- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Extraction and Quantification of Cyclic Di-GMP from Pseudomonas aeruginosa
Published: Vol 3, Iss 14, Jul 20, 2013 DOI: 10.21769/BioProtoc.828 Views: 15026
Reviewed by: Fanglian He

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

β-lactamase (Bla) Reporter-based System to Study Flagellar Type 3 Secretion in Salmonella
Fabienne F. V. Chevance and Kelly T. Hughes
Jun 20, 2023 1764 Views
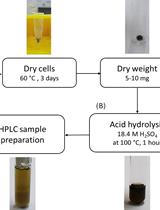
Determination of Poly(3-hydroxybutyrate) Content in Cyanobacterium Synechocystis sp. PCC 6803 Using Acid Hydrolysis Followed by High-performance Liquid Chromatography
Janine Kaewbai-ngam [...] Tanakarn Monshupanee
Aug 20, 2023 1808 Views
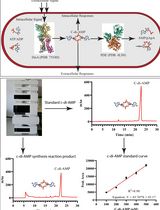
An HPLC-based Assay to Study the Activity of Cyclic Diadenosine Monophosphate (C-di-AMP) Synthase DisA from Mycobacterium smegmatis
Avisek Mahapa [...] Dipankar Chatterji
Dec 20, 2024 1779 Views
Abstract
Cyclic di-GMP (c-di-GMP) has emerged as an important intracellular signaling molecule, controlling the transitions between planktonic (free-living) and sessile lifestyles, biofilm formation, and virulence in a wide variety of microorganisms. The following protocol describes the extraction and quantification of c-di-GMP from Pseudomonas aeruginosa samples. We have made every effort to keep the protocol as general as possible to enable the procedure to be applicable for the analysis of c-di-GMP levels in various bacterial species. However, some modifications may be required for the analysis of c-di-GMP levels in other bacterial species.
Keywords: HPLCMaterials and Reagents
- Bacterial culture
- Ethanol (95-100%)
- Tris base
- EDTA
- NaCl
- KCl
- Na2HPO4.7H2O
- KH2PO4 (pH 7.2)
- 0.5 M Ethylenediaminetetraacetic acid (EDTA) disodium salt solution (pH 8.0)
- Bis-(3'-5')-cyclic diguanylic monophosphate (c-di-GMP) (Biolog)
- Ammonium acetate (MS grade)
- Methanol (HPLC grade)
- Nanopure water (18 Ohm)
- Protein determination reagents
- Phosphate-buffered saline (PBS) (see Recipes)
- HPLC Solvent A (see Recipes)
- HPLC Solvent B (see Recipes)
- TE buffer (see Recipes)
Equipment
- Syringes (1 ml)
- Syringe filters (2 μm) (Upchurch Scientific, catalog number: B-100 )
- Microfuge tubes (unpolished, to reduce static)
- Spectrophotometer
- Refrigerated microcentrifuge
- Vacuum concentrator/Centrifugal evaporator (e.g. SpeedVac)
- Reverse-phase C18 Targa column (2.1 x 40 mm, 5 μm) (The Nest Group, catalog number: TR-0421-C185 )
- Heat block or water bath
- High performance liquid chromatography (HPLC) system and software for HPLC peak analysis (e.g. Agilent 1100 HPLC)
- Sonicator
- Homogenizer
Software
- ChemStation for LC (Agilent Technologies)
Procedure
- Extraction of c-di-GMP
- Grow bacterial cells to desired growth stage under required experimental conditions. Proceed directly with the extraction, with no waiting periods or incubation of cells on ice, as this may drastically alter the c-di-GMP levels.
