- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Isolation of Cells from Human Intestinal Tissue
Published: Vol 4, Iss 7, Apr 5, 2014 DOI: 10.21769/BioProtoc.1092 Views: 17847
Reviewed by: Ivan ZanoniSavita NairAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
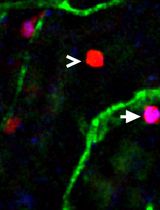
Utilizing EdU to Track Leukocyte Recruitment to the Brain
Zoie K. Lipfert [...] David P. Sullivan
Dec 5, 2025 1600 Views
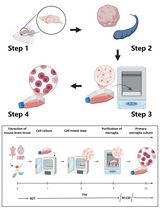
Revisiting Primary Microglia Isolation Protocol: An Improved Method for Microglia Extraction
Jianwei Li [...] Guohui Lu
Dec 5, 2025 1572 Views
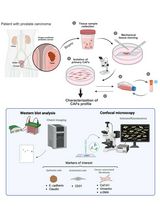
Non-Enzymatic Isolation of Cancer-Associated Fibroblasts From Human Prostate Tumor Explants
Giulia Gangarossa [...] Paola Chiarugi
Mar 5, 2026 148 Views
Abstract
The intestinal lamina propria contains a dense network of T cells, dendritic cells (DCs) and macrophages, which play an important role in local innate and adaptive immune responses. We have recently identified distinct subsets of DCs (Persson et al., 2013) and macrophages (Bain et al., 2013) in the human intestine. In addition, we have studied T cells in healthy and diseased intestine. Here, we describe two methods for isolating these cell populations: 1) enzymatic treatment and 2) migration based isolation. The enzymatic method can be used to isolate T cells, DC and macrophages, whereas the migration based ‘walk-out’ protocol is suitable for DC isolation, as these cells migrate out from the tissues.
Materials and Reagents
- Tissue specimens of small (terminal ileum) and large intestine
- RPMI 1640 (Life Technologies, catalog number: 21875-034 )
- Fetal Bovine Serum (FBS) (Sigma-Aldrich, catalog number: F7424 )
- HEPES (Life Technologies, Gibco®, catalog number: 15630-080 )
- Penicillin and Streptomycin (Life Technologies, catalog number: 15140-122 )
- HBSS (Life Technologies, catalog number: 14180-046 )
- EDTA (Life Technologies, catalog number: AM9261 )
- Liberase TM (Roche Diagnostics, catalog number: 05401127001 )
- DNase I (Sigma-Aldrich, catalog number: D4263 )
- Collagenase 1A (0.2 μm-filtered) (Sigma-Aldrich, catalog number: C9891 )
- FACS antibodies
- T cells
CD3-PE-Cy7 (SK7) (BD biosciences)
Pacific blue (PB)-CD8 (RPA-T8) (BD biosciences)
Quantum dot (QD) 605-CD4 (S3.5) (Life Technologies, InvitrogenTM)
LIVE/DEAD® Fixable Near IR Dead Cell Stain Kit (Life Technologies, InvitrogenTM)
- DC/macrophages
CD3-PE-Cy5 (UCHT1) (eBioscience)
CD19-PE-Cy5 (HIB19) (eBioscience)
CD11c-PE-Cy7 (3.9) (eBioscience)
CD103-PE (Ber-ACT8) or CD103-eFluor647 (B-Ly7) (eBioscience)
CD14-eFluor450 (61D3) (eBioscience)
HLA-DR-APCeFluor780 (L43) (eBioscience)
CD20 PE-Cy5 (2H7) (BioLegend)
TCRab-PE-Cy5 (IP26) (BioLegend)
Biotin-or PE-Cy7 CD172a (SE5A5) (BioLegend)
CD56-PE-Cy5 (Alpha Diagnostic Intl)
Biotin- or FITC-CD11c (MJ4-27G12) (Miltenyi Biotec)
CD141-PE (AD5-14H12) (Miltenyi Biotec)
CD45 V500 (HI30) (BD Biosciences)
Biotinylated antibodies were detected using streptavidin conjugated to PE-Cy7 (eBiosicence) or QDot605 (Life Technologies, InvitrogenTM).
Dead cells were excluded from analysis using propidium iodide PI (Life Technologies, Molecular Probes®).
