- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Assessing Swarming of Aerobic Bacteria from Human Fecal Matter
(*contributed equally to this work) Published: Vol 11, Iss 9, May 5, 2021 DOI: 10.21769/BioProtoc.4008 Views: 3899
Reviewed by: Juan Facundo Rodriguez AyalaAna S. AlmeidaAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
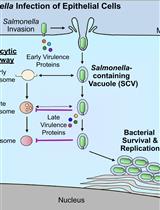
Bacterial Pathogen-mediated Suppression of Host Trafficking to Lysosomes: Fluorescence Microscopy-based DQ-Red BSA Analysis
Mădălina Mocăniță [...] Vanessa M. D'Costa
Mar 5, 2024 2895 Views
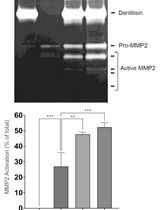
Purification of Native Dentilisin Complex from Treponema denticola by Preparative Continuous Polyacrylamide Gel Electrophoresis and Functional Analysis by Gelatin Zymography
Pachiyappan Kamarajan [...] Yvonne L. Kapila
Apr 5, 2024 2088 Views
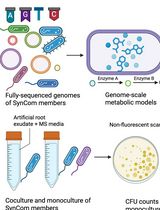
In Silico Prediction and In Vitro Validation of Bacterial Interactions in the Plant Rhizosphere Using a Synthetic Bacterial Community
Arijit Mukherjee [...] Sanjay Swarup
Nov 5, 2025 1655 Views
Abstract
Swarming – swift movement across a surface via flagella propulsion – is a unique property of many bacteria. The role of swarming, particularly among bacterial populations of the human gut microbiome, is not yet fully understood; although, it is becoming an area of increased scientific and clinical inquiry. To further characterize bacterial swarming in human health, an effective assay for swarming that utilizes complex material, such as fecal matter, is necessary. Until now, the vast majority of swarming assays have only been able to accommodate bacteria grown in culture, most often Pseudomonas. These assays tend to use a standard lysogenic broth (LB) agar medium; however, the reagents involved have not been tailored to the inoculation of complex material. In this paper, we offer a specialized protocol for eliciting the swarming of bacteria from frozen human fecal samples. We describe the simple, yet reproducible steps required to perform the assay, identifying an ideal volume of 7.5 μl for inoculation of material, as well as an ideal agar concentration of 0.4%. This protocol typically allows researchers to identify swarming within 24 h after incubation in a standard incubator.
Keywords: Bacterial swarmingBackground
Swarming is a unique bacterial property that ensures mass movement across a surface in a collective fashion (Verstraeten et al., 2008; Kearns, 2010). The exact host consequences of bacteria that swarm are not yet fully known; however, recent research has aimed to elucidate such a role in pathogenesis (Barak et al., 2009). It has been theorized that swarming enables bacteria to adhere to and disperse from localized spots of infection (Kearns, 2010), as well as to escape engulfment by macrophages (Amendola, 1998). For instance, the swarming of Proteus mirabilis has been implicated in catheter-associated urinary tract infections (Jones et al., 2004). Moreover, bacteria of many species become resistant to a range of antibiotics when they swarm, in a manner that appears unrelated to normal antibiotic efflux mechanisms (Butler et al., 2010).
More recently, our group has shown for the first time that swarming bacterial activity from human fecal matter is a specific indicator of intestinal inflammation and/or polyp formation (De et al., 2021). This represents a novel advancement since patient feces may now be used to inform clinicians in the assessment of “vague abdominal” symptoms among patients at risk for inflammatory bowel disease, ulcerative colitis, and polyps, among other conditions. This knowledge could help to guide clinical decision-making, such as determining which patients need a colonoscopy and further evaluation versus those who do not, ultimately reducing overall healthcare costs. As a result, the importance of optimizing an assay that detects swarming bacteria from feces is critical to the validation of our findings. Thus far, assays that focus on detecting swarming bacteria have been restricted to the study of cultured single strains of bacteria, namely Pseudomonas (Tremblay et al., 2008; Morris et al., 2011; Ha et al., 2014; Morales-soto et al., 2015). Ha et al., (2014), Morris et al., (2011), and Tremblay et al., (2008) describe thorough protocols for assaying bacterial swarming in Pseudomonas aeruginosa (Tremblay and Deziel, 2008; Morris et al., 2011; Ha et al., 2014); however, little, if any, attention has been focused on designing a protocol for eliciting bacterial swarming from human fecal samples, which are often the most convenient materials from which researchers can learn more about the microbial communities of the human gut. In this protocol, which we used in our group’s recent publication in De et al. (2021), we describe an assay to observe swarming in the shortest amount of time and with the least consumption of laboratory reagents. To the best of our knowledge, this is the first comprehensive description and optimization of an assay protocol that detects swarming bacteria from fecal matter.
