- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Production, quantification, and infection of Amazonian Phlebovirus (Bunyaviridae)
(*contributed equally to this work) Published: Vol 11, Iss 13, Jul 5, 2021 DOI: 10.21769/BioProtoc.4072 Views: 4049
Reviewed by: Woojong LeeAlexandros AlexandratosAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Isolation of Viral Biofilms From HTLV-1 Chronically Infected T Cells and Integrity Analysis
Coline Arone [...] Delphine Muriaux
Dec 20, 2024 1828 Views

Generation, Propagation, and Titering of Dicistrovirus From an Infectious Clone
Junzhou Shen [...] Eric Jan
Feb 20, 2025 2187 Views
Identification and Sorting of Adipose Inflammatory and Metabolically Activated Macrophages in Diet-Induced Obesity
Dan Wu [...] Weidong Wang
Oct 20, 2025 2232 Views
Abstract
Phlebotomine vectors, sand flies of the order Diptera, are known to transmit Leishmania parasites as well as RNA viruses (arboviruses) to humans. The arbovirus, Icoaraci Phlebovirus (BeAN 24262 - ICOV), used in this study was isolated from Nectomys rodents, a mammalian species that is the same natural sylvatic reservoir of Leishmania (Leishmania) amazonensis. This Leishmania species is distributed in primary and secondary forests in Brazil and other countries in America and causes localized and diffuse anergic skin lesions. In our recent studies, we observed an aggravation of the protozoan infection by ICOV through the modulation of cytokine expression, such as IL-10 and IFN-β, enhancing the parasite load and possibly the pathogenesis. Efficient viral production and quantitation had to be developed and standardized to ensure that immuno-molecular assays provide consistent and reproducible viral infection results. The standardization of these procedures becomes a particularly useful tool in research, with several applications in understanding the interaction between the host cell and Phlebovirus, as well as co-infections, allowing the study of intracellular signaling pathways. Here, we detail a protocol that allows the production and quantitation of the Icoaraci Phlebovirus using BHK-21 cells (baby hamster kidney cells) and subsequent infection of peritoneal macrophages from C57BL/6 mice.
Keywords: PhlebovirusBackground
In Brazil, approximately 210 different types of arboviruses have been isolated, of which 196 were found in the Brazilian Amazon (Rosa et al., 1998; Azevedo et al., 2007), one of the largest reserves of arboviruses in the world. The Bunyaviridae family is composed of more than 350 viruses divided into five genera, among which Orthobunyavirus, Phlebovirus, Nairovirus, and Hantavirus contain species pathogenic to humans and animals, while the genus Tospovirus is pathogenic only to plants (Walter and Barr, 2011; Ly and Ikegami, 2016).
In most human cases, Phlebovirus cause a wide spectrum of symptoms from mild febrile illness to hemorrhagic fever and death (Bird et al., 2009; Ikegami and Makino, 2011; Brisbarre et al., 2015). A specific member of Phlebovirus, the Rift Valley Fever virus, is potentially dangerous to pregnant domesticated animals such as cattle, goat, and sheep, with infection resulting in fetal malformation and abortion (Swanepoel and Coetzer, 2004). In this work, we used the Icoaraci Phlebovirus (BeAN 24262), which is prevalent in small forest animals (Causey and Shope, 1965). All Phlebovirus are morphologically similar: 80-120 nm in diameter, icosahedral symmetry, and enveloped with two surface glycoproteins (Gn and Gc) (Freiberg et al., 2008). The single-stranded RNA (ssRNA) is segmented and packaged within ribonucleoprotein particles (RNPs) by protein N and associated with an RNA-dependent RNA polymerase (RdRp). The small (S) segment uses the ambisense coding strategy, coding for a nucleocapsid protein (N) and non-structural protein (NSs) (Simons et al., 1990; Giorgi et al., 1991).
Studies in animal models suggest a protective effect of type I interferon (IFN) in infection caused by Phlebovirus (Mendenhall et al., 2009). Most viruses have developed the ability to express proteins capable of preventing the immune response (Habjan et al., 2009). This antagonistic activity has been observed for several non-structural viral proteins (Van Knippenberg et al., 2010) by mechanisms that explore different targets in IFN signaling pathways. Although Phlebovirus exhibit different levels of pathogenicity, the NSs protein is considered an important virulence factor for all members of the genus, as well as other members of the Bunyaviridae family (Habjan et al., 2009; Ikegami et al., 2009).
