- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Analysis of Mycobacterial Protein Secretion
Published: Vol 4, Iss 12, Jun 20, 2014 DOI: 10.21769/BioProtoc.1159 Views: 14173
Reviewed by: Fanglian HeRon Saar-DoverAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

From Llama to Nanobody: A Streamlined Workflow for the Generation of Functionalised VHHs
Lauren E.-A. Eyssen [...] Raymond J. Owens
Mar 20, 2024 6302 Views
Purification of Native Dentilisin Complex from Treponema denticola by Preparative Continuous Polyacrylamide Gel Electrophoresis and Functional Analysis by Gelatin Zymography
Pachiyappan Kamarajan [...] Yvonne L. Kapila
Apr 5, 2024 2121 Views
Thermus thermophilus CRISPR Cas6 Heterologous Expression and Purification
Junwei Wei [...] Yingjun Li
Jul 20, 2025 2202 Views
Abstract
Mycobacterium tuberculosis (Mtb) is the causative agent of tuberculosis. Analysis of proteins secreted by Mtb has been of interest to the field of tuberculosis research since certain secreted proteins interact with the host to promote virulence, while others may be important antigens or serve as biomarkers of infection. Here, we describe a protocol to prepare whole cell extracts (WCE) and short term culture filtrate (CF) from Mtb or the vaccine strain Mycobacterium bovis- bacillus Calmatte- Guérin (BCG) (Mehra et al., 2013). These are both slow growing mycobacteria, but the same basic procedure can easily be adapted to analyze secreted proteins from rapidly growing mycobacteria, such as Mycobacterium smegmatis (Msmeg), a non-pathogenic species commonly used in the laboratory. The fractions obtained can be analyzed by western blotting to examine proteins of interest or by mass spectrometry if antibodies are not available or to examine the entire secretome. Genetic knockout mutants for the gene of interest serve as a negative control. Additionally, levels of a cytosolic protein such as the chaperone GroEL or the pyruvate dehydrogenase E2 component sucB (Rv2215/dlaT) should be assessed in the CF fraction to rule out the possibility that a positive signal in CF is due to bacterial lysis (see Figure 1). By varying the growth conditions of the strain, this in vitro secretion assay can be used to examine conditions that alter the secretome. We are thankful to Magnus Stiegedal for helpful tips on TCA (trichloroacetic acid) precipitation.
Keywords: Mycobacteria culture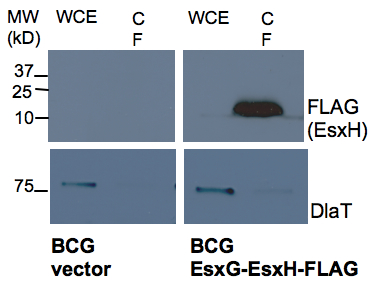
Figure 1. Western analysis of secretion of EsxH by BCG. BCG containing an empty vector control and EsxG-EsxH-FLAG expression construct (FLAG tag at C terminal of EsxH) were analyzed for presence EsxH by anti-FLAG western in WCE and CF prepared as described in the protocol. DlaT was used as a loading control to indicate the degree of bacterial lysis.
Materials and Reagents
Note: All work with live Mtb must be performed in a Biosafety Level 3 (BSL3) facility.
