of HBSS.
- Pellet cells at 500 x g for 3 min at room temperature. Resuspend cells in 2 ml/sample of pre-warmed labeling medium. Mix gently.
- Incubate the cells for 20 min at 37 ℃ incubator.
- Pellet cells at 500 x g for 3 min at room temperature and resuspend cells in 100 μl/sample of pre-warmed labeling medium. Keep the cells at 37 ℃, either in a water bath or an incubator during the labeling and chase periods.
- Pulse for 2 min at 37 ℃ with [35S] methionine (100 μCi/ml) (see Note 2).
- Add 400 μl/sample of chase medium. Pipet up and down gently to ensure proper mixing.
- Immediately take out 500 μl for the 0 min sample. Transfer to an eppendorf tube filled with 500 μl of 2x ice cold stop buffer (see Note 3).
- Spin cells at 500 x g for 2 min at 4 ℃ and freeze pellet.
- Repeat steps 7-8 for every time point.
- Proceed to cell lysis or keep the pellet frozen in -80 ℃.
- Wash cells (2 x 106 per sample) with 2 ml of HBSS.
- Cell lysis and immunoprecipitation
- Add 1ml cold lysis buffer to each cell pellet.
- Apply mechanical shearing force to the cell lysate by passing it through a 26-gauge needle attached to a 1 ml syringe (repeat 5-10 times for thorough lysis). Incubate on ice for 15 min. Spin at 16,000 x g for 15 min at 4 ℃ to clarify the lysate. Transfer the clarified cell extracts to a new tube.
- Add antibody against protein of interest to equal volume of cell extracts and rotate overnight at 4 ℃ (see Note 4).
- Add 30 μl of Protein A/G beads and incubate for another 2 h at 4 ℃.
- Wash the immunoprecipitates twice with 1 ml of lysis buffer.
- Add 50 μl of sample buffer without reducing agents (such as DTT or 2-mercaptoenthanol). See Note 5.
- Heat the samples 65 °C for 5 min. Centrifuge briefly before proceeding to non-reducing SDS-PAGE.
- Add 1ml cold lysis buffer to each cell pellet.
- SDS-PAGE
- Load 25 μl of the immunoprecipitates on SDS-polyacrylamide gel.
- Immerse the gel in fixing buffer for 15-30 min at room temperature.
- Immerse the gel in amplify solution for 15 min at room temperature.
- Immerse the gel in gel drying solution for another 15 min at room temperature.
- Dry the gel on filter paper on top of a gel dryer of choice.
- [35S] methionine-labeled proteins can be visualized after exposure to a Phosphor imaging screen.
- Load 25 μl of the immunoprecipitates on SDS-polyacrylamide gel.
Representative data
- The protocol described here was used to examine the maturation kinetics of pentameric IgM complexes during their passage through the secretory pathway. IgM assembly begins with the coupling of a heavy chain (H) and a light chain (L), resulting in a monomeric heavy and light chain intermediate (HL). A H2L2 unit is assembled and followed by large multimers of “H2L2”. At later points of the chase time, the signals for these IgM intermediates, especially the high molecular weight species, will decrease due to the successful assembly and secretion of mature IgM.
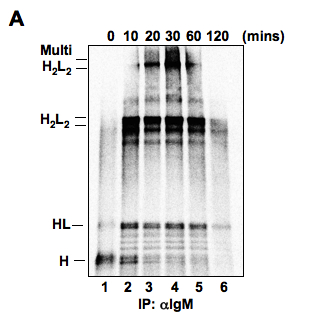
Figure 1. I.29 μ+ mouse lymphoma cells were pulse labeled with 35S-methoninie for 2 min and chased for the indicated times. The cell extracts were immunoprecipitated with αIgM and the immunoprecipitates were resolved by non-reducing SDS/PAGE. IgM assembly intermediates are indicated.
Notes
- In order to ensure equal number of cells are used in each samples, it is necessary to determine the number of chase time points (x) and the volume of chase medium before starting the experiment. We normally prepare enough cells and medium for x (number of chase time points) plus 1 to account for any fluid loss during the experiment.
- The incubation time with radioactive amino acids should be optimized depending on the target protein and the cells used. Subsequently, keeping the labeling step consistent, especially the number of cells and ‘pulse’ time with radioactive amino acids, will reduce variability in the end results.
- Alternatively, the cells can be added directly to ice cold 2x lysis buffer and cell lysis can commence immediately.
- The amount of antibody and cell extracts to be added and the incubation time for antibody-antigen binding should be determined and optimized according to each antibody and antigen. Incubation time can vary from 1 h to overnight at 4 ℃.
- Denaturing sample buffer with reducing agents can be used if disulfide bond formation is not being monitored.
- Dialyzed FBS is used to prevent contamination of ‘cold’ methionine and cysteine. The amount added to the medium should be determined according to the cell line used.
- Cyclohexamide (CHX) is added to reduce the level of newly translated protein. CHX as well as 'cold' methionine and cysteine should be added fresh to the medium.
- NEM is an alkylating agent that covalently attaches to free SH-groups found on cysteines. It is added to prevent disulfide bonds from forming once the chase period has ended. In pulse-chase experiments that track the formation of disulfide bonds, NEM must be added to the stop and lysis buffers.
Recipes
- Labeling medium
RPMI lacking methionine or cysteine
1% Penicilin and Streptomycin
1% glutamine
5-10% dialyzed FBS (see Note 6) - Chase medium (see Note 7)
Labeling medium (see above) plus
5 mM cysteine
5 mM methionine
1 mM CHX - 2x stop buffer
HBSS with 40 mM NEM (see Note 8) - Lysis buffer
50 mM Tris (pH 7.4)
150 mM NaCl
0.5% NP-40
0.5% Na deoxycholate
20 mM NEM
Protease inhibitors - Gel fixing solution
25% methanol
15% acetic acid
Acknowledgments
The protocol presented here was adapted from van Anken et al. (2009). This work was partly supported by grants from the National Institutes of Health (CA082057, CA31363, CA115284, CA180779, AI105809, and AI073099); the Hastings Foundation; the Fletcher Jones Foundation and BaCaTec (J.U.J.). We also thank the Jung laboratory members for their support and discussions.
References
- van Anken, E., Pena, F., Hafkemeijer, N., Christis, C., Romijn, E. P., Grauschopf, U., Oorschot, V. M., Pertel, T., Engels, S., Ora, A., Lastun, V., Glockshuber, R., Klumperman, J., Heck, A. J., Luban, J. and Braakman, I. (2009). Efficient IgM assembly and secretion require the plasma cell induced endoplasmic reticulum protein pERp1. Proc Natl Acad Sci U S A 106(40): 17019-17024.
- Wong L. Y., Brulois K., Toth Z., Inn K. S., Lee S. H., O'Brien K., Lee H., Gao S. J., Cesarman E., Ensser A. and Jung J. U. (2013). The product of Kaposi's sarcoma-associated herpesvirus immediate early gene K4.2 regulates immunoglobulin secretion and calcium homeostasis by interacting with and inhibiting pERP1. J Virol 87(22): 12069-12079.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Wong, L., Liang, Q., Brulois, K. and Jung, J. (2014). Pulse Chase of Suspension Cells. Bio-protocol 4(13): e1170. DOI: 10.21769/BioProtoc.1170.
Category
Immunology > Antibody analysis > Antibody detection
Biochemistry > Protein > Labeling
Biochemistry > Protein > Structure
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link













