- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Flow Cytometric Analysis of Autophagic Activity with Cyto-ID Staining in Primary Cells
Published: Vol 4, Iss 7, Apr 5, 2014 DOI: 10.21769/BioProtoc.1090 Views: 27571
Reviewed by: Anonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Monitoring of Sperm-Independent Calcium Oscillations in Immature Oocytes of Mice
Sae Horiike [...] Hidehiko Ogawa
Feb 5, 2026 245 Views
Time-Lapse Into Immunofluorescence Imaging Using a Gridded Dish
Nick Lang [...] Andrew D. Stephens
Feb 20, 2026 238 Views

How to Train Custom Cell Segmentation Models Using Cell-APP
Anish J. Virdi and Ajit P. Joglekar
Feb 20, 2026 344 Views
Abstract
Flow cytometry allows very sensitive and reliable high-throughput analysis of autophagic flux. This methodology permits to screen cells in flow and capture multi-component images. Using this technology autophagic flux may be analysed accurately in both suspension as well as adherent cells upon trypsinization independent of how heterogeneous the autophagosomal content might be. The method is based on Cyto-ID staining of autophagic compartments (pre-autophagosomes, autophagosomes, and autophagolysosomes) in live cells using Cyto-ID® Autophagy Detection Kit. Autophagic compartments are intermediate constituents of a dynamic lysosomal degradation process and their intracellular abundance at a particular time point is a function of the established equilibrium between their generation and degradation. Determination of autophagic flux facilitates the discrimination between early induction of autophagosome formation and late inhibition of autophagosome maturation as both results in an ultimate increase in autophagosomal presence. Cyto-ID assay is based on the usage of a specific dye that selectively stains autophagic compartments and therefore allows determination of autophagic flux as accumulation of stained compartments in basic or activated conditions [rapamycin (1-5 µmol/L), PP242 (1-5 µmol/L) or Hanks’ Balanced Salt Solution containing 6 mmol/L glucose (starvation medium)] after blockage of autophagolysosomal degradation using lysosomotropic compounds such as ammonium chloride (NH4Cl) (10-20 mmol/L) or chloroquine (CQ) (5-10 µmol/L). ΔMFI Cyto-ID = MFI Cyto-ID (+CQ/NH4Cl) - MFI Cyto-ID (-CQ/NH4Cl).
Materials and Reagents
- Cells lines of interest (HepG2, HUH7, CMK, K562 etc.) or primary cells [for example: murine bone marrow-derived dendritic cells (BMDCs)]
- Cyto-ID® Autophagy Detection Kit (Enzo Life Sciences, catalog number: ENZ-51031-K200 )
- Eagle's minimal essential medium (EMEM) (ATCC, catalog number: 30-2003 ) containing 10% fetal bovine serum (FBS) (Biochrom, catalog number: S0615 ) with 100 U/100 μg/ml penicillin/streptomycin (Life Technologies, Gibco®, catalog number: 15140-122 )
- RPMI 1640 with L-glutamine (Lonza, catalog number: BE12-702F ) containing 10% FBS with 100 U/100 μg/ml penicillin/streptomycin
- DMEM (low glucose, pyruvate, no glutamine, no phenol red) (Life Technologies, Gibco®, catalog number: 11880-028 )
- Dulbecco’s Phosphate Buffered Saline (DPBS) (Biochrom, catalog number: L1825 )
- 1x 0.05% Trypsin-EDTA (phenol red) (Life Technologies, catalog number: 25300 )
- Hanks Balanced Salt Solution (HBSS) (Life Technologies, Gibco®, catalog number: 14025 ) containing 6 mM glucose (starvation medium)
- Rapamycin from Streptomyces hygroscopicus (1-5 µmol/L) (Sigma-Aldrich, catalog number: R0395 )
- PP242 hydrate (1-5 µmol/L) (Sigma-Aldrich, catalog number: P0037 )
- 3-methyladenine (3-MA) (3-10 mmol/L) (Sigma-Aldrich, catalog number: M9281 )
- Wortmannin (30-100 nmol/L) (Sigma-Aldrich, catalog number: W3144 )
- LY294002 (7-20 µmol/L) (Sigma-Aldrich, catalog number: L9908 )
- Nocodazole (12-50 µmol/L) (Sigma-Aldrich, catalog number: M1404 )
- Vinblastine (12-50 µmol/L) (Sigma-Aldrich, catalog number: V1377 )
- Ammonium chloride (NH4Cl) (10-20 mmol/L) (Sigma-Aldrich, catalog number: A0171 )
- Hydrohychloroquine sulphate (HCQ) (5-10 µmol/L) (Sigma-Aldrich, catalog number: H0915 )
- Chloroquine (CQ) (5-10 µmol/L) (Sigma-Aldrich, catalog number: C6628 )
- Dimethyl sulfoxide DMSO (Sigma-Aldrich, catalog number: D8418 )
Equipment
- 37 °C, 5% CO2 humidified incubator
- Centrifuge
- FACSCalibur, LSR II (BD) or analogous equipment
Procedure
- Maintain cells under standard tissue culture conditions at 37 °C, 5% CO2 in a humidified incubator. Keep cell density below 1 x 106/ml.