Materials and Reagents
Petri dish, stackable lid, 100 mm × 15 mm, sterile, polystyrene (Fisherbrand, Fisher Scientific, catalog number: FB0875713)
Fecal specimen collection cup, 650 ml (Fisherbrand, Commode Specimen Collection System, catalog number: 23-038032)
Microcentrifuge tube, 1.5 ml (Fisherbrand, Fisher Scientific, catalog number: 05-408-130)
Adjustable 0.5-10 μl pipette (Eppendorf Research, catalog number: EPPR4402)
10 μl pipette Tips, racked, sterile (Thomas Scientific, catalog number: 1148U48)
Agar, bacteriological grade (Research Products International, CAS number: 9002-18-0)
Sodium chloride (Fischer Chemicals, Fisher Scientific, CAS number: 7647-14-5)
Tryptone, microbiologically tested (Sigma-Aldrich, CAS number: 91079-40-2)
Yeast extract (Acumedia, Neogen Corporation, catalog number: 7184B)
Distilled water (Thermo Fisher Scientific, catalog number: 15230001)
Agar LB plates (see Recipes)
Note: All materials and reagents are stored at room temperature.
Equipment
Epson Expression 1600 Scanner (Epson, Expression 1600, catalog number: E1600-PRO)
Incubator (Thermo Scientific, NAPCO Series 8000DH Incubator, catalog number: 3541)
-80°C freezer (Thermo Fisher Scientific, Revco ExF, catalog number: EXF40086D)
Autoclave (Amsco Scientific, Eagle, catalog number: SV120)
Hotplate (Thermolyne, Cimarec, catalog number: HP131225)
Biological hood (Thermo Fisher Scientific, 1300 Series Class II, Biological Safety Cabinet, catalog number: 1353)
Software
ImageJ Version 1.59e (National Institutes of Health, https://imagej.nih.gov/ij/download.html)
Procedure
Prepare the agar LB plates with a final agar concentration of 0.4% (see Recipes).
Remove fecal samples from the freezer
Fresh human fecal samples were obtained in 2015 (under protocols IRB# 2009-446 and 2015-4465) from healthy donors without intestinal distress and donors known to suffer from intestinal inflammation. Specimens were collected in sterile fecal specimen cups without preservatives. Specimens were then kept at 4°C for up to 15 min before transport to the laboratory at ambient temperature for processing, which consisted of aliquoting the specimen into a 100-μl microcentrifuge and storage at -80°C since 2015.
Remove the fecal samples from the freezer and allow to thaw at room temperature on the bench.
Under a biological hood, use a pipette to inoculate 7.5 μl healthy sample on one side of a plate (as a control) and 7.5 μl disease sample on the other half of the plate.
Note: If the samples are not pre-homogenized, homogenization is preferable using a homogenizer for uniformity.
Take care to avoid puncturing the agar with the pipette tip, as this may impede the ability to identify future swarming.
Leave the cover off the Petri dish and allow to dry under a biological hood for 10 min.
Incubation
(Optional) Once dry, take an image of the plate on the scanner, which will serve as a “before image” for later reference. See Figure 1A below.
Place the plate upright (not inverted) in an incubator, set to 37°C and a humidity of 80%.
Remove the plate from the incubator after 24 h and assess visually for the presence of swarming.
(Optional) Scan an image of the plate and label as “after image” for record-keeping. See Figure 1B below.
(Optional) Input the before and after images into the NIH’s ImageJ program to quantitatively measure the folds of expansion. Use the “Free Select” tool to highlight an area in the ‘before image’, then highlight the area of swarming in the ‘after image’. This method allows measurement of the pixels of growth and enables quantitation even when swarming occurs in irregular directions.

Figure 1. Bacterial swarming on LB agar plates before and after incubation. A. 7.5 μl fecal samples were inoculated onto 20 ml 0.4% LB agar plates, which were then scanned. B. Plates were placed in an incubator, set to 37°C and a humidity of 80%, and were removed and scanned after 24 h.
Data analysis
An agar concentration of 0.4% was found to elicit the most swarming (compared with agar concentrations of 0.5%, 0.6%, and 0.7%), as was a sample volume of 7.5 μl (compared with volumes of 2.5 μl or 5.0 μl). At the same time, the thawing method for the samples (on ice or at room temperature) was found to have no significant impact on the presence of swarming. These conclusions were drawn from the statistical analysis of 10 independent experiments (performed in duplicate) for 6 separate patient samples. One-way ANOVA was performed in GraphPad Prism (v8) to determine significant differences between agar and inoculant volume conditions.