Note: Extraction at mid-exponential phase is recommended, as using early exponential phase cells will yield c-di-GMP levels too low for accurate detection. To extract c-di-GMP from planktonic cells, allow P. aeruginosa to grow in either Lennox broth or Vogel-Bonner minimal medium (VBMM) for 6 h to mid-exponential phase in flasks at 37 °C and 220 rpm. Inoculate using a 5% inoculum size of an overnight culture. Biofilm c-di-GMP levels can be determined from biofilms grown for 3-5 days under flowing conditions; see References 2-4 for more detail. - Determine the optical density of the bacterial culture at 600 nm (OD600).
Note: If working with biofilm samples, include a step to homogenize (10 sec on high) the cultures to disrupt cell aggregates prior to OD600 determination. - Obtain a bacterial culture volume equivalent to 1 ml of OD600 = 1.8 (e.g. If the OD600 = 0.9, spin down 2 ml of culture).
Note: This biomass has been optimized for the analysis of c-di-GMP in P. aeruginosa strains PAO1 and PA14 planktonic and biofilm samples. Analysis of c-di-GMP levels in other strains or species may require the initial biomass harvested for extraction to be adjusted. - Centrifuge (16,000 x g, 2 min, 4 °C) the respective culture volume. Discard the supernatant.
- Wash the cell pellet with 1 ml ice-cold PBS (16,000 x g, 2 min, 4 °C). Discard the supernatant.
- Repeat step A-5.
- Resuspend the cell pellet in 100 μl ice-cold PBS and incubate at 100 °C for 5 min.
- Add ice-cold ethanol (stored at -20 °C until use) to a final concentration of 65% (186 μl of 100% ethanol or 217 μl of 95% ethanol) and vortex for 15 sec.
- Centrifuge sample (16,000 x g, 2 min, 4 °C), and remove and retain the supernatant containing extracted c-di-GMP in a new microfuge tube. Store the supernatant on ice or at -80 °C until step A-10. Retain the cell pellet.
- Using the cell pellet, repeat twice the extraction procedure in steps A-7~A-9. Pool the supernatants obtained from the three extractions into one microfuge tube. Retain the cell pellet after the final extraction step. The cell pellet can be stored at -20 °C until step B-3-a.
- Dry the combined supernatants using a vacuum concentrator/centrifugal evaporator. Following evaporation, a white pellet should be visible. This sample, containing the extracted c-di-GMP, can be stored at -80 °C until step B-2-a.
- Grow bacterial cells to desired growth stage under required experimental conditions. Proceed directly with the extraction, with no waiting periods or incubation of cells on ice, as this may drastically alter the c-di-GMP levels.
- Quantification of c-di-GMP
This procedure has been optimized for the detection of c-di-GMP using an Agilent 1100 HPLC equipped with an autosampler, degasser, pressure regulator, prefilter, and UV/Vis detector set to 253 nm. Separation was carried out using a reverse-phase C18 Targa column (2.1 x 40 mm; 5 μm) and a flow rate of 0.2 ml/min. Solvents containing methanol and ammonium acetate (see Recipes for solvents A and B) were used. The following gradient was used to elute c-di-GMP: 0 to 9 min, 1% B ( = 1% solvent B and 99% solvent A); 9 to 14 min, 15% B; 14 to 19 min, 25% B; 19 to 26 min, 90% B; 26 to 40 min, 1% B. This gradient resulted in the elution of c-di-GMP at approximately 14-15 min.
Note: Analysis of c-di-GMP levels using a different reverse-phase column, flow rate, or HPLC system may require the optimization of HPLC separation gradients.- Generation of a standard curve
- Using commercially available c-di-GMP, prepare the following standards in nanopure water: 1, 2, 5, 10 and 20 pmo/μl.
- To generate the standard curve, inject 20 μl per standard per HPLC run. Use 20 μl of nanopure water as a negative control (0 pmol μl-1 c-di-GMP). For an example of c-di-GMP detection (Figure 1).