Several studies have shown the importance of the relationship between viruses and Leishmania parasites in the establishment of reciprocal adaptive advantages that promote survival and persistent infection (Ives et al., 2011; Hartley et al., 2012; Vivarini et al., 2015; Eren et al., 2016; Rath et al., 2018). Our group previously demonstrated that the induction of antiviral pathways by L. amazonensis increases macrophage infection (Pereira et al., 2010; Vivarini et al., 2015). Different Leishmania species associated with host immunological factors determine distinct clinical manifestations of leishmaniasis (Alvar et al., 2012). The production of leishmanicidal mediators, such as IL-12, IL-6, and reactive oxygen species (ROS), by innate immune cells contributes to infection control (Dayakar et al., 2019), but Leishmania parasites are able to subvert the signaling pathways that trigger these mechanisms and instead induce pathways favoring the infectious process. Using Icoaraci Phlebovirus as a model, we demonstrated that this virus potentiates infection by L. amazonensis in macrophages. Increased macrophage infection requires toll-like receptor 3 (TLR3) and protein kinase R (PKR) pathways due to the production of type I IFNs induced by the Leishmania and amplified during co-infection with the virus (Rath et al., 2019). Our results highlight the importance of complex and multifactorial analysis of infectious processes, especially in endemic areas where overlap occurs, as observed between L. amazonensis and Icoaraci Phlebovirus.
Given the socioeconomic impact of Phlebovirus infection and the potential co-infection in cutaneous leishmaniasis, the present protocol for the production, quantitation, and infection of Icoaraci Phlebovirus aims to address the molecular mechanisms associated with the interaction between sand fly-transmitted arboviruses and Leishmania parasite infection in vitro and in vivo in the cell host. Subsequent research using this methodology may reveal important aspects related to the host immune response, helping to outline the pathophysiological phenomena and clinical conditions and clarifying scenarios that may indicate how an antiviral response to double-stranded RNA (dsRNA) can mediate immune-inflammatory and immunopathogenic processes in Phlebovirus infection and co-infections.
Materials and Reagents
Cell culture vessels, plates, and tubes
25-cm2 cell culture flask (Corning, catalog number: CLS430639)
100-mm plates (SARSTEDT, catalog number: 82.1472.001)
6-well cell culture plates (Corning, catalog number: 83.3920)
24-well cell culture plates (Corning, catalog number: 83.3922)
15-ml and 50-ml Falcon tubes (BD Falcon)
Cell counting slides with grids
1.5-ml Eppendorf tubes
Eco Nitrile Gloves (SuperMax)
Filter tips, low-retention: 1,000 µl, 200 µl, 20 µl, 10 µl (Axygen, catalog numbers: 31000LRS, 3200LRS, 320LRS, 310LRS, respectively)
5-ml serological pipettes (SARSTEDT, catalog number: 86.1253.001)
10-ml serological pipettes (SARSTEDT, catalog number: 86.1254.001)
0.22-µm cell culture medium filtration unit (Corning, catalog number: CLS430513-12EA)
Cells
BHK-21 cells (baby hamster kidney cells, ATCC: CCL-10)
Cells are maintained in 25-cm2 cell culture flasks with complete DMEM (DMEM – high glucose and supplemented with 10% heat-inactivated FBS: see Recipe 1) at 37°C in a humidified atmosphere containing 5% CO2. After reaching confluence, cells are subcultured to maintain cell viability. For this, remove the medium and rinse the cells with 3 ml warm PBS without calcium and magnesium. Dissociate the cells using 1 ml trypsin-EDTA solution for 1 min. Add 4 ml complete DMEM, transfer to a new 15-ml conical tube, and centrifuge the cells for 5 min at 300 × g. Resuspend the cell pellet in fresh complete medium and split 1:20 in new flasks.
