- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
In situ Hybridization (ISH) in Preparasitic and Parasitic Stages of the Plant-parasitic Nematode Meloidogyne spp.
Published: Vol 8, Iss 6, Mar 20, 2018 DOI: 10.21769/BioProtoc.2766 Views: 9848
Reviewed by: Eugenio LlorensAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Quantitative Estimation of Auxin, Siderophore, and Hydrogen Cyanide Production in Halo and Drought-Tolerant Bacterial Isolates for Cucumber Growth
Zeinab Fotoohiyan and Ali Salehi Sardoei
Oct 5, 2025 1342 Views
Reproducible Emu-Based Workflow for High-Fidelity Soil and Plant Microbiome Profiling on HPC Clusters
Henrique M. Dias [...] Christopher Graham
Jan 20, 2026 403 Views
Abstract
The spatio-temporal expression pattern of a gene provides important indications to better understand its biological function. In situ hybridization (ISH) uses a labeled complementary single-stranded RNA or DNA probe to localize gene transcripts in a whole organism, a whole organ or a section of tissue. We adapted the ISH technique to the plant parasite Meloidogyne spp. (root-knot nematode) to visualize RNAs both in free-living preparasitic juveniles and in parasitic stages settled in the plant tissues. We describe each step of the probe synthesis, digoxigenin (DIG) labeling, nematode extraction from plant tissue, and ISH procedure.
Keywords: Gene expression patternBackground
So far, the stable transformation of plant-parasitic nematode(s) has not been successful. ISH enables the analysis of spatio-temporal gene expression in vivo in whole-mount Meloidogyne spp. nematodes. These root-knot nematodes hatch in the soil as microscopic vermiform juveniles (J2) and infect host plant roots. J2s penetrate the root and migrate to the root vascular cylinder cells. The juveniles settle in the root and develop into J3 and J4 parasitic juveniles that induce the differentiation specialized feeding cells. The nematode eventually develops into a pear-shaped female that will release hundreds of eggs on the root surface. Here, we report a detailed protocol to detect single RNA molecules in preparasitic whole mount J2s and parasitic stages. ISH on parasitic stages requires an additional procedure the day before extraction of the nematodes from infected roots. We describe the detection of transcripts using digoxigenin (DIG)-labeled cDNA probes in nematode whole mount tissues.
Materials and Reagents
- Nitrile gloves
- RNase-free microcentrifuge tubes (1.5 ml) (e.g., Thermo Fisher Scientific, Applied BiosystemsTM, catalog number: AM12450 )
- Sieve of 2.5 mm/250 µm/40 µm/10 µm
- 50 ml tubes (Corning, Falcon®, catalog number: 352070 )
- Paper towel
- Microscope slides and cover slip (e.g., Fisher Scientific, catalog number: 12-544-1 )
- Autoclaved razor blades
- RNase-free filtered tips, e.g.,
20 µl tips (Mettler-Toledo, Rainin, catalog number: 17007957 )
200 µl tips (Mettler-Toledo, Rainin, catalog number: 17002927 )
1,000 µl tips (Mettler-Toledo, Rainin, catalog number: 17014361 ) - Root-knot nematodes (e.g., M. incognita Morelos strain, M. enterolobii Godet strain) preparasitic J2s (10,000) or parasitic stages (50)
- DIG-labelled ISH probe(s)
Note: See Procedure A for probe synthesis and storage. - Forward and reverse primers designed to allow an amplicon size around 200 bp (e.g., SePOP Desalted oligos from Eurogentec)
- PCR-grade dNTPs (Thermo Fisher Scientific, InvitrogenTM, catalog number: 10297117 )
- Pfu DNA Polymerase (Promega, catalog number: M7741 )
- QIAquick PCR Purification Kit (QIAGEN, catalog number: 28104 )
- QIAquick Gel extraction kit (QIAGEN, catalog number: 28704 )
- Digoxigenin (DIG) DNA Labeling Kit (Roche Diagnostics, catalog number: 11175025910 )
- TE buffer pH 8 (10 mM Tris-HCl, 1 mM EDTA) (e.g., Sigma-Aldrich, catalog number: 93283 )
- RNaseZap® RNase Decontamination Solution (Thermo Fisher Scientific, InvitrogenTM, catalog number: AM9780 )
- 70% ethanol
- Tap water
- Acetone (e.g., VWR, catalog number: 20065.293 )