- T cells
- R10 medium (see Recipes)
Equipment
- Cell strainer (100 μm) (Thermo Fisher Scientific, catalog number: 22363549 )
- 50 ml Falcon tube
- Polyester filters cut in 10 x 10 cm squares (mesh count 27 threads/cm, mesh opening 250 μm, thread diameter 120 μm) (Tekniska Precisionsfilter JR AB)
- Petri dish (SARSTEDT AG, catalog number: 82.1473 )
- Ultra low attachment 24-well culture plate (Sigma-Aldrich, catalog number: CLS3473 )
- Pipette
- Needle/forceps
- Surgical scissor/scalpel
- Beakers with lid (VWR International, catalog number: 216-2694 )
- 37 °C, 5% CO2 cell culture incubator
- Shaker
- Centrifuge
- Microscope
- Multicolour FACS analyser (BD Lsr II flow cytometer)
Software
- FlowJo software (Tree Star Inc)
Procedure
- Enzymatic isolation protocol
- Surgical specimens of human intestine are collected in R10 medium. The tissue is kept on ice and the isolation procedure started optimally within 1 h.
- Underlying muscular layers and fat are removed with scissors and the remaining tissue is cut into small (< 5 mm) pieces with surgical knives (Figure 1). The tissue is placed into a 50 ml Falcon tube. The maximum amount of tissue in one tube should be approximately 4 g.
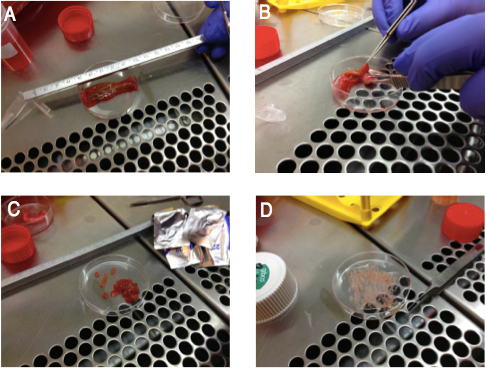

Figure 1. Preparation of terminal ileum for enzymatic or migration based isolation protocols. A) Tissue is placed on a Petri dish and cut longitudinally to expose the mucosa (now facing upwards). B) The soft mucosal layer (containing the immune cells) is held by forceps and the underlying rigid muscular layer together with fat is then removed by cutting with scissors. C) Prior to EDTA treatment, the mucosal layer obtained in B) is cut into approximately 5 mm sized pieces. D) After the EDTA treatment, the tissue is cut into even smaller (1 mm) pieces. These can be then added directly in the bottom of a cell culture well (migration protocol) or in a beaker with enzymes (enzymatic protocol).
- Epithelial cells are removed by incubating tissue fragments in 15 ml of HBSS supplemented with 5% FCS, 10 mM HEPES, 100 U/ml penicillin, 100 μg/ml streptomycin and EDTA (2 mM) for 15 min at 37 °C. After the incubation, the tissue is shaken vigorously by hand for 15 sec (foam will appear) and the epithelial cells in suspension are removed by filtering through a nylon filter. The tissue pieces are collected from the filter and placed again in 50 ml Falcon tubes.
- Repeat step A3 for a total of three times.
- Confirm the denuded appearance of the tissue by light microscopy (Figure 2).
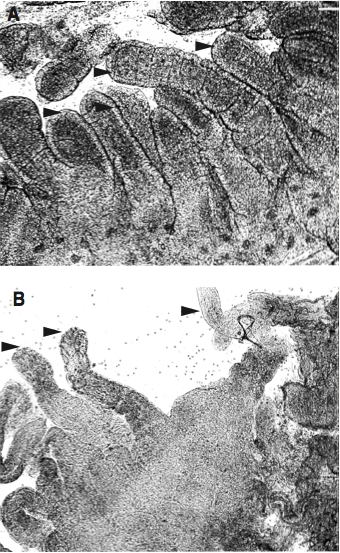
Figure 2. Light microscope images of human terminal ileum A) before and B) after treatment with EDTA to remove epithelial cells. Arrowheads in A) point to numerous villi with the clearly visible epithelial cell layers still intact. In B) the arrow heads point to the missing epithelium. Tiny pieces (1 mm) of intestine were placed directly on glass objective slides together with a drop of R10 medium and a glass coverslip. Images were acquired with a 5x objective and a digital camera attached to the microscope.
- Transfer remaining tissue pieces to a Petri dish. Cut in very fine pieces (< 1 mm) using a needle/forceps and a surgical scissor/scalpel. Look carefully if there are any big pieces left, cut more (Figure 1).