Phlebovirus
The Icoaraci Phlebovirus (BeAN 24262-ICOV) was obtained from Dr. Pedro Vasconcelos (Instituto Evandro Chagas/SVS/MS).
The virus was isolated from the liver, spleen, kidney, and heart of Nectomys squamipes in Belém, Pará, Brazil, and subsequently inoculated intracerebrally in 2-day-old Swiss mice. The samples were lyophilized and stored at -80°C. Before use, reconstitute the samples in 0.5 ml Puck’s medium.
Cell culture reagents
0.25% trypsin-EDTA solution (Sigma-Aldrich, catalog number: T4049), stored at -20°C
Dulbecco’s Modified Eagle’s Medium (DMEM) with 4.5 g/L glucose and L-glutamine (Lonza, catalog number: 12-604Q)
100× penicillin-streptomycin solution (HyClone-GE, catalog number: SV30010)
Puck’s medium (PAN Biotech, catalog number: P04-51100)
Phosphate-buffered saline (PBS) (Lonza, catalog number: 17-516Q)
Fetal bovine serum (FBS) (Thermo Fisher Scientific, GibcoTM, catalog number: 10270106)
UltraPureTM DNase/RNase-free distilled water (Thermo Fisher Scientific, InvitrogenTM, catalog number: 10977-035)
Methylcellulose (Sigma-Aldrich, catalog number: M7027)
100-bp DNA ladder, 500 µg/ml (Promega, catalog number: G201A)
6× gel loading dye (Promega, catalog number: G190A)
Standard agarose for routine gel electrophoresis (UniScience, catalog number: AGR-LE-100)
Ethidium bromide (Sigma-Aldrich, catalog number: E7637)
Tris-base (Promega, catalog number: H5131)
Glacial acetic acid (Sigma-Aldrich, catalog number: A6283)
UltraPureTM 0.5 M EDTA (Ethylenediaminetetraacetic acid, disodium salt dihydrate), pH 8.0 (Thermo Fisher Scientific, catalog number: 15575020)
Crystal violet (Sigma-Aldrich, catalog number: C0775)
Sodium thioglycolate (Sigma-Aldrich, catalog number: T0632), stored at room temperature
Direct-zolTM RNA MiniPrep Plus kit (Zymo Research, catalog number: R2070)
ImProm-IITM Reverse Transcription System (Promega, catalog number: A3800)
Random primers (Promega, catalog number: C1181)
GoTaq® DNA polymerase (Promega, catalog number: M3001)
Complete DMEM (see Recipes)
Semi-solid medium (see Recipes)
Crystal violet solution (see Recipes)
10× TAE Buffer (1 L) (see Recipes)
1× TAE Buffer (1 L) (see Recipes)
1.5% agarose gel (see Recipes)
100-bp DNA ladder (50 µg/ml) (see Recipes)
3% sodium thioglycolate (see Recipes)
Equipment
P1000 (Gilson, catalog number: F123602)
P200 (Gilson, catalog number: F123601)
P20 (Gilson, catalog number: F123600)
P2 (Gilson, catalog number: F144801)
Lab refrigerator set at 4°C (Electrolux, model: IF55)
Lab freezer set at -20°C (Prosdocimo, model: F21)
Lab freezer set at -80°C (Panasonic, model: MDF-U56VC-PA)
Incubator at 37°C with 5% CO2 (Sanyo inCU-Safe, model: MCO-17AC)
Laminar flow hood (Esco Class II BSC- AirStream)
Centrifuge 5418 (Eppendorf) at room temperature; set here at 24°C
Centrifuge (ThermoFisher Scientific, model: D-37520)
NanoDropTM ND-1000 Spectrophotometer (ThermoFisher Scientific)
Balance (BEL Analytical Equipment, model: S622)
Veriti 96-Well thermal cycler (Applied Biosystems PCR instruments, model: 9902)
Mini-gel migration system (Loccus Biotecnology, model: LCH-7X8)
Electrophoresis power supply (GIBCO BRL, Life Technologies, 250) and power cables
U.V. transilluminator (Fisher Scientific, model: 88A)
Procedure
Figure 1 illustrates the steps for virus production, quantitation, and infection. A detailed description of the procedures is found in the following items.
