- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Rhodobacter capsulatus Gene Transfer Agent (RcGTA) Activity Bioassays
Published: Vol 3, Iss 2, Jan 20, 2013 DOI: 10.21769/BioProtoc.317 Views: 10789

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
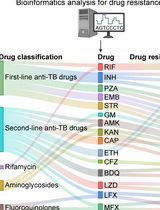
Identification of Mycobacterium tuberculosis and its Drug Resistance by Targeted Nanopore Sequencing Technology
Chen Tang [...] Guangxin Xiang
Feb 5, 2025 1985 Views
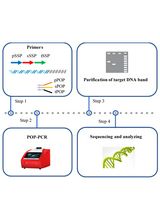
Protocol to Identify Unknown Flanking DNA Using Partially Overlapping Primer-based PCR for Genome Walking
Mengya Jia [...] Haixing Li
Feb 5, 2025 1369 Views
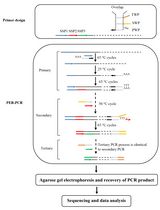
Protocol to Mine Unknown Flanking DNA Using PER-PCR for Genome Walking
Zhou Yu [...] Haixing Li
Feb 20, 2025 1498 Views
Abstract
RcGTA, a small phage-like particle produced by Rhodobacter capsulatus, was initially identified in culture filtrates as a DNase-resistant form of DNA transfer between R. capsulatus cells. This gene transfer assay has been used to identify RcGTA-producing strains and to help determine the roles of genes thought to be responsible for RcGTA production.
Keywords: TransductionMaterials and Reagents
- GTA donor strain (see Notes 1)
- Recipient strain (see Notes 1)
- RCV broth (Beatty and Gest, 1981)
- YPS broth (Wall et al., 1975)
- YPS agar (2 plates per recipient-donor combination, one plate per donor, and one plate per recipient)
- 0.22 μm filtered GTA buffer (Solioz and Marrs, 1977)
- 0.45 μm low protein binding (PVDF) syringe filters (e.g. EMD Millipore, catalog number: SLHV033RB )
- Plate spreader (Including 95% ethanol + flame)
- 1 ml syringes
- Test tubes for aerobic culturing (e.g. Fisher brand brand, catalog number: 14-961-30 ) w/caps
- Culture tubes for anaerobic/photosynthetic culturing (e.g. Fisher brand, catalog number: 14-959-37A w/screw caps)
- Polypropylene sterile culture tubes (e.g. Simport, catalog number: T405-2A )
- Optional (if transferring photosynthesis marker): Chambers/packs for anaerobic plate growth
- RCV Medium (see Recipes)
- YPS Medium (see Recipes)
- GTA buffer (see Recipes)
Equipment
- Shaking incubator
- Microcentrifuge
- Incandescent light-box or light incubator
- 1 ml pipettes
- 1.5 ml Microcentrifuge tubes
Procedure
- Preparation:
- 3 days prior to the assay, inoculate GTA donor strains in RCV broth and grow aerobically overnight at 35 °C (200-250 rpm).
- 2 days prior, measure optical densities (OD) of overnight cultures of GTA donor strains and normalize them by dilution with RCV broth. It is simplest to dilute all ODs to match the lowest, as the actual OD is irrelevant so long as all cultures are at the same final density. Use a 1% v/v inoculum of normalized donor to inoculate YPS broth without antibiotics to grow anaerobically and photosynthetically over two days at 35 °C without shaking, mixing occasionally by inverting the culture tubes. These culture tubes should be filled to the brim with YPS broth and sealed tightly, to create anaerobic conditions, and placed equidistant from the light source. Incandescent bulbs are better than fluorescent bulbs, but the heat generated from the bulbs must be dissipated (e.g. by having the culture tubes in a water tank) in order to maintain culturing temperature.
- 1 day prior, inoculate recipient strain in RCV broth and grow aerobically overnight at 35 °C (200-250 RPM).
- Assay:
- Pass donor strain cultures through a 0.45 μm filter, collecting the filtrates in polypropylene tubes.
- Centrifuge 1 ml of recipient cultures, decant the supernatant and re-suspend in an equal volume of GTA buffer.
- Mix the following in a polypropylene tube:
-Filtrate Controls (for each Donor filtrate): 0.5 ml GTA buffer, 0.1 ml filtrate.
-Recipient Controls (for each Recipient Strain): 0.5 ml GTA buffer, 0.1 ml recipient cells.
-Experimental samples: 0.4 ml GTA buffer, 0.1 ml filtrate, 0.1 ml recipient cells. - Incubate tubes at 35 °C for 1 h with no shaking.
- Add 0.9 ml RCV broth to each tube.
- Incubate tubes at 35 °C for 3 h with shaking at ~200 rpm.
- Transfer the 1.5 ml mixtures from the polypropylene tubes to individual 1.5 ml microcentrifuge tubes.
