- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Longitudinal Bioluminescent Quantification of Three Dimensional Cell Growth
Published: Vol 3, Iss 23, Dec 5, 2013 DOI: 10.21769/BioProtoc.993 Views: 9481
Reviewed by: Lin Fang

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
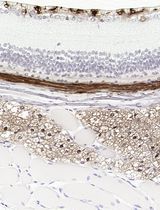
Improved Immunohistochemistry of Mouse Eye Sections Using Davidson's Fixative and Melanin Bleaching
Anne Nathalie Longakit [...] Catherine D. Van Raamsdonk
Nov 20, 2025 1565 Views
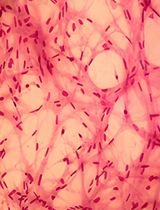
A Simplified 3D-Plasma Culture Method for Generating Minimally Manipulated Autologous Equine Muscle-Derived Progenitor Cells
Hélène Graide [...] Didier Serteyn
Dec 5, 2025 1251 Views
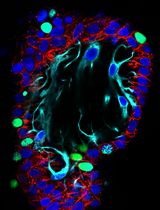
Simple and Rapid Model to Generate Differentiated Endometrial Floating Organoids
Adriana Bajetto [...] Tullio Florio
Feb 5, 2026 92 Views
Abstract
The use of three-dimensional (3D) cell culture systems is widely accepted as representing a more physiologically relevant means to propagate mammary epithelial and breast cancer cells. However, 3D cultures systems are plagued by several experimental and technical limitations as compared to their traditional 2D counterparts. For instance, quantifying the growth of mammary epithelial or breast cancer organoids longitudinally is particularly troublesome using standard [3H]thymidine or MTT assay systems, or using computer-assisted area calculations. Likewise, the nature of the multicellular aggregates and organoids formed by breast cancer cells under 3D conditions precludes efficient recovery of the cells from 3D matrices, an event that is time consuming and leads to spurious results. The assay described here utilizes stable expression of firefly luciferase as means to quantify the longitudinal outgrowth of cells propagated within a 3D matrices. The major advantages of this technique include its high-throughput nature and ability to longitudinally track single wells over a defined period of time, thereby decreasing the costs associated with assay performance. Finally, this technique can be readily combined with drug treatments and/or genetic manipulations to assay their effects on the growth of 3D organoids.
Materials and Reagents
- Murine 4T1 mammary carcinoma cells (ATCC, catalog number: CRL-2539 ) or any cell line of interest engineered to stably express firefly luciferase under the control of a constitutively-active promoter such cytomegalovirus.
Note: Several Luciferase encoding plasmids are commercially available and typically employ pcDNA3.1- or pBabe-based backbones to deliver firefly or renilla luciferases. In either scenario, antibiotic administration is used to select and maintain stable expression of luciferase in reporter cells.
- Cultrex: Reduced growth factor (RGF) basement membrane extract (BME) (Trevigen, catalog number: 3433-005-01 )
- Ice
- Dulbecco’s Modified Eagle Medium (DMEM) (Life Technologies, catalog number: 10313-021 )
- Penicillin/Streptomycin (Pen/Strep) (Gibco®, catalog number: 15140 )
- Fetal bovine serum (Sigma-Aldrich, catalog number: F1051 )
- D-luciferin, Potassium Salt (15 mg/ml) in sterile H2O (Gold Bio, catalog number: LUCK-1G )
Equipment
- 2D culture dishes
- White walled, clear bottom 96-well culture dishes (Corning, Costar®, catalog number: 3610 )
- Luminometer capable of reading 96-well plate format (Promega GloMax-Multi Detection System or similar bioluminescent plate reader).
- Hemocytometer or other means of cell counting
- 37 °C/5% CO2 cell incubator
Procedure
- Cells are grown in DMEM supplemented with 10% FBS and Pen/Strep (full growth media).
- Cells are harvested from actively proliferating, sub-confluent 2D culture dishes.