Figure 1. Different steps for the production, quantitation, and infection of Icoaraci Phlebovirus. Figure created in the Mind the Graph platform.
Production of the Icoaraci Phlebovirus
Plate 4 × 106 BHK-21 cells in 100-mm plates (7 ml/plate). In order to obtain a good virus concentration, seed approximately 30 plates. For this step, use BHK cells that have undergone no more than five passages.
After exactly 24 h, wash the cells once with 2 ml warm PBS and proceed with viral absorption (next step).
Dilute the virus in 1 ml DMEM without FBS [multiplicity of infection (MOI) 0.01] and absorb the inoculum for 2 h at room temperature. The inoculum volume is not sufficient to cover the whole plate surface, so please move the plate carefully every 20 min to allow the inoculum to move across the plate.
Remove the inoculum, add 7 ml DMEM (supplemented with 10% FBS) (Recipe 1), and incubate at 37°C for 3 days.
Collect the supernatant and centrifuge at 18,000 × g for 1 h at 4°C to concentrate the virus.
Discard the supernatant and resuspend the pellet in 300 µl PBS. Split the volume between 3 cryovials and store at -80°C until the titration.
Quantitation of the Icoaraci Phlebovirus
Viral titers are determined using a plaque assay. For this, plate 1.5 × 106 BHK-21 cells/well in a 6-well plate. Cells should be used up to a maximum of passage 3.
After 24 h, infect the cells (~80% confluence) with 200 µl 10-fold serial dilution of ICOV stock, as described in the previous section, and incubate for 2 h at room temperature. Again, the inoculum volume is not sufficient to cover the well surface, so please move the plate carefully every 20 min to allow the inoculum to move across the plate.
Using a pipette, remove the inoculum completely and cover the monolayer with 2 ml semi-solid medium composed of complete DMEM plus 1% methylcellulose (Recipe 2). In this step, it is very important to ensure complete removal of the inoculum in order for the semi-solid medium to be evenly distributed throughout the cell monolayer.
Three days post-infection, fix and stain the cells with 2 ml 0.1% crystal violet solution containing 10% formaldehyde (Recipe 3) under slight agitation for 12 h at room temperature.
Wash the wells delicately with running water to remove the fixative/dye and allow the plate to dry upside down at room temperature for 24 h. Viral plaques can be counted on the white-light transilluminator, and titers are expressed as plaque-forming units (PFU)/ml (Figure 2).
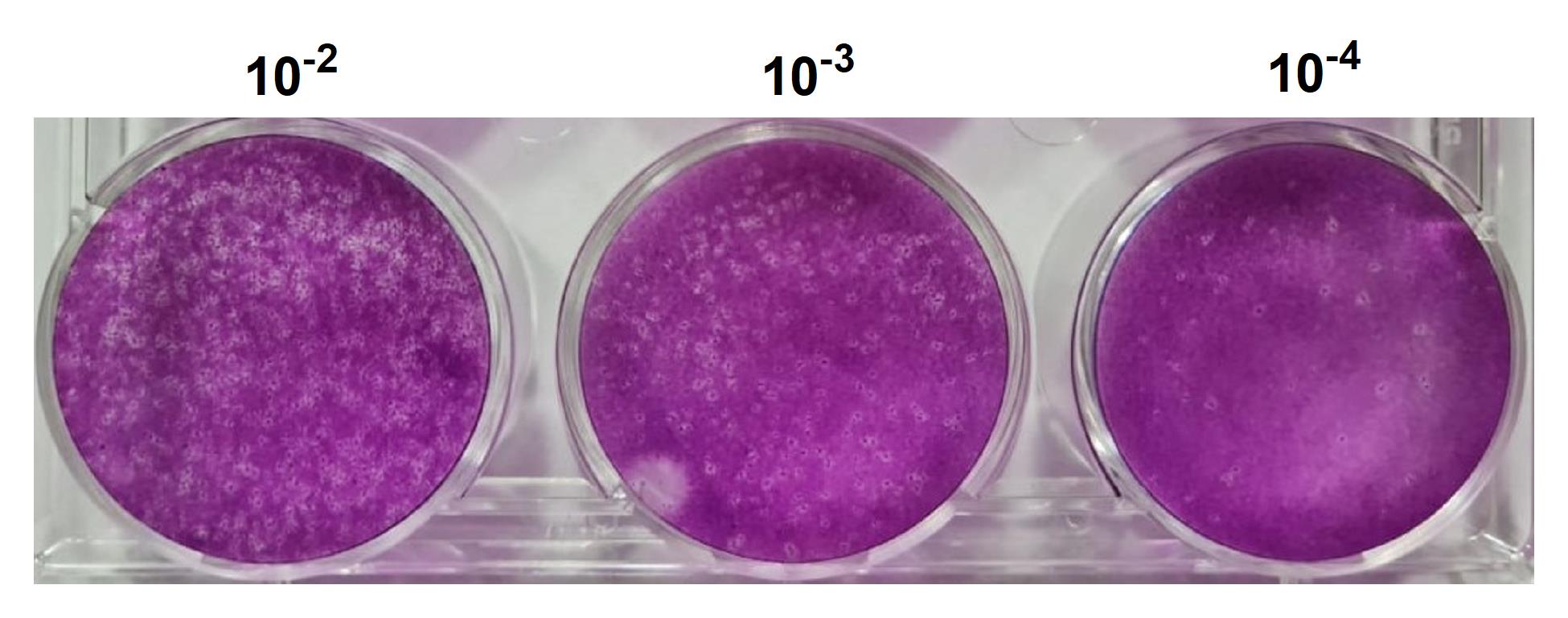