- Middlebrook 7H9 Broth (Difco, catalog number: 271310 )
- Tween-80 (Sigma-Aldrich, catalog number: P4780 )
- Glycerol (Sigma-Aldrich, catalog number: G5516 )
- Albumin-dextrose-catalase (ADC) (BD, catalog number: 212352 )
- Oleic-albumin-dextrose-catalase (OADC) (BD, catalog number: 212351 )
- Potassium phosphate (monobasic) (KH2PO4) (Sigma-Aldrich, catalog number: P9791 )
- L-asparagine monohydrate (Sigma-Aldrich, catalog number: A8381 )
- Citric acid monohydrate (Sigma-Aldrich catalog number: C1909 )
- Ferric ammonium citrate (Sigma-Aldrich catalog number: F5879 )
- Zinc sulfate monohydrate (ZnSO4.H2O) (Sigma-Aldrich, catalog number: 96495 )
- Magnesium sulfate heptahydrate (MgSO4.7H2O) (Sigma-Aldrich, catalog number: 230391 )
- Chelex 100 resin (Bio-Rad Laboratories, catalog number: 142-2822 )
- 100x Halt Protease Inhibitor single use cocktail (Pierce, catalog number: 1860932 )
- 100% trichloroacetic acid (TCA) (pre-chilled prior to use) (Sigma-Aldrich, catalog number: 49010 )
- Acetone (pre-chilled prior to use) (Sigma-Aldrich, catalog number: 32201 )
- Phosphate buffered saline (PBS) (Life Technologies, Gibco®, catalog number: 10010-023 )
- Bromophenol Blue, sodium salt (US Biological, catalog number: 12370 )
- Tris (MP Biomedicals, catalog number: 02194855 )
- Sodium Dodecyl Sulfate (SDS) (US Biological, catalog number: 18220 )
- Ethylenediaminetetraacetic acid (EDTA) (Sigma-Aldrich, catalog number: E6758 )
- β-mercaptoethanol (2-ME) (Sigma-Aldrich, catalog number: M6250 )
- 7H9 complete media (see Recipes)
- Sauton’s media (see Recipes)
- Chelated sauton’s media (an alternative minimal media for mycobacterial growth) (see Recipes)
- Protein extraction buffer (see Recipes)
- 5x SDS-PAGE sample buffer (see Recipes)
Equipment
- Autoclave
- Steriflip-GV filter units (0.22 µM pore size) (Millipore, catalog number: SE1M179M6 )
- 20 ml syringes (BD, catalog number: 302830 )
- 0. 22 µM syringe filter units (33 mm) (Millipore, catalog number: SLGV033RS )
- Disposable Sterile Filter system (1L, 0.22 µm pore size) (Corning, catalog number: 09761104 )
- 0.1 mm zirconia/silica beads (Bio Spec Products, catalog number: 11079101z )
- 30 ml square media bottles (Nalgene®, catalog number: NE/2019-0030 )
- 125 ml square media bottles (Nalgene®, catalog number: NE/2019-0125 )
- 50 ml falcon tubes (Corning, catalog number 430290 )
- 15 ml falcon tubes (Corning, catalog number: 430052 ) with plug seal caps
Note: These 50 ml tubes are compatible with organic solvents and high speed centrifugation. Falcon tubes with these features can be used from different vendors. - Microtubes (2 ml screw cap with O rings) (SARSTEDT AG, catalog number: 72.693 )
- Spectrophotometer
- Centrifuge with swinging bucket rotor for spinning down bacterial cultures (for example, Beckman Coulter, model: Allegra X-15R ; bench top centrifuge with SX4750 rotor)
Notes:- Msmeg and BCG should be handled according to institutional standards of practice for biosafety.
- Mtb cultures should be handled in biosafety level 3 facilities according to institutional standards of practice.
- Centrifuging BCG and Mtb requires appropriate aerosol containment.
- Msmeg and BCG should be handled according to institutional standards of practice for biosafety.
- Beckman Aerosolve® canisters to contain aerosols during centrifugation of mycobacterial cultures (e.g. Beckman Coulter, catalog number: BK359232 )
- 37 °C shaking incubator
- Aerosol containment units for shaking BCG and Mtb liquid cultures in the shaking incubator
Note: Incubator should be placed in BSL3 facility for Mtb cultures. - High speed centrifuge for 50 ml polypropylene falcons used for TCA precipitation of CF (e.g. Beckman Coulter centrifuge with JLA16.2 rotor)
- 50 ml falcon adaptors for rotor JLA16.2
- Bead beater (Bio Spec Products, model: Minibead beater 16; http://www.biospec.com/product/34/mini_beadbeater/)
- Standard table top centrifuge with refrigeration
- Heating block for Eppendorf tubes (set to 95 °C)
- pH meter
Acronyms
- Mtb: Mycobacterium tuberculosis
- BCG: Mycobacterium bovis bacillus Calmatte- Guérin
- Msmeg: Mycobacterium smegmatis
- WCE: Whole cell extract
- CF: Culture filtrate
- DlaT: Rv2215/pyruvate dehydrogenase E2 component sucB protein of mycobacteria
- TCA: Trichloroacetic acid
- BSC: Biosafety cabinet
- OD600: Absorbance or Optical Density at wavelength of 600 nm
Procedure
- Inoculation and growth of mycobacterial cultures
- Inoculate liquid cultures of mycobacteria from frozen stocks, prepared from mid-log phase mycobacterial cultures (OD600 between 0.5 to 1.0) that were frozen in 18% glycerol at -80 °C.
- For inoculation, the doubling rate of the mycobacterial species needs to be kept in mind. Msmeg doubles approximately every 3 h, whereas BCG and Mtb double approximately every 20 h in 7H9 complete media at 37 °C.