- Methanol (e.g., VWR, catalog number: 20847.307 )
- Anti-Digoxigenin-AP-Fab fragments Labelling Mix (Roche Diagnostics, catalog number: 11093274910 )
- BCIP 5-bromo-4-chloro-3-indolyl-phosphate, 4-toluidine salt (Roche Diagnostics, catalog number: 11383221001 )
- NBT 4-Nitro blue tetrazolium chloride (Roche Diagnostics, catalog number: 11383213001 )
- Pectinex® (Sigma-Aldrich, catalog number: P2611 )
- Celluclast® (Sigma-Aldrich, catalog number: C2730 )
- Sodium phosphate dibasic (Na2HPO4) (Sigma-Aldrich, catalog number: S7907 )
- Potassium phosphate, monobasic (KH2PO4) (Sigma-Aldrich, catalog number: P9791 )
- Sodium chloride (NaCl) (Sigma-Aldrich, catalog number: S7653 )
- Magnesium sulfate (MgSO4·7H2O) (Sigma-Aldrich, catalog number: 63138 )
- Sucrose (e.g., Sigma-Aldrich, catalog number: S9378 )
- 10x PBS (Thermo Fisher Scientific, InvitrogenTM, catalog number: AM9624 )
- 37% formaldehyde solution (Sigma-Aldrich, catalog number: F15587 )
Note: This product has been discontinued. - Proteinase K 20 mg/ml (Thermo Fisher Scientific, InvitrogenTM, catalog number: AM2546 )
- Formamide, deionized (Sigma-Aldrich, catalog number: F9037 )
- SSC buffer 20x (Thermo Fisher Scientific, InvitrogenTM, catalog number: AM9770 )
- Boehringer blocking reagent (Roche Diagnostics, catalog number: 11096176001 )
- SDS (Thermo Fisher Scientific, InvitrogenTM, catalog number: 15553027 )
- Denhart’s solution (Sigma-Aldrich, catalog number: D2532 )
- EDTA pH 8 (Thermo Fisher Scientific, InvitrogenTM, catalog number: AM9261 )
- Salmon sperm DNA (Thermo Fisher Scientific, InvitrogenTM, catalog number: 15632011 )
- tRNA from baker’s yeast (Sigma-Aldrich, catalog number: R8759 , type X-SA)
- Maleic acid (Fisher Scientific, catalog number: 10348843)
Manufacturer: Acros Organics, catalog number: 125230051 . - Sodium chloride (NaCl) (Sigma-Aldrich, catalog number: S7653 )
- Sodium hydroxide (NaOH) (VWR, catalog number: 28245.298 )
- Tris-base (e.g., Sigma-Aldrich, catalog number: T1503 )
- Concentrated HCl (VWR, catalog number: 20248.295 , type 35%)
- Deionized water
- Nuclease-free water (not DEPC-Treated; Thermo Fisher Scientific, InvitrogenTM, catalog number: AM9938 )
- For nematode extraction (see Recipes)
- Pectinex® and Celluclast® mix
- M9-buffer pH 7
- 50% sucrose in M9 buffer
- RNase-free 1x PBS
- Pectinex® and Celluclast® mix
- For probe, fixation and hybridization, nuclease-free (see Recipes)
- Fixative buffer
- 1 mg/ml Proteinase K solution
- tRNA from Baker’s yeast
- Hybridization buffer (HB)
- Washing buffer 1 (4x SSC + 0.1% SDS)
- Washing buffer 2 (0.1x SSC + 0.1% SDS)
- 10% Boehringer blocking reagent
- Filtered maleic acid buffer, pH 7.5
- Tris-HCl pH 9.5
- Alkaline phosphatase detection buffer pH 9.5 (APB)
- Fixative buffer
Equipment
- Pipetmens (e.g., Gilson, models: P2 , P20 , P200 , P1000 )
- Thermocycler (e.g., Biometra, model: T3000 )
- NanoDrop (e.g., Agilent Technologies, model: Agilent 2100 Bioanalyser , catalog number: G2939)
- Microcentrifuge at room temperature (RT) (e.g., Hitachi Koki, model: CT15E ) with fixed-angle rotor (e.g., Hitachi Koki, model: T15A61 )
- Centrifuge for 50 ml tube (e.g., Eppendorf, model: 5804 R ) with swing-bucket rotor (e.g., Eppendorf, model: A-4-44 )
- 50 ml glass beaker (Corning, PYREX®)
- Fume hood
- Aquarium air pump (Rena Aquatic Supply, model: Model 301 )
- Glass plate
- Mini hybridization oven (Appligene, catalog number: 001050414 )
- Vortexer (e.g., Baxter, catalog number: S8223-1 )
- Orbital shaker (VWR)
- 37 °C water bath
- Fridge at 4 °C
- -20 °C freezer
- -80 °C freezer
- Autoclave
- Microscope (ZEISS, model: Axioplan 2 )
Procedure
- PCR digoxigenin (DIG)-labeled cDNA antisense and control (sense) probe synthesis and purification
- PCR amplify a selected specific fragment from the targeted gene (~200 bp) using cDNA or, in absence of introns, genomic DNA as a matrix, with forward and reverse primers 1 µM each, dNTPs 250 µM and proof-reading Pfu polymerase.