Figure 2. Quantitation of Icoaraci Phlebovirus. The supernatant from 48 h after infection was collected, diluted to three different concentrations (10-2, 10-3, and 10-4), and titrated in a BHK-21 monolayer as described above. After 3 days, the cells were fixed and stained with crystal violet.
Obtaining and cultivating peritoneal macrophages
Mouse colonies are maintained at the Department of Immunology, UFRJ in accordance with the rules established by the National Council for the Control of Animal Experimentation. The animal experimentation protocols used in this project were approved by the Animal Use Ethics Committee (Protocol No. 046/20).
Peritoneal macrophages from 8-week-old C57 Black-6 (C57BL/6) mice are elicited with 2 ml 3% sodium thioglycolate (intraperitoneal injection, Recipe 8) for 4 days, and cells are obtained after euthanization by injecting 8 ml DMEM into the peritoneal cavity. The mice were inclined at a 45° angle with the head down, positioning the intestines cranially, away from the administration area. The needle was then inserted in the lower right quadrant of the abdomen at a 30-45° angle for either the inoculation of thioglycolate or to obtain macrophages.
Centrifuge the cells at 400 × g for 10 min at 4°C.
Resuspend the cells in 1 ml DMEM and count using a hemocytometer under the microscope (10× dilution in the Neubauer chamber with DMEM).
Plate the macrophages on glass coverslips in 24-well plates (2 × 105 cells per well) or 6-well plates (4 × 106 cells per well).
After a 1-h adherence at 37°C, wash the cells with PBS and add complete DMEM. The macrophages are kept for 1 day at 37°C in a 5% CO2 atmosphere and subsequently subjected to infection.
Infection
For peritoneal macrophage ICOV infections, adsorption is performed for 1 h at 37°C using an MOI of 1.
Dilute the virus in DMEM without FBS and use 100 µl for 24-well plates and 1 ml for 6-well plates.
Remove the inoculum with a pipette and add 1 ml (24-well plate) or 2 ml (6-well plate) complete DMEM.
The subsequent treatment time, conditions, or other additional infections for which the experimental model is proposed may vary. For example, in Leishmania infections, we allow the parasite to interact for 4 h after viral adsorption and then lyse the cells for analysis of gene expression by qPCR.
Analysis of murine macrophage infection by semiquantitative PCR for viral detection
To detect and confirm the viral infection and replication in peritoneal macrophages, a semiquantitative PCR must be performed.
After 4, 24, or 48 h of infection, isolate and purify the total RNA using the Direct-zolTM RNA MiniPrep Plus kit according to the manufacturer's instructions.