- Plate 150 μl of each filtrate + recipient mix on plates to represent 10% of the total. This is not necessary for the recipient and filtrate controls.
- Centrifuge all tubes, decant supernatant, and resuspend pellets in the small (~100 μl) volume that remains. Spread these resuspensions on plates to represent 90% and 100% for experimental and control assays, respectively. These plates should be selective for the transfer of the marker, or grown in conditions that select for a transferred marker.
- Grow for 2-3 days at 30-35 °C.
- Count colonies. Determine ratios of colonies on experimental plates over the number of colonies found on a positive control (e.g. a wild type strain).
Notes
- One common bioassay employs monitoring the transfer of the puhA gene to puhA deletion mutant DW5 (Wong et al., 1996), with selection for transfer of the puhA gene being the ability to grow photosynthetically. This bioassay shows no spontaneous mutation background. Another bioassay is the transfer of rifampicin resistance, which is a property of some strains of R. capsulatus, to rifampicin-sensitive strains such as the natural isolate strain B10 (Weaver et al., 1975). This bioassay has a low but detectable rate of spontaneous mutation background that must be accounted for in the data analysis.
- It is essential to compare transfer of the same marker, as marker sizes can affect bioassay results (Hynes et al., 2012), presumably affecting both packaging and recombination rates.
- Bioassay absolute numbers can vary greatly depending on growth state of donor and recipient cells, batch of media (e.g. the batch of Yeast Extract was used for YPS), so it is essential to compare ratios within one bioassay, and perform multiple independent replicates. When performing bioassays for or into strains/mutants with impaired growth or viability, it can help to normalize transfer rate to the number of viable cells (either donors or recipients, whichever is impaired) by performing viable cell counts alongside the bioassay.
Recipes
- RCV Medium
7.5 mM (NH4)2SO4
30 mM DL-malate, pH 6.8 with NaOH
54 μM EDTA
0.8 mM MgSO4
0.51 mM CaCl2
43 μM FeSO4
3 μM Thiamine-HCl
9.5 μM MnSO4
45 μM H3BO3
0.2 μM Cu(NO3)2
0.83 μM ZnSO4
3 μM NaMoO4
4.5 mM KH2PO4
5.1 mM K2HPO4
pH to 6.8 - YPS Medium
3.0 g/L Yeast Extract
3.0 g/L Peptone
2 mM MgSO4
2 mM CaCl2
pH to 7.0 with NaOH or HCl, as needed
Add 1.5% Agar for solid media - GTA Buffer
10mM Tris-HCl (pH 7.8) with NaOH
1.0 mM MgCl2
1.0 mM CaCl2
1.0 mM NaCl
500 μg/ml BSA (Fraction V)
Filter sterilize with 0.22 μm filter
Acknowledgments
APH was supported by fellowships from Memorial University School of Graduate Studies and the Natural Sciences and Engineering Research Council (NSERC) of Canada. This research in ASL’s lab was supported by grants from NSERC, the Canada Foundation for Innovation, and the Newfoundland and Labrador Research & Development Corporation. This protocol is based on work originally reported in: Hynes et al. (2012).
References
- Beatty, J. T. and H. Gest (1981). Generation of succinyl-coenzyme A in photosynthetic bacteria. Arch Microbiol 129(5): 335-340.
- Hynes, A. P., R. G. Mercer, D. E. Watton, C. B. Buckley and A. S. Lang (2012). DNA packaging bias and differential expression of gene transfer agent genes within a population during production and release of the Rhodobacter capsulatus gene transfer agent, RcGTA. Mol Microbiol 85(2): 314-325.
- Solioz, M. and B. Marrs (1977). The gene transfer agent of Rhodopseudomonas capsulata. Purification and characterization of its nucleic acid. Arch Biochem Biophys 181(1): 300-307.
- Wall, J. D., P. F. Weaver and H. Gest (1975). Genetic transfer of nitrogenase-hydrogenase activity in Rhodopseudomonas capsulata. Nature 258(5536): 630-631.
- Weaver, P. F., J. D. Wall and H. Gest (1975). Characterization of Rhodopseudomonas capsulata. Arch Microbiol 105(3): 207-216.
- Wong, D. K., W. J. Collins, A. Harmer, T. G. Lilburn and J. T. Beatty (1996). Directed mutagenesis of the Rhodobacter capsulatus puhA gene and orf 214: pleiotropic effects on photosynthetic reaction center and light-harvesting 1 complexes. J Bacteriol 178(8): 2334-2342.
Article Information
Copyright
© 2013 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Hynes, A. P. and Lang, A. S. (2013). Rhodobacter capsulatus Gene Transfer Agent (RcGTA) Activity Bioassays. Bio-protocol 3(2): e317. DOI: 10.21769/BioProtoc.317.
Category
Molecular Biology > DNA > Transformation
Microbiology > Microbial genetics > DNA
Microbiology > Microbial genetics > Transformation
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link