- The cultures are inoculated in 10 ml of 7H9 complete media in 30 ml square media bottles. Include appropriate antibiotics if required e.g. antibiotics can be included at appropriate concentration for selecting a plasmid when growing a transformed mycobacterial strain.
- The mycobacterial cultures are incubated at 37 °C with slow shaking at a speed of 90 rpm. BCG and Mtb cultures are placed in aerosol containment in a shaking incubator.
- For Msmeg, inoculate 200 µl from glycerol stock and grow overnight to saturation. Dilute the over-night culture 1:40 into fresh 7H9 complete media and grow to mid-log phase. For BCG and Mtb, inoculate 0.5 ml from frozen stock. It will take several days (~4 days) for the culture to reach mid-log phase and the exact time depends upon the initial inoculum density and growth conditions.
- For inoculation, the doubling rate of the mycobacterial species needs to be kept in mind. Msmeg doubles approximately every 3 h, whereas BCG and Mtb double approximately every 20 h in 7H9 complete media at 37 °C.
- Measure the OD600 of the culture. Determine the culture volume required to give a final starting OD600 of at least 0.25 when resuspended in 25 ml of Sauton’s media [culture volume in ml = 25 ml x desired OD600 (e.g. 0.25)/OD600 of the experimental culture].
- Transfer this volume of culture media to a 15 ml falcon tube and pellet the bacterial cells by centrifuging at 1500 x g for 7 min at room temperature. Remove the supernatant and resuspend the pellet in PBS. Again centrifuge at 1,500 x g for 7 min. This wash step is important to remove albumin carried over from 7H9 complete media.
Notes:- As per our institutional standards of practice for centrifuging Mtb and BCG cultures, falcon tubes are placed in centrifuge adaptors that are put in Aerosolve® canisters (mentioned in Equipment, item 14) inside class II biosafety cabinet (BSC II).
- These are removed from the hood and loaded into the rotor and centrifuged.
- After spin, the whole Aerosolve® canister with the falcon tubes is carried back to the BSC II hood and opened.
- This procedure contains aerosols produced during centrifugation. Mtb cultures should be handled in BSL3 facility.
- As per our institutional standards of practice for centrifuging Mtb and BCG cultures, falcon tubes are placed in centrifuge adaptors that are put in Aerosolve® canisters (mentioned in Equipment, item 14) inside class II biosafety cabinet (BSC II).
- Remove the supernatant and resuspend the bacterial pellet in 1 ml of Sauton’s media. Transfer this to 125 ml square media bottle containing 24 ml of Sauton’s media (include appropriate antibiotics in Sauton’s media as required). Incubate in the shaking incubator at 37 °C.
Notes:- Sauton’s media is a minimal media in which mycobacteria grow more slowly than they do in 7H9 complete media.
- Alternatively, if the effect of metal ions on secretion is to be studied, one can use chelated Sauton’s media as the minimal media and add defined concentrations of metal ions.
- Sauton’s media is a minimal media in which mycobacteria grow more slowly than they do in 7H9 complete media.
- Monitor OD600 of the culture. Grow the culture to between 0.5 to 0.8 OD600. It takes about 6 h for Msmeg and 48 h for BCG and Mtb This procedure could be modified to inoculate the bacteria at a lower density and allow them to grow for a longer period of time, depending upon the goals of the experiment.
- Inoculate liquid cultures of mycobacteria from frozen stocks, prepared from mid-log phase mycobacterial cultures (OD600 between 0.5 to 1.0) that were frozen in 18% glycerol at -80 °C.
- Harvesting the bacterial culture and preparation of the WCE
- After incubating in Sauton’s media as described above, determine the OD600.
- Calculate the volume corresponding to 5 OD600 units. For example, if the OD600 of the culture is 0.5, then 10 ml will correspond to 5 OD units (desired volume = 5/OD600 of the experimental culture).
- Transfer 5 OD600 of the culture into a 15 ml falcon tube (to be used to prepare the WCE). Transfer the remaining culture into a 50 ml falcon tube (to be used to precipitate proteins from the CF). Centrifuge both tubes at 2,000 x g for 5 min at 4 °C.
- Pool the supernatant from both the 15 and 50 ml falcons into one unused 50 ml falcon tube and store on ice until you are ready to process it further (see below, preparation of CF in step C).
- Wash the pellet in the 15 ml falcon with 5 ml of PBS by resuspending and centrifuging at 2,000 x g, 5 min, 4 °C.
- Discard the supernatant and resuspend the pellet again in 1 ml of PBS and transfer it to a 2 ml Sarstedt tube with O ring in the screw cap (for containing aerosols during bead beating).