Quantitate the RNA using the NanoDrop. The expected concentration should be 100-200 ng/µl, and the 260/280 ratio should be 1.8-2.0, ensuring good RNA quality.
First-strand cDNA synthesis is performed in a reaction containing Improm-II Reverse Transcriptase, a mix of dNTPs, and random primers, as described in the manufacturer's instructions.
RT-PCR is performed using 10 µM primers for the N protein gene of ICOV (sense 5’-AGGTGAGGCTGTAAATCTTG-3’ and antisense 5’-TCACATCATCCTTCCAAGTG-3’), 2.5 U GoTaq DNA polymerase, 1.5 mM MgCl2, 10 mM dNTPs, 2 µl cDNA, and 1× appropriate buffer in a final volume of 50 µl. The PCR cycle is as follows: 95°C for 1 min, 48°C for 1 min, and 72°C for 1 min in 40 amplification cycles.
PCR products are separated on a 1.2% agarose gel containing 0.5 µg/ml ethidium bromide in 1× TAE buffer (Recipe 5) at 80-100 V and photographed in a UV transilluminator. The product has 100 bp (Figure 3).
Figure 3. Peritoneal macrophages from wild-type C57BL/6 mice were infected with stationary-phase promastigotes of L. (L.) amazonensis at a ratio of 5 parasites/cell, Icoaraci Phlebovirus (ICOV), or co-infected with both for 4 h, 24 h, or 48 h. Total RNA was extracted and analyzed by semiquantitative PCR for the Icoaraci N protein.
Recipes
Complete DMEM
To make 500 ml complete DMEM:
445 ml 1× DMEM
50 ml 10% FBS (heat-Inactivated)
5 ml 1× penicillin-streptomycin (100×)
Semi-solid medium
Complete DMEM plus 1% methylcellulose
Add 1 g methylcellulose to 100 ml distilled water in an autoclavable flask.
Before autoclaving, add a stir bar in order to mix the solution and maintain sterility. For complete solubilization, store the solution overnight at 4°C and then leave the flask in a magnetic stirrer.
Note: If a lumpy solution forms, cool the temperature of the solution and mix again. The procedure should be repeated until the solution is homogeneous.
Add cold methylcellulose to the cold complete DMEM (v/v) and thoroughly mix to ensure a homogeneous mixture. Prior to use, wait until the bubbles dissipate and the temperature raises to room temperature.
Crystal violet solution
0.1% crystal violet in PBS solution
10% formaldehyde
10× TAE Buffer (1 L)
Weigh 48.5 g Tris-base, 11.4 ml glacial acetic acid and 20 ml 0.5 M EDTA pH 8.0
Add distilled water up to 1 L
Store at room temperature
1× TAE Buffer (1 L)
Add 100 ml 10× TAE (Recipe 5) to 900 ml distilled water
Store at room temperature
1.5% agarose gel
Weigh agarose powder (1.5 g) in an autoclaved glass bottle
Add 100 ml 1× TAE
Heat in a microwave until the agarose is homogeneously dissolved
Cool down at room temperature for a few minutes
100-bp DNA Ladder (50 µg/ml)
For 240 µl DNA Ladder in a 1.5-ml tube on ice, add 176 µl UltraPureTM DNase/RNase-free distilled water
Add 40 µl 6× gel loading dye and 24 μl 100-bp DNA ladder at 500 µg/ml
Store at 4°C
3% sodium thioglycolate
Weigh sodium thioglycolate powder (3.0 g) in an autoclaved glass bottle
Add water up to 100 ml
Filter the solution with a 0.22-µm cell culture medium filtration unit
Store at room temperature
Acknowledgments
We thank Professor Pedro Fernando da Costa Vasconcellos for providing the Phleboviruses, which are fundamental for carrying out this work. We thank Professor Clarissa Damaso for the support in the virology methodologies. We also thank all members of the Molecular Parasitology laboratory for helpful discussion. This research was supported by the National Council for Scientific and Technological Development (CNPq) and Fundação Carlos Chagas Filho de Amparo à Pesquisa do Estado do Rio de Janeiro (FAPERJ).
Competing interests
The authors declare no competing interests.