- Purify the PCR1 fragment with QIAquick PCR Purification Kit if a single band is obtained or purify the band from the gel with QIAquick gel extraction kit, elute with 10 µl elution buffer and quantify on gel or using NanoDrop.
- Perform the DIG labeling step through a second PCR by using DIG labeled dUTPs using 50-100 ng of PCR1, 2 µM forward or reverse primer for sense or anti-sense probe, respectively, DIG-dUTP/dNTP mix (DIG DNA labeling kit).
- Purify PCR product with QIAquick PCR Purification Kit and resuspend in 10 µl TE buffer at pH 8.0.
- For quality control, run small amount (1 µl) of the labeled probe (PCR2) on an agarose gel. In addition, probes can be tested using dot blot to analyze the sensitivity or Northern blot for specificity (Liao et al., 2014), prior to in situ hybridization (ISH).
- DIG-labeled cDNA probes are stored at -20 °C. They are stable for more than a year.
- PCR amplify a selected specific fragment from the targeted gene (~200 bp) using cDNA or, in absence of introns, genomic DNA as a matrix, with forward and reverse primers 1 µM each, dNTPs 250 µM and proof-reading Pfu polymerase.
- Day 1. Preparasitic or parasitic root-knot nematode (RKN) extraction from roots and fixation for ISH
- To extract parasitic stages from infected roots, rinse roots with tap water, cut them into ~1 cm sticks.
- Immerse roots in the Pectinex® and Celluclast® mix and incubate the mixture for 4 h (to overnight) at RT under gentle agitation (orbital shaker, 50 rounds per minute).
- Prepare 2 Corning tubes containing 40 ml of 50% sucrose/M9 and store them in the fridge at 4 °C.
- After digestion, pour the liquid through a 2.5 mm sieve and collect the filtrate.
- Pour the liquid through a 250 µm sieve and collect the filtrate.
- Pour the liquid through a 40 µm sieve.
- Collect the parasitic RKNs on the 40 µm sieve with a minimum volume of tap water (~8 ml).
- Purify the nematodes by sucrose gradient centrifugation: add slowly 4 ml of the collected nematode on the top of cold 50% sucrose/M9 mix (2 tubes) (Figure 1).
- Centrifuge at 720 x g, at 4 °C for 5 min, without a break.
- Collect the layer of nematodes with a P1000 and wash them carefully adding 40 ml M9 buffer in a 50 ml glass beaker (Figure 1).