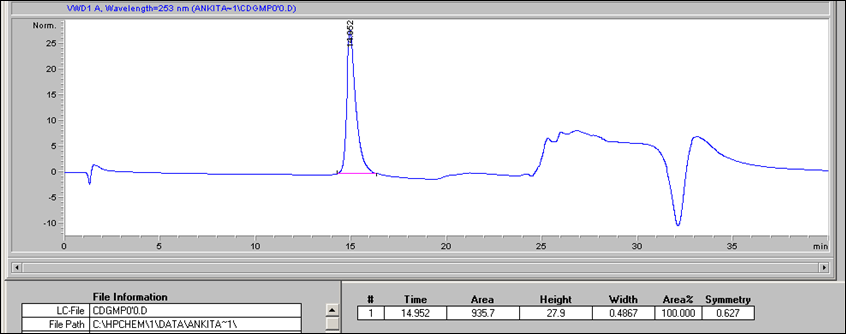
Figure 1. Example of c-di-GMP standard elution profile. 20 μl of a 10 pmo/μl c-di-GMP standard (200 pmol total) were separated using a reverse-phase C18 Targa column (2.1 x 40 mm; 5 μm) at a flow rate of 0.2 ml/min with the following gradient: 0 to 9 min, 1% B; 9 to 14 min, 15% B; 14 to 19 min, 25% B; 19 to 26 min, 90% B; 26 to 40 min, 1% B. Peak at 15 min corresponds to the elution of c-di-GMP. The c-di-GMP peak was found to have an area of 935.7. - Prepare a standard curve by plotting the c-di-GMP amount in pmol (e.g. 200 pmol for the 20 μl of the 10 pmol/μl standard) vs the peak areas. An example of a standard curve is given in Figure 2. Peak areas can be determined using various programs, including ChemStation for LC, which was used here.
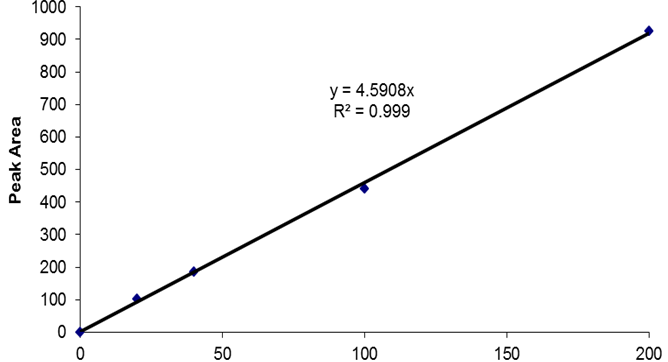
Figure 2. Example of a c-di-GMP HPLC standard curve. The standard curve was generated by plotting the peak areas obtained following the separation of 20 μl aliquots of c-di-GMP standards (here: 0, 1, 2, 5, and 10 pmo/μl) versus the total c-di-GMP amounts in pmol. Peak areas were obtained using the ChemStation for LC software.
- Using commercially available c-di-GMP, prepare the following standards in nanopure water: 1, 2, 5, 10 and 20 pmo/μl.
- Analysis of samples
- Resuspend the dried extracts from step A-11 in 200 μl nanopure water and vortex for 1 min.
- Centrifuge the solution at max speed (≥ 16,000 x g) to remove insoluble material.
- Using a 2 μm HPLC syringe filter attached to a 1 ml syringe, filter the sample supernatants into a new microfuge tube.
Note: Small sample volume loss may occur, but will not interfere with downstream application, as only a limited sample volume (20 μl) is subjected to HPLC analysis. - Analyze 20 μl per sample using the HPLC program used for the standards above. Repeat using biological replicates.
Note: This procedure has been optimized for P. aeruginosa PAO1 and PA14. Analysis of c-di-GMP levels in other strains or species may require the adjustment of sample volumes. - Determine the peak area for each sample and determine the c-di-GMP amounts using the standard curve established with the commercially available c-di-GMP in step B-1-c.
- Resuspend the dried extracts from step A-11 in 200 μl nanopure water and vortex for 1 min.