Ethics
All animal experiments were performed in accordance with protocols approved by the National Council for Animal Experiment Control and Animal Use Ethics Committee (Protocol No. 046/20 – ID: 01200.001568/2013-87 Validity: 08/30/2022).
References
- Alvar, J., Vélez, I. D., Bern, C., Herrero, M., Desjeux, P., Cano, J., Jannin, J., den Boer, M. WHO Leishmaniasis Control Team. (2012). Leishmaniasis worldwide and global estimates of its incidence. PLoS One 7(5):e35671.
- Azevedo, R. S., Nunes, M. R., Chiang, J. O., Bensabath, G., Vasconcelos, H. B., Pinto, A. Y., Martins, L. C., Monteiro, H. A., Rodrigues, S. G. and Vasconcelos, P. F. (2007). Reemergence of Oropouche fever, northern Brazil. Emerg Infect Dis 13(6): 912–915.
- Bird, B. H., Ksiazek, T. G., Nichol, S. T. and Maclachlan, N. J. (2009). J Am Vet Med Assoc. 234: 883-893.
- Brisbarre, N., Plumet, S., Cotteaux-Lautard, C., Emonet, S.F., Pagès, F. and Leparc-Goffart, I. (2015). A rapid and specific real time RT-PCR assay for diagnosis of Toscana virus infection. J Clin Virol 66:107-111.
- Causey, O.R. and Shope, R. E. (1965). Icoaraci, a new virus related to naples phlebotomus fever virus. Proc Soc Exp Biol Med 118: 420-1.
- Dayakar, A., Chandrasekaran, S., Kuchipudi, S. V. and Kalangi, S. K. (2019). Cytokines: Key Determinants of Resistance or Disease Progression in Visceral Leishmaniasis: Opportunities for Novel Diagnostics and Immunotherapy. Front immunol 10: 670.
- Eren, R.O., Reverte, M., Rossi, M., Hartley, M. A., Castiglioni, P., Prevel, F., Martin, R., Desponds, C., Lye, L. F., Drexler, S. K., Reith, W., Beverley, S. M., Ronet, C. and Fasel, N. (2016). Mammalian innate immune response to a Leishmania-Resident RNA virus increases Macrophage survival to promote parasite persistence. Cell Host Microbe 20(3): 318-328.
- Freiberg, A. N, Sherman, M. B., Morais, M. C., Holbrook, M. R. and Watowich, S. J. (2008). Three-dimensional organization of Rift Valley fever virus revealed by cryoelectron tomography. J Virol 82(21):10341-10348.
- Giorgi, C., Accardi, L., Nicoletti, L., Gro, M. C., Takehara, K., Hilditch, C., Morikawa, S. and Bishop, D. H. (1991). Sequences and coding strategies of the S RNAs of Toscana and Rift Valley fever viruses compared to those of Punta Toro, Sicilian Sandfly fever, and Uukuniemi viruses. Virol 180(2):738-753.
- Habjan, M., Pichlmair, A., Elliott, R. M., Overby, A. K., Glatter, T., Gstaiger, M., Superti-Furga, G., Unger, H. and Weber, F. (2009). NSs protein of rift valley fever virus induces the specific degradation of the double-stranded RNA-dependent protein kinase. J Virol 83(9):4365-4375.
- Hartley, M. A., Ronet, C., Zangger, H., Beverley, S. M. and Fasel, N. (2012). Leishmania RNA virus: when the host pays the toll. Front Cell Infect Microbiol 2: 99.
- Ikegami, T., Narayanan, K., Won, S., Kamitani, W., Peters, C. J. and Makino, S. (2009). Rift Valley fever virus NSs protein promotes post-transcriptional downregulation of protein kinase PKR and inhibits eIF2α phosphorylation. PLoS Pathog 5(2): e1000287.
- Ikegami T and Makino S. (2011). The Pathogenesis of Rift Valley Fever. Viruses 3: 493-519.
- Ives, A., Ronet, C., Prevel, F., Ruzzante, G., Fuertes-Marraco, S., Schutz, F., Zangger, H., Revaz-Breton, M., Lye, L. F., Hickerson, S. M., Beverley, S. M., Acha-Orbea, H., Launois, P., Fasel, N. and Masina, S. (2011). Leishmania RNA virus controls the severity of mucocutaneous leishmaniasis. Science 331(6018): 775-778.