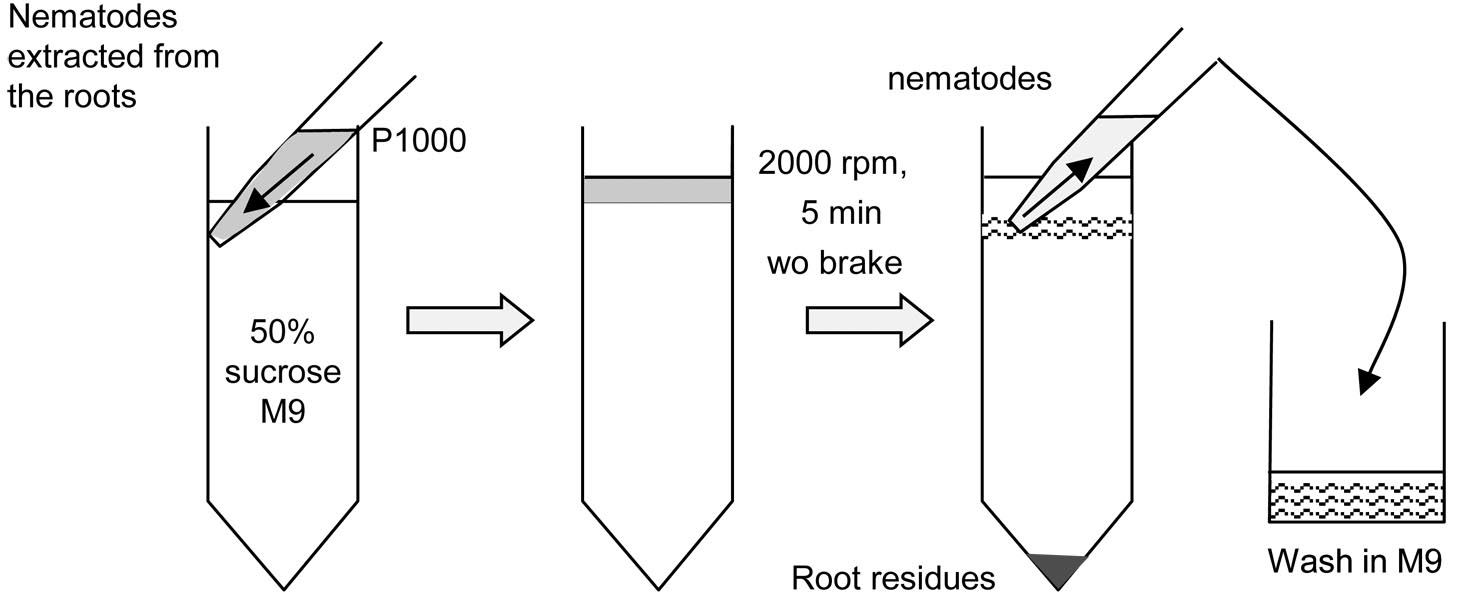
Figure 1. Extraction of parasitic stages by sucrose gradient - To remove sucrose, wash them twice with 50 ml M9, on the 10 µm sieve, in a 50 ml glass beaker and finally transfer to a 50-ml Corning tube.
- Collected nematodes are at the parasitic juvenile’s stage 2 (J2), 3 (J3) or 4 (J4), or adult females. Freshly hatched RKN preparasitic J2 are obtained from eggs as described in Caillaud and Favery (2016).
- Under a fume hood, with gloves, resuspend the nematodes (10,000 J2s or at least 50 parasitic stages) in 10 ml of fixative buffer in a 50-ml Corning tube. The fixation lasts overnight (16 h) for the preparasitic J2 and 48 h for parasitic stages at RT.
- To extract parasitic stages from infected roots, rinse roots with tap water, cut them into ~1 cm sticks.
- Day 2. (or Day 4 for parasitic stages) Nematode cutting and permeabilization for ISH
- Wipe down bench top and pipettors with RNaseZap®.
- Clean a glass plate with soap, rinse with nuclease-free water, spray RNaseZap® and rinse with 70% ethanol. Dry well with a paper towel.
- Centrifuge the 50 ml tube containing nematodes soaked in fixative buffer at 1,620 x g, slow deceleration.
- Withdraw fixative buffer but keep 500 µl and transfer to 1.5 ml Eppendorf tubes.
- Under the fume hood, wearing gloves, spread the nematodes on the glass plate, and cut the nematodes into two to three pieces using the cutting system (vibrating aquarium pump + autoclaved razor blades) described in Figure 2 and Video 1. Hold it with both hands. Check under the microscope the efficiency of the cutting. Do not cut too much, in particular the parasitic stages.