- Calculations
To normalize the c-di-GMP levels, total cellular protein levels from the extraction procedure must be determined.- Resuspend the cell pellet from step A-10 in 500 μl TE buffer.
- Sonicate the suspension for a total of 1 min using 10 sec bursts at 5 W followed by 15 sec off. Sonication should be carried out on ice.
- Determine the protein concentrations using a protein determination assay.
- Normalize the c-di-GMP levels to total cellular protein levels (i.e. pmol/mg) using the following calculations:
- Total c-di-GMP in pmol
= (pmol c-di-GMP) x 10
A factor of 10 is used here, as only 1/10th of the c-di-GMP extract (20 μl out of 200 μl) was used for HPLC analysis - Total protein in mg
= (mg/ml protein) * 0.5 ml
→Normalized c-di-GMP (pmol/mg)
= Total c-di-GMP/Total protein
- Total c-di-GMP in pmol
- Resuspend the cell pellet from step A-10 in 500 μl TE buffer.
- Generation of a standard curve
Recipes
- PBS
137 mM NaCl
2.7 mM KCl
4.3 mM Na2HPO4·7H2O
1.4 mM KH2PO4 (pH 7.2) - HPLC Solvent A
10 mM ammonium acetate in water
Do not adjust pH - HPLC Solvent B
10 mM ammonium acetate in methanol
Dissolve ammonium acetate salt in methanol
Do not adjust pH - TE buffer
10 mM Tris-HCl (pH 8.0)
1 mM EDTA
Acknowledgments
The c-di-GMP extraction procedure using heat and ethanol is based on previously published protocols (Amikam et al., 1995; Simm et al., 2004). The HPLC-based method for the detection and quantitation of c-di-GMP is a modification of protocols published by Thormann and Spormann (Thormann et al., 2006) and Ueda and Wood (2009). This work was supported by a grant from NIH (1RO1 A107525701A2).
References
- Amikam, D., Steinberger, O., Shkolnik, T. and Ben-Ishai, Z. (1995). The novel cyclic dinucleotide 3'-5' cyclic diguanylic acid binds to p21ras and enhances DNA synthesis but not cell replication in the Molt 4 cell line. Biochem J 311 ( Pt 3): 921-927
- Morgan, R., Kohn, S., Hwang, S. H., Hassett, D. J. and Sauer, K. (2006). BdlA, a chemotaxis regulator essential for biofilm dispersion in Pseudomonas aeruginosa. J Bacteriol 188(21): 7335-7343
- Petrova, O. E., Schurr, J. R., Schurr, M. J. and Sauer, K. (2012). Microcolony formation by the opportunistic pathogen Pseudomonas aeruginosa requires pyruvate and pyruvate fermentation. Mol Microbiol 86(4): 819-835.
- Simm, R., Morr, M., Kader, A., Nimtz, M. and Romling, U. (2004). GGDEF and EAL domains inversely regulate cyclic di-GMP levels and transition from sessility to motility. Mol Microbiol 53(4): 1123-1134.
- Thormann, K. M., Duttler, S., Saville, R. M., Hyodo, M., Shukla, S., Hayakawa, Y. and Spormann, A. M. (2006). Control of formation and cellular detachment from Shewanella oneidensis MR-1 biofilms by cyclic di-GMP. J Bacteriol 188(7): 2681-2691.
- Ueda, A. and Wood, T. K. (2009). Connecting quorum sensing, c-di-GMP, pel polysaccharide, and biofilm formation in Pseudomonas aeruginosa through tyrosine phosphatase TpbA (PA3885). PLoS Pathog 5(6): e1000483.
Article Information
Copyright
© 2013 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Roy, A. B., Petrova, O. E. and Sauer, K. (2013). Extraction and Quantification of Cyclic Di-GMP from Pseudomonas aeruginosa. Bio-protocol 3(14): e828. DOI: 10.21769/BioProtoc.828.
Category
Microbiology > Microbial biochemistry > Other compound
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link