- Transfer the fragments into plastic beakers making sure that all pieces are transferred from the Petri dish.
- Wash the tissue fragments with 20 ml of R10 to remove any remaining EDTA, which may otherwise interfere with enzyme activity in the subsequent step. The pieces will sediment to the bottom of the beaker, remove the R10 with a pipette.
- For isolation of macrophages and DC, the tissue fragments are digested for 2 x 60 min at 37 °C in 10 ml of R10 containing Liberase TM (0.13 Wünsch units/ml) and DNase I (20 IU/ml) with magnetic stirring in plastic beakers with lid. Change the medium and enzymes after the first 60 min and then incubate another 60 min. The cell suspensions are pooled after filtering through 100 µM cell strainers. For T cells, incubate in 20 ml of R10 containing collagenase 1A (1 mg/ml) and DNase I (10 U/ml) with agitation for 60 min. The obtained cell suspension is filtered through 70 µM cell strainer.
- The single cell suspension is washed in 20 ml of R10 and pelleted at 177 x g for 10 min.
- Approximately 4 x 106 T cells, 5,000 CD11c+CD103+ DC and 50,000 CD14+ cells can be isolated from each gram of tissue as calculated by cell sorting (weighed after removal of muscular layer). These numbers are however only an estimate as the donor/preparation variation is high.
- Example of flow cytometry analysis from retrieved cell populations is shown in Figure 3.
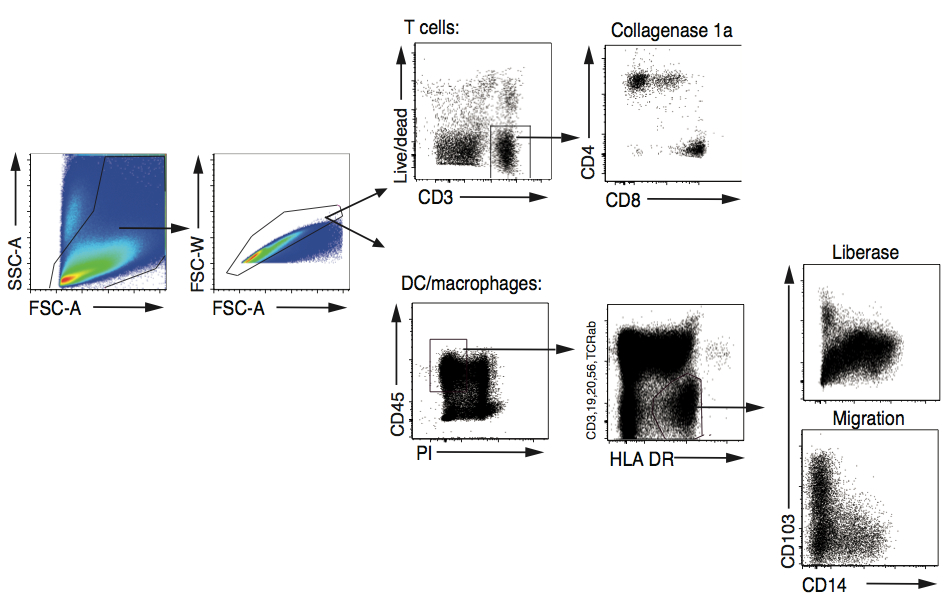
Figure 3. Isolation of human T cells, DC and macrophages. Human intestine was treated with either Collagenase 1A to isolate T cells or Liberase TM to isolate DC macrophages, as described above. The resulting cell suspensions were stained for multicolour FACS analysis. After gating for size and granularity on the FSC/SSC profile, single, live CD3+ cells were gated and analysed for the expression of CD4 and CD8. For DC/macrophage analysis, live CD45+Lin-HLADR+ events were analysed for the expression of CD103 and CD14 showing both CD103+ DC and CD14hi/CD14lo macrophages. Comparison between liberase and migration based isolation method shows less CD14+ macrophages when the cells were allowed to migate out from the tissue. (Lin= CD3, CD19, CD20, CD56, TCRab)
- Surgical specimens of human intestine are collected in R10 medium. The tissue is kept on ice and the isolation procedure started optimally within 1 h.
- Migration based isolation protocol
- The intestine is collected, cleaned and the epithelial cells removed as in the previous section resulting in tiny pieces (< 1 mm) of denuded tissue.
- Approximately 5 pieces of tissue/well are placed in 2 ml of R10 medium in ultra low attachment 24-well culture plates.