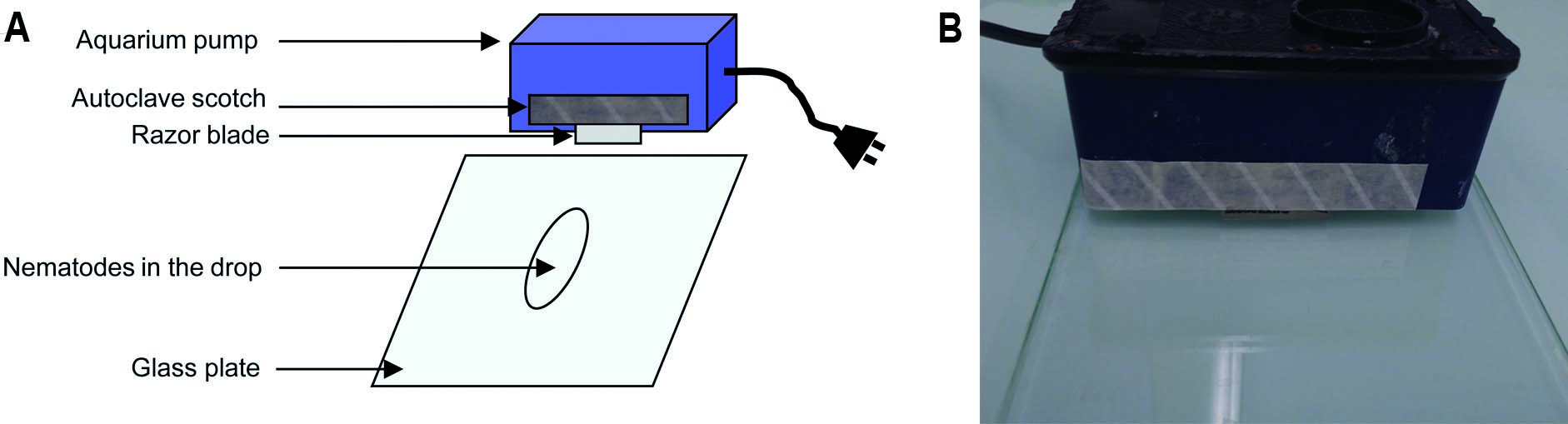
Figure 2. Nematode cutting before ISH. Diagram (A) and picture of the experimental device (B).Video 1. Nematode cutting for ISH - Add few drops of 1x RNase-free PBS buffer to collect the nematode sections on the glass plate and transfer into an Eppendorf tube.
- Spin down the sample (15,000 x g, 1 min, slow deceleration). Remove the buffer.
- Suspend in 1 ml of 1x RNase-free PBS, and then centrifuge the tube at 15,000 x g for 1 min with slow deceleration. Repeat Steps C7 and C8 three times.
- For nematode permeabilization, add 1 ml of cold 1 mg/ml Proteinase K solution to the nematodes. Keep Proteinase K in a cold block and do not vortex.
- Incubate the tube at 37 °C for 1 h under rotation (in a hybridization oven).
- Spin down the sample (15,000 x g, 1 min, slow deceleration). Keep the pellet.
- Wash the sample in 1 ml of 1x PBS and centrifuge the tube at 15,000 x g for 1 min with slow deceleration. Keep the pellet. Repeat this step three times. In the last wash, remove as much 1x PBS as possible to obtain a dry pellet.
- Place the pellet at -80 °C for 15 min.
- Add 1 ml cold methanol (precool at -80 °C).
- Vortex to suspend the nematodes and leave them 2 min at RT.
- Spin down the sample (15,000 x g, 1 min, slow deceleration).
- Remove methanol.
- Add 1 ml cold acetone (precool at -80 °C).
- Place the tube at -80 °C for 15 min.
- Spin down the sample (15,000 x g, 1 min, slow deceleration).
- Wipe down bench top and pipettors with RNaseZap®.
- Day 2 (or Day 4 for parasitic stages): (in situ) hybridization
- Set the temperature of the hybridization oven up to 50 °C
- Preheat the hybridization buffer (HB) at 50 °C.
- Remove acetone.
- Add 500 µl of pre-heat HB on the nematode pellet.
- Spin down the sample (15,000 x g, 1 min, slow deceleration) and remove the supernatant.
- Add 500 µl of pre-heated HB on the nematode pellet.
- Incubate for 30 min at 50 °C under rotation in the oven.
- Meanwhile, denature the probes at 100 °C for 5 min (water bath) and transfer onto ice immediately (let the temperature drop swiftly to prevent re-annealing). Keep them on ice for 3 min.
- Spin down the sample (15,000 x g, 1 min, slow deceleration).
- Set the temperature of the oven at 42 °C.
- Suspend the nematodes in 100 µl HB per probe (e.g., 500 µl for 5 probes).
- Dispatch 100 µl HB into individual nuclease free tubes.
- Add 5 µl of each probe.
- Perform the hybridization overnight at 42 °C (40-50 °C, see Notes) under agitation in the oven.