- Ly, J. H., Ikegami, T. (2016). Rift Valley fever virus NSs protein functions and the similarity to other bunyavirus NSs proteins. Virol J 13:118.
- Mendenhall, M., Wong, M.-H., Skirpstunas, R., Morrey, J. D. and Gowen, B. B. (2009). Punta Toro Virus (Bunyaviridae, Phlebovirus) Infection in Mice: Strain Differences in Pathogenesis and Host Interferon Response. Virology 395(1): 143-151.
- Mind the Graph platform. www.mindthegraph.com
- Pereira, R. M., Teixeira, K. L., Barreto-de-Souza, V., Calegari-Silva, T. C., De-Melo, L. D., Soares, D. C., Bou-Habib, D. C., Silva, A. M., Saraiva, E. M. and Lopes, U. G. (2010). Novel role for the double-stranded RNA-activated protein kinase PKR: modulation of macrophage infection by the protozoan parasite Leishmania. FASEB J 24(2): 617-626.
- Rath, C. T., Schnellrath, L. C., Damaso, C. R., de Arruda, L. B., Vasconcelos, P., Gomes, C., Laurenti, M. D., Calegari Silva, T. C., Vivarini, Á. C., Fasel, N., Pereira, R. and Lopes, U. G. (2019). Amazonian Phlebovirus (Bunyaviridae) potentiates the infection of Leishmania (Leishmania) amazonensis: Role of the PKR/IFN1/IL-10 axis. PLoS Negl Trop Dis 13(6): e0007500.
- Rosa, J. F. S. T., Rosa, A. P. A T., Vasconcelos, P. F. C., Pinheiro, F. P., Rodrigues, S. G., Rosa, E. S. T., Dias, L. B. and Cruz, A. C. R. (1998). Arboviruses isolated in the Evandro Chagas Institute, including some described for the first time in the Brazilian Amazon region, their known hosts, and their pathology for man. In: Rosa, A. P. A. T., Vasconcelos, P. F. C. and Rosa, J. F. S. T. (Eds.). An overview of arbovirology in Brazil and neighbouring countries. Instituto Evandro Chagas, Belém, 18-31.
- Simons, J. F., Hellman, U. and Pettersson, R. F. (1990). Uukuniemi virus S RNA segment: ambisense coding strategy, packaging of complementary strands into virions, and homology to members of the genus Phlebovirus. J Virol 64(1): 247-255.
- Swanepoel, R. and Coetzer, J. A. W. (2004). Rift Valley fever. In: Coetzer, J. A. W. and Tustin, R. C. (Eds.). Infectious diseases of livestock with special reference to southern Africa. 2. Cape Town: Oxford University Press, 1037-1070.
- Van Knippenberg, I., Carlton-Smith, C. and Elliott, R. M. (2010). The N-terminus of Bunyamwera orthobunyavirus NSs protein is essential for interferon antagonism. J Gen Virol 91: 2002–2006.
- Vivarini, Á., Pereira, R., Barreto-de-Souza, V., Temerozo, J. R., Soares, D. C., Saraiva, E. M., Saliba, A. M., Bou-Habib, D. C. and Lopes, U. G. (2015). HIV-1 Tat protein enhances the intracellular growth of Leishmania amazonensis via the ds-RNA induced protein PKR. Sci Rep5: 16777.
- Walter, C. T. and Barr, J. N. (2011). Recent advances in the molecular and cellular biology of bunyaviruses. J Gen Virol 92: 2467-2484.
Article Information
Copyright
© 2021 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Rath, C. T., Vivarini, A. C., Pereira, R. M. and Lopes, U. G. (2021). Production, quantification, and infection of Amazonian Phlebovirus (Bunyaviridae). Bio-protocol 11(13): e4072. DOI: 10.21769/BioProtoc.4072.
Category
Microbiology > Microbial cell biology > Virus propagation
Cell Biology > Cell isolation and culture > Virus isolation
Immunology > Immune cell isolation > Macrophage
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link