- Cells are trypsinized, washed in excess full-growth media, and pelleted by gentle centrifugation. Afterward, the resulting cell pellets are resuspended and allowed to recover in full growth media for 2 h.
- During this time, coat the 96-well dish with 50 μl of 100% Cultrex per well and allow to gel at 37 °C.
Note: Cultrex must be maintained on ice at all times to prevent solidification of the matrices that transpires as the gel warms to >4 °C.
- Count the cells using the hemocytometer and dilute them to 1,000 cells in 150 μl of full growth media, which is supplemented with 4% Cultrex. Thoroughly mix cells and media/4% Cultrex solution and subsequently plate the cells on top of the solidified Cultrex cushions.
Note: The 4% Cultrex solutions do not solidify, and as such, cells contained within these mixtures will readily attach to solidified Cultrex cushions, thereby permitting top layer media changes and/or replacements throughout the experiment.
- Prior to plating the cells, other compounds such as growth factors or chemical inhibitors may be mixed with the cells and 4% Cultrex solutions. Each experimental condition should be plated in triplicate.
Note: All test compounds must be screened initially by short-term exposure to verify that these agents do not directly impact the expression of CMV-driven luciferase and/or the activity of luciferase:luciferin reactions.
- Two hours after plating the cells, obtain initial time zero (T0) luminescence readings by adding 2 μl of D-luciferin (15 mg/ml) and gently tap the side of the plate to mix.
Note: Culture lid is open while in the plate reader, and as such, it is imperative that the plate reader remain clean and well-sanitized, and potentially be located in a laminar flow hood to prevent unwanted cell contamination. Culture medium does not need to be replaced at this time.
- Place culture in a 37 °C incubator with 5% CO2.
- Four days after plating, multicellular organoids will have begun to form. Obtain a second luminescence reading as described in step 7. Afterward, change and refresh all media and experimental test components, being extremely careful not to disrupt the cells and solidified Cultrex cushion.
Note: At this point, therapeutic protocols can be initiated to monitor the anticancer activities of various chemotherapies against established organoids.
- For all experimental conditions, repeat steps 7-9 at days 8 and 11 post-plating.
Note: These time points may vary depending dramatically based on the relative growth characteristics of the cell line under study. As such, longitudinal luciferase readings, dosing regiments, and cell plating densities need to be determined empirically for each cell line to be studied to maximize signal-to-noise ratios.
- Use Excel to normalize each luminescence value to the initial T0 value derived after plating cells.
Acknowledgments
We thank the members of the Schiemann laboratory for their helpful comments and suggestions. Bioluminescent 3D-organotypic longitudinal growth assays were adapted from Wendt et al. (2011) and Wendt et al. (2013). Research support was provided in part by the National Institutes of Health to M.K.W. (CA166140), and to W.P.S. (CA129359 and CA177069).
References
- Wendt, M. K., Schiemann, B. J., Parvani, J. G., Lee, Y. H., Kang, Y. and Schiemann, W. P. (2013). TGF-beta stimulates Pyk2 expression as part of an epithelial-mesenchymal transition program required for metastatic outgrowth of breast cancer. Oncogene 32(16): 2005-2015.
- Wendt, M. K., Taylor, M. A., Schiemann, B. J. and Schiemann, W. P. (2011). Down-regulation of epithelial cadherin is required to initiate metastatic outgrowth of breast cancer. Mol Biol Cell 22(14): 2423-2435.
Article Information
Copyright
© 2013 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Wendt, M. K. and Schiemann, W. P. (2013). Longitudinal Bioluminescent Quantification of Three Dimensional Cell Growth. Bio-protocol 3(23): e993. DOI: 10.21769/BioProtoc.993.
Category
Cancer Biology > General technique > Cell biology assays > Proliferation analysis
Cell Biology > Cell isolation and culture > 3D cell culture
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