- Set the temperature of the hybridization oven up to 50 °C
- Day 3 (or Day 5 for parasitic stages): Washes and revelation
- Spin down the sample (15,000 x g, 1 min, slow deceleration). Discard supernatant.
- Wash the pellet with 1 ml washing buffer 1 (4x SSC, 0.1% SDS), 10 min at RT under rotation.
- Spin down the sample (15,000 x g, 1 min, slow deceleration). Discard supernatant.
- Repeat Steps E2 and E3 twice.
- Wash the pellet with 1 ml washing buffer 2 (0.1x SSC, 0.1% SDS), 10 min at 50 °C under rotation.
- Spin down the sample (15,000 x g, 1 min, slow deceleration). Discard supernatant.
- Repeat Steps E5 and E6 twice.
- Spin down the sample (15,000 x g, 1 min, slow deceleration). Discard supernatant.
- Wash 30 sec with 1 ml of filtered maleic acid buffer (1x).
- Spin down the sample (15,000 x g, 1 min, slow deceleration). Discard supernatant.
- Incubate the nematodes at 37 °C for 30 min (oven) in 1 ml of 1x Boehringer blocking reagent (see Recipes).
- Incubate the nematodes at 37 °C for 3 h in 1 ml of the anti-DIG antibody coupled with Alkaline Phosphatase (1:500) in 1x Boehringer blocking reagent.
- Spin down the sample (15,000 x g, 1 min, slow deceleration). Discard supernatant.
- Wash three times each with 1 ml 1x filtered maleic acid buffer at 37 °C during 15 min.
- Wash briefly with 1 ml Alkaline Phosphatase detection buffer (APB).
- Reveal the signal by incubating the nematodes for 6 to 8 h at RT in 1 ml APB + 3.75 µl BCIP + 5 µl NBT without agitation.
- Put a drop on a microscope slide, add a cover slip.
- Observe under a microscope (Axioplan2, 10x objective).
- If no signal is detected, continue the incubation overnight at 4 °C.
- Wash the nematode with nuclease-free water once.
- Put a drop on a microscope slide, add a cover slip.
- Observe under the microscope and image nematodes.
- Spin down the sample (15,000 x g, 1 min, slow deceleration). Discard supernatant.
Data analysis
Nematodes are examined with differential interference contrast microscopy. The signal observed should be compared to the images obtained with the control sense probe (negative control) (Figure 3). A positive control should be added. We usually used the MiPG gene encoding a polygalacturonase expressed in the two sub ventral glands (Jaubert et al., 2002a). We recommend taking pictures of at least 40 stained nematodes, analyzing the cellular localization of detected RNA spots, and to count the number of stained nematodes. Each ISH experiment should be reproduced independently. ISH could be performed after siRNA gene silencing of the target gene to ensure specificity and/or to evaluate the silencing effect. Moreover, control experiments using a different set of probes hybridizing the same target gene could be done to ensure the specificity of the probes to the target gene.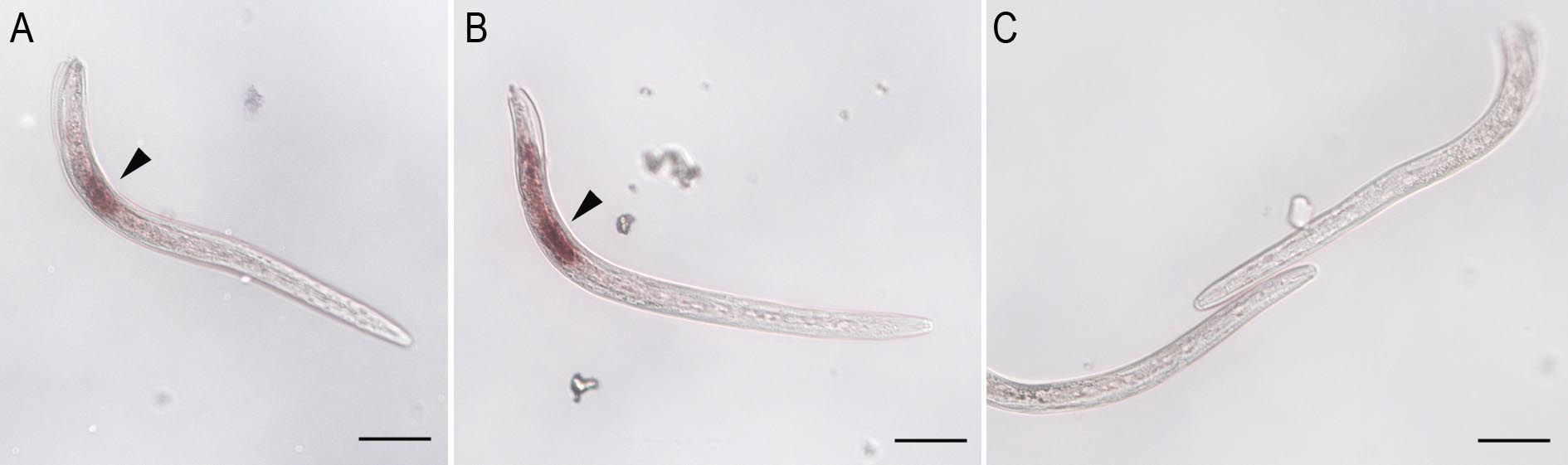