- The tissues are incubated for 18 h at 37 °C in 5% CO2 to allow cells to migrate out of the tissue.
- Place the plate on ice for 30 min in order to detach any plate bound cells. The tissue pieces are discarded and the medium containing migrated cells, is collected from the wells and cells pelleted by centrifugation at 177 x g for 10 min at 4 °C.
- Example of staining of the DC subsets is shown in Figure 4.
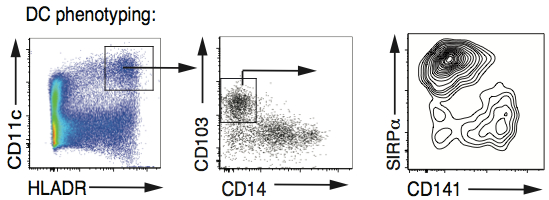
Figure 4. Characterisation of CD103+ DC subsets in the human intestine. DC were allowed to migrate out of human small intestinal preparations as described above. The cells were stained for multicolour FACS analysis with antibodies to CD3, CD19, CD20, CD56, TCRαβ(Lin), CD45, HLA-DR, CD11c, CD14 and CD103. The contour plots show Live CD45+Lin-HLADR+CD11c+CD103+CD14- events. DCs were assessed for expression of SIRPα and CD141.
- The intestine is collected, cleaned and the epithelial cells removed as in the previous section resulting in tiny pieces (< 1 mm) of denuded tissue.
Notes
- Both the enzymatic and migration based protocols can also be applied to small biopsy specimens. In this case, tissue does not need to be further trimmed for removal of fat or size reduction. In addition, the epithelial cell removal step should not include shaking and filtering as the amount of starting material can be very low. Instead, the biopsies are kept in the HBSS/EDTA solution for 45 min at 37 °C with low speed magnetic stirring. After the incubation, discard the medium (and the epithelial cells) and proceed as in step A6. We recommend needles instead of pipette tips for the handling of the tiniest biopsies.
- Collagenase 1A preserves the CD4 epitope on T cells, whereas Liberase cleaves CD4.
- FACS Staining
- Cells were preincubated on ice for 10 min in FACS buffer (PBS/2% FBS) containing 5% mouse serum to prevent unspecific antibody binding.
- Cell staining was performed in 100 μl aliquots for 30 min at 4 °C in the dark, followed by a wash in FACS buffer before being analyzed using an LSR II or FACSAria I and FlowJo software.
- Cells were preincubated on ice for 10 min in FACS buffer (PBS/2% FBS) containing 5% mouse serum to prevent unspecific antibody binding.
Recipes
- R10 medium
RPMI 1640 supplemented with:
10% FBS
10 mM HEPES
100 U/ml penicillin
100 µg/ml streptomycin
50 μg/ml gentamicin
Acknowledgments
This work was supported by grants from the Swedish Medical Research Council, the Göran Gustafsson, Crafoordska, Torsten and Ragnar Söderbergs, Kocks, Österlund, Swartz, Richard and Ruth Julins, and the IngaBritt and Arne Lundbergs Foundations, the Royal Physiographic Society, a clinical grant from the Swedish National Health Service, and the Swedish Foundation for Strategic Research FFL-2 program.
References
- Bain, C. C., Scott, C. L., Uronen-Hansson, H., Gudjonsson, S., Jansson, O., Grip, O., Guilliams, M., Malissen, B., Agace, W. W. and Mowat, A. M. (2013). Resident and pro-inflammatory macrophages in the colon represent alternative context-dependent fates of the same Ly6Chi monocyte precursors. Mucosal Immunol 6(3): 498-510.
- Persson, E. K., Uronen-Hansson, H., Semmrich, M., Rivollier, A., Hagerbrand, K., Marsal, J., Gudjonsson, S., Hakansson, U., Reizis, B., Kotarsky, K. and Agace, W. W. (2013). IRF4 transcription-factor-dependent CD103(+)CD11b(+) dendritic cells drive mucosal T helper 17 cell differentiation. Immunity 38(5): 958-969.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Uronen-Hansson, H., Persson, E., Nilsson, P. and Agace, W. (2014). Isolation of Cells from Human Intestinal Tissue. Bio-protocol 4(7): e1092. DOI: 10.21769/BioProtoc.1092.
Category
Immunology > Immune cell isolation > Maintenance and differentiation
Cell Biology > Cell isolation and culture > Cell isolation
Cell Biology > Cell movement > Cell migration
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