Caution: Prior to analysis cell should be kept for several hours (min 12 h) in fresh medium to avoid potential activation of autophagy due to nutrients exhaustion. Generally culture medium contains autophagy affecting substances: amino acids, glucose, growth factors, hormones etc. Take care when comparing autophagic flux in different conditions to normalize for all the necessary factors. Normalize also for the solvent used when analysing the effect of different substances on autophagy – for example DMSO, ethanol etc. might affect autopagic flux.
- Incubate the cells for the desired time and under the conditions of interest. Negative control subjected to vehicle treatment should be included.
Caution: When analysing prolonged periods of time and/or under conditions potentially affecting cells numbers or viability, differences in nutrient consumption and therefore abundance might occur and influence your results as autophagic activity is highly related to the nutritional status.
- Positive control [rapamycin (1-5 µmol/L), PP242 (1-5 µmol/L), Hanks’ Balanced Salt Solution containing 6 mmol/L glucose (starvation medium)] and negative control [3-methyladenine (3-MA) (3-10 mmol/L), wortmannin (30-100 nmol/L), LY294002 (7-20 µmol/L), nocodazole (12-50 µmol/L), vinblastine (12-50 µmol/L), ammonium chloride (NH4Cl) (10-20 mmol/L), hydrohychloroquine (HCQ) or chloroquine (CQ) (5-10 µmol/L)] may be also included.
- Upon completion of treatment period, wash the cells with DPBS. For suspension cells use centrifugation, for adherent cells wash as they are adherent (do not trypsinize).
- Resuspend suspension cells, or add to adherent cells Cyto-ID Green containing indicator free cell culture medium, containing 5% FBS. Suggested cell density for staining stage is 105 to 106 cells/ml. Suggested Cyto-ID Green concentration is 1 μl of Cyto-ID Green Detection Reagent in 1 ml cell culture medium.
- Mix well and incubate for 30 min under standard tissue culture conditions at 37 ºC, 5% CO2 in the dark.
- At the end of staining procedure, wash away the Cyto-ID containing medium. Wash once more with DPBS and resuspend suspension cells or trypsinize, wash and resuspend adherent cells in ice cold DPBS containing 2% FBS.
Note: Caution upon completion of staining step whenever possible keep always on ice and in the dark.
- Analyze samples using flow cytometer and plot data of cell counts as FITC (FL1) fluorescence intensity. Table 1 may be used to facilitate the interpretation of the results.
Table 1. Analysis of unknown substance XX plus
Medium
Activator
I
→
↑↑
II
↑↑
↑↑↑
III
↓↓
↓ or ↑
Notes:- →↑↓: ΔMFI Cyto-ID = MFI Cyto-ID (+CQ/NH4Cl) - MFI Cyto-ID (-CQ/NH4Cl).
CQ/NH4Cl stays for either CQ or NH4Cl.
Caution: The calculation of ΔMFI Cyto-ID requires both +CQ/NH4Cl and -CQ/NH4Cl condition for each individual sample!
- Suggested incubation with ammonium chloride (NH4Cl) (10–20 mmol/L) or chloroquine (CQ) (5–10 µmol/L) is 5 h.
- (Activator): autophagy induction [rapamycin (1-5 µmol/L), PP242 (1-5 µmol/L) or starvation medium]. Suggested incubation 5 h. (Caution: The concentration of the activator should be established for the particular cell type.)