Recipes
Agar LB plates
1 g Tryptone
0.5 g Yeast extract
0.5 g NaCl
0.4 g Agar
100 ml dH2O
Combine the water and dry ingredients while stirring on a hotplate, until the water begins to boil, then autoclave to sterilize (121°C, 15 PSI, 65 min)
After autoclaving, allow the 0.4% LB agar solution to cool until warm to the touch, stirring continuously to prevent clumping or hardening
Pour 20 ml 0.4% LB agar solution into sterile Petri dishes
Allow LB agar plates to cool and harden on the bench at RT for at least one h
Acknowledgments
Research was conducted under a Student Research Fellowship Award from the Crohn’s and Colitis Foundation of America (CCFA) #640806 awarded to A.B, as well as #431602 awarded to S.M. The data acquisition was also partially supported by the Broad Medical Research Program (BMRP) grant #362520 at CCFA to S.M. The authors would like to thank Arpan De and Hao Li for their advice, suggestions for variables to explore, and help with formatting and editing. This protocol was modified from work completed by our group, published in March of 2021 in Gastroenterology (De et al., 2021) https://doi.org/10.1053/j.gastro.2021.03.017.
Competing interests
The authors have no competing interests or financial disclosures to report.
Ethics
All described protocols have been approved by the Albert Einstein College of Medicine Institutional Review Board (IRB# 2009-446 and 2015-4465). Informed consent was given by the human subjects from whom fecal samples were acquired.
References
- Amendola, A., Geisenberger, O., Anderson, J. B., Givskov, M., Schleifer, K. H. and Eberl, L. (1998) Serratia liquefaciens swarm cells exhibit enhanced resistance to predation by Tetrahymena sp. FEMS Microbiol Lett 164(1): 69-75.
- Barak, J. D., Gorski, L., Liang, A. S. and Narm, K. E. (2009). Previously uncharacterized Salmonella enterica genes required for swarming play a role in seedling colonization. Microbiology 155(Pt 11): 3701-3709.
- Butler, M. T., Wang, Q. and Harshey, R. M. (2010) Cell density and mobility protect swarming bacteria against antibiotics. Proc Natl Acad Sci 107(8):3776-3781.
- De, A., Chen, W., Li, H., Wright, J. R., Lamendella, R., Lukin, D. J., Szymczak, W. A., Sun, K., Kelly, L., Ghosh, S., Kearns, D. B., He, Z., Jobin, C., Luo, X., Byju, A., Chatterjee, S., San Yeoh, B., Vijay-Kumar, M., Tang, J. X., Prajapati, M. et al. (2021). Bacterial Swarmers Enriched during Intestinal Stress Ameliorate Damage. Gastroenterology S0016-5085(21)00524-2.
- Ha, D. G., Kuchma, S. L. and O'Toole, G. A. (2014). Plate-based assay for swarming motility in Pseudomonas aeruginosa. Methods Mol Biol 1149: 67-72.
- Jones, B. V., Young, R., Mahenthiralingam, E. and Stickler, D. J. (2004) Ultrastructure of Proteus mirabilis swarmer cell rafts and role of swarming in catheter-associated urinary tract infection. Infect Immun 72(7): 3941-3950.
- Kearns, D. B. (2010). A field guide to bacterial swarming motility. Nat Rev Microbiol 8(9): 634-644.
- Morales-Soto, N., Anyan, M. E., Mattingly, A. E., Madukoma, C. S., Harvey, C. W., Alber, M., Deziel, E., Kearns, D. B. and Shrout, J. D. (2015). Preparation, imaging, and quantification of bacterial surface motility assays. J Vis Exp 98.
- Morris, J. D., Hewitt, J. L., Wolfe, L. G., Kamatkar, N. G., Chapman, S. M., Diener, J. M., Courtney, A. J., Leevy, W. M. and Shrout, J. D. (2011). Imaging and analysis of Pseudomonas aeruginosa swarming and rhamnolipid production. Appl Environ Microbiol 77(23): 8310-8317.
- Tremblay, J. and Deziel, E. (2008). Improving the reproducibility of Pseudomonas aeruginosa swarming motility assays. J Basic Microbiol 48(6): 509-515.
- Verstraeten, N., Braeken, K., Debkumari, B., Fauvart, M., Fransaer, J., Vermant, J. and Michiels, J. (2008). Living on a surface: swarming and biofilm formation. Trends Microbiol 16(10): 496-506.
Article Information
Copyright
© 2021 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Byju, A. S., Patel, D., Chen, W. and Mani, S. (2021). Assessing Swarming of Aerobic Bacteria from Human Fecal Matter. Bio-protocol 11(9): e4008. DOI: 10.21769/BioProtoc.4008.
Category
Microbiology > Microbe-host interactions > Bacterium
Cell Biology > Cell movement > Cell motility
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