- Spin on a table top centrifuge at 3,000 x g, 5min and discard the supernatant. To the pellet add 300 µl of protein extraction buffer with protease inhibitors. Resuspend the pellet by pippetting.
- Dispense 100 µl of 0.1 mm zirconia/silica beads into a 1.7 ml eppendorf and add this to the Sarstedt tube containing the pellet.
- Using a bead beater contained within a class II bio-safety cabinet (please read the bead beater manual for the operating procedure and also see the link provided in Equipment, item 19), bead beat the sample for 1 min at 3,450 oscillations/min and place it on ice for 2 min to cool. Repeat this twice. This sample is the WCE. Centrifuge the lysate at full speed for 2 min in a microcentrifuge to settle the froth before opening the tube.
Notes:- Alternate bead beaters may have different speeds and hence the user should optimize bead beating (speed and time) for maximal lysis of mycobacterial pellet.
- The degree of lysis can be established by plating the bacteria before and after bead beating for colony forming units.
- Alternate bead beaters may have different speeds and hence the user should optimize bead beating (speed and time) for maximal lysis of mycobacterial pellet.
- For SDS-PAGE, add 60 µl of 5x SDS-PAGE sample buffer to the Sarstedt tube and then gently mix by inversion. Incubate at 95 °C for 5 min. The heating step helps to denature the sample and also sterilize mycobacterial lysate after bead beating. The mycobacterial WCE can be stored at -80 °C.
Note: The WCE of Mtb can generally be transferred out of BSL3 facility at this step as long as that is consistent with the institutional standards of practice. - Before use, thaw at room temperature and then heat at 95 °C for 5 min. 20 µl of the WCE is often sufficient for SDS-PAGE and western analysis, although this amount will depend upon the target protein levels and the sensitivity of the detection method.
- After incubating in Sauton’s media as described above, determine the OD600.
- TCA precipitation of the CF
- Filter the culture supernatant from step B9 through 0.22 µM filters using a 25 ml syringe into a fresh 50 ml falcon tube.
Notes:- Alternatively, it can be filtered by vacuum using 50 ml Steriflip-GV filter units (0.22 µM) to remove any bacteria from the supernatants.
- This can be done twice depending on the turbidity of the supernatant.
- Double filtration may also be required depending upon the standard operating procedures of the facility to transfer the CF from the BSL3 facility.
- Alternatively, it can be filtered by vacuum using 50 ml Steriflip-GV filter units (0.22 µM) to remove any bacteria from the supernatants.
- Pre-chill trichloroacetic acid (TCA) on ice. Add TCA to a final concentration of 12% to the CF (3 ml of 100% TCA to the 25 ml sample). Invert the tube several times to mix and put the falcon tube at 4 °C overnight. Ensure the falcon tubes used for precipitation are compatible with organic solvents and high speed centrifugation.
- Spin the CF at 15,000 x g for 15 min at 4 °C.
- Discard the supernatant carefully to avoid loosing the pellet.
- Wash the remaining TCA by adding 5 ml acetone (pre-chilled at 4 °C) slowly along the sides of the tube.
- Spin at 15,000 x g for 15 min at 4 °C.
- Discard the supernatant very carefully at this stage since the pellet is not adhered tightly to the tube.
- Leave the tubes open in a Class II BSC for 10-20 min for faster drying of the pellet. Do not over dry to avoid difficulty in solubilizing of the pellet.
- Resuspend the pellet in 150 µl 1x SDS-PAGE sample buffer. You may need to place tubes in a 65 °C water bath briefly to completely resuspend.
- If samples turns yellow due to incomplete removal of TCA, add a few microliters of 1 M Tris-HCl (pH 8) until they turn blue again.
- Store sample at -70 °C. Thaw before use. 10 µl of the sample may be enough for SDS-PAGE and western analysis, although the amount will depend upon the amount of the target protein and the sensitivity of the detection method.
- Filter the culture supernatant from step B9 through 0.22 µM filters using a 25 ml syringe into a fresh 50 ml falcon tube.
Notes
- For applications incompatible with TCA precipitations, Centricon Plus-70 centrifugal filter units can be used to concentrate the CF from step B9 as per the manufacturer’s instructions. This can be followed by buffer exchange to desired buffers. Care must be taken to use Centricons with membranes of molecular weight cut-off less than the molecular weight of the target proteins to be analyzed in the CF.