- Analysis of unknown substance X should include co incubation with one or more activators. Table 1 illustrates three possible scenarios. a) Scenario (I) unknown substance X does not impact cellular autophagic activity; b) Scenario (II) X is an activator; c) Scenario (III) X is an inhibitor of autophagy.
- Caution: Although considered the most accurate sensitive and reliable method for analysis of autophygic flux, flow cytometry determination of Cyto-ID stained cells should be combined with alternative methods with non overlapping limitations such as electron microscopy (EM) fluorescence microscopy (FM), western blotting (WB) etc.
- Example data:
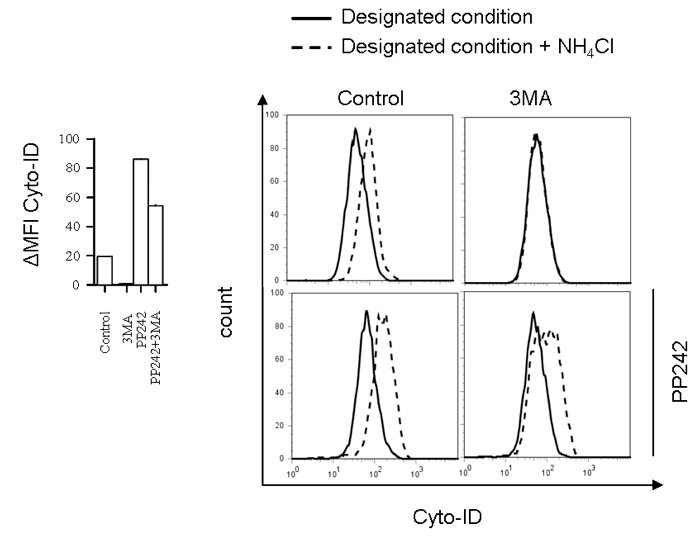
Figure 1. Flow cytometry analysis of autophagic flux in BMDC incubated with autophagy inhibitor 3MA (10 mM) for 10 h in the presence or absence of autophagy activator PP242 for the last 5 h (left) with representative histograms (right)
- →↑↓: ΔMFI Cyto-ID = MFI Cyto-ID (+CQ/NH4Cl) - MFI Cyto-ID (-CQ/NH4Cl).
Acknowledgments
Stankov, M and Behrens, G were supported by the DFG (BE-2089/2-1 and EXC62/1). Klusmann, J was supported by a grant from the Wilhelm Sander-Foundation (2011.057.1), the German Research Foundation (DFG; KL-2374/1-1). Klusmann, J is a fellow of the Emmy Noether-Programme from the DFG (KL-2374/2-1). Leverkus, M by the DFG (LE-953/8-1 and LE-953/6-1) and the Mildred-Scheel-Stiftung (#109891).
References
- Cyto-ID® Autophagy Detection Kit Manual (Enzo Life Sciences). http://www.enzolifesciences.com/fileadmin/enzo/Manuals/51031-K200.pdf.
- Mizushima, N., Yoshimori, T. and Levine, B. (2010). Methods in mammalian autophagy research. Cell 140(3): 313-326.
- Sinicrope, F. A., Sirko, A., Siu, P. M. et al. (2012). Guidelines for the use and interpretation of assays for monitoring autophagy. Autophagy 8(4): 445-544.
- Stankov, M. V., El Khatib, M., Kumar Thakur, B., Heitmann, K., Panayotova-Dimitrova, D., Schoening, J., Bourquin, J. P., Schweitzer, N., Leverkus, M., Welte, K., Reinhardt, D., Li, Z., Orkin, S. H., Behrens, G. M. and Klusmann, J. H. (2014). Histone deacetylase inhibitors induce apoptosis in myeloid leukemia by suppressing autophagy. Leukemia 28(3): 577-588.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Stankov, M., Panayotova-Dimitrova, D., Leverkus, M., Klusmann, J. and Behrens, G. (2014). Flow Cytometric Analysis of Autophagic Activity with Cyto-ID Staining in Primary Cells. Bio-protocol 4(7): e1090. DOI: 10.21769/BioProtoc.1090.
Category
Microbiology > Antimicrobial assay > Autophagy assay
Cell Biology > Cell-based analysis > Flow cytometry
Cell Biology > Cell imaging > Live-cell imaging
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