Recipes
- 7H9 complete media (1 L)
7H9 powder 4.7 g
50% glycerol 4.0 ml
20% Tween 80 2.5 ml
Water to 900 ml
Dissolve 7H9 powder in water and add glycerol and Tween 80
Adjust the amount of water to give a final volume of 900 ml
Add 100 ml of OADC for Mtb or 100 ml of ADC for Msmeg and BCG
Sterilize through 0.22 µM filter and stored at 4 °C (the media can be stored for up to six months. If supplemented with additional additives, like antibiotics or salts, the media should be stored as appropriate for the compounds if they are sensitive to light or degradation)
Note: Alternatively, autoclave after dissolving 7H9 and glycerol in 897.5 ml water and then supplement with sterile tween-80 and 100 ml of ADC/OADC and store at 4 °C as noted above. - Sauton’s media (1 L)
Potassium phosphate, monobasic 0.5 g
L-asparagine monohydrate 4.0 g
Citric acid monohydrate 2.0 g
Ferric ammonium citrate 0.05 g 1% Zinc sulfate monohydrate 0.1 ml
Magnesium sulfate heptahydrate 0.5 g 100% Glycerol 60 ml
Water to 1 L
20% Tween 80 2.5 ml
Dissolve the above components except for the Tween 80 in 900 ml water
Adjust pH to 7 with 5 N NaOH
Make up the volume to 1 L with water
Autoclave, cool, and add 2.5 ml of sterile 20% Tween 80.
Stored at 4 °C as mentioned above for 7H9 complete media - Chelated Sauton’s media (1 L)
Potassium phosphate, monobasic 0.5 g
L-asparagine monohydrate 4.0 g
Citric acid monohydrate 2.2 g
Glycerol 60 ml
20% Tween 80 2.5 ml
Water to 1 L
Chelex 100 resin 10 g
Dissolve the components in water
Adjust the pH to 7.4 with 5 N NaOH and make up the volume to 1 L with water
Add 10 g Chelex 100 resin and stir the media for 1-2 days at room temperature
Sterilize through a 0.22 µM filter unit
Dissolve 1 g of MgSO4.7H2O in 5 ml water and sterilize it by filtration and add to the sterilized media
Stored the media at 4 °C as mentioned above for 7H9 complete media
Defined concentrations of metals like iron, copper and zinc can be added back after chelation. - Protein extraction buffer (10 ml)
1 M Tris-Cl (pH 7.5) 0.5 ml
0.5 M EDTA 0.1 ml
20% SDS 0.3 ml
Water 9.1 ml
This can be prepared and stored at 4 °C.
Immediately prior to use, add Halt Protease Inhibitor cocktail to a final concentration of 1x and keep on ice. - 5x SDS-PAGE sample buffer (50 ml)
0.5 M Tris (pH 6.8) 12.5 ml
SDS 5 g
100% glycerol 25 ml
Bromophenol blue 0.030 g
β- mercaptoethanol 500 µl
Water to 50 ml
Dissolve SDS in Tris buffer and some water, followed by addition of glycerol and bromophenol blue.
Mix and make up the volume to 50 ml with water
Add 500 µl of β- mercaptoethanol
Freeze in aliquots at -20 °C
Acknowledgments
This protocol was originally used in the published work Mehra et al. (2013). This published work was supported by grants and fellowships from the NIH (R01 AI087682), the Doris Duke Charitable Foundation, the Infectious Disease Society of America, the Michael Saperstein Medical Scholars Research Fund (New York University School of Medicine), Potts Memorial Foundation and the American Society of Microbiology.
References
- Mehra, A., Zahra, A., Thompson, V., Sirisaengtaksin, N., Wells, A., Porto, M., Koster, S., Penberthy, K., Kubota, Y., Dricot, A., Rogan, D., Vidal, M., Hill, D. E., Bean, A. J. and Philips, J. A. (2013). Mycobacterium tuberculosis type VII secreted effector EsxH targets host ESCRT to impair trafficking. PLoS Pathog 9(10): e1003734.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Mehra, A. and Philips, J. A. (2014). Analysis of Mycobacterial Protein Secretion . Bio-protocol 4(12): e1159. DOI: 10.21769/BioProtoc.1159.
- Mehra, A., Zahra, A., Thompson, V., Sirisaengtaksin, N., Wells, A., Porto, M., Koster, S., Penberthy, K., Kubota, Y., Dricot, A., Rogan, D., Vidal, M., Hill, D. E., Bean, A. J. and Philips, J. A. (2013). Mycobacterium tuberculosis type VII secreted effector EsxH targets host ESCRT to impair trafficking. PLoS Pathog 9(10): e1003734.
Category
Microbiology > Microbial biochemistry > Protein > Isolation and purification
Microbiology > Microbial cell biology > Cell isolation and culture
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link