Figure 3. Tissue expression of M. incognita EFFECTOR1 (MiEFF1) in preparasitic J2 and parasitic juveniles. A-B. Transcripts from MiEFF1 (Minc17998) were localized in the salivary dorsal gland (arrowhead) of preparasitic J2s. C. The sense MiEFF1 control probe yielded no labeling of nematode tissues. MiEFF1 tissue expression in a parasitic juvenile was presented in Jaouannet et al., 2012. Scale bars = 50 µm.
Notes
- For nematode purification and fixation, prepare fresh solutions immediately before use.
- Prepare two Corning tubes containing 40 ml of 50% sucrose/M9 and store them in the fridge at 4 °C.
- For ISH, wear gloves. All solutions are RNAse Free, do not pipette directly into the Master solutions.
- All the experiments performed with the fixative and hybridization buffers should be performed under the fume hood.
- Be very careful when using the razor blade.
- Day 1 Place aliquots of methanol and acetone at -80 °C for nematode permeabilization step of the next day.
- For nematode permeabilization, do not vortex the Proteinase K and keep it in a cold block.
- The temperature of hybridization can be adjusted to obtain the best balance between the specificity and quantity of the signal.
- Digoxigenin (DIG)-labeled RNA probes can be used instead of cDNA probes as in Rosso et al. (1999).
- Use nuclease-free, not DEPC-treated, water.
- For revelation, horseradish peroxidase (HRP) can be used in replacement of alkaline phosphatase (AP). We chose AP because the signal from AP substrates gradually increases over time, is very stable and can last several days.
Recipes
For nematode extraction
- Pectinex® and Celluclast® mix
15 ml Celluclast®
30 ml Pectinex®
Tap H2O up to 200 ml - M9-buffer
42.3 mM Na2HPO4
22 mM KH2PO4
85.6 mM NaCl
1 mM MgSO4·7H2O - 50% sucrose in M9
50 g sucrose
M9 buffer up to 100 ml - RNase-free 1x PBS
1 ml 10x PBS
9 ml nuclease-free distilled water
- Fixative buffer
1 ml 37% formaldehyde solution
1x PBS up to 10 ml
Notes:- Prepare and manipulate under the fume hood.
- If 37% formaldehyde solution was stored at 4 °C; warm it to RT prior to opening.
- Prepare and manipulate under the fume hood.
- 1 mg/ml Proteinase K solution
1 ml Proteinase K solution (20 mg/ml)
1x PBS up to 20 ml - tRNA from Baker’s yeast
Dissolve 500 U yeast tRNA in 1 ml of nuclease-free water
Aliquot and store at -80 °C - Hybridization buffer (HB)
Note: Prepare and manipulate under the fume hood and store at -20 °C.
5 ml deionized formamide (99.5%, 4 °C)
0.5 ml SSC buffer (20x, RT)
1 ml Boehringer blocking reagent (10%, 4 °C)
2 ml SDS (10%, RT)
200 µl Denhart’s solution (50x, -20 °C)
20 µl EDTA pH 8 (0.5 M, RT)
200 µl salmon sperm DNA (10 mg/µl, -20 °C)
62 µl tRNA from Baker’s yeast (500 U/ml, -80 °C)
Nuclease-free water up to 10 ml - Washing buffer 1 (4x SSC + 0.1% SDS)
400 µl SSC buffer (20x, RT)
20 µl SDS (10%, RT)
Nuclease-free water up to 2 ml - Washing buffer 2 (0.1x SSC + 0.1% SDS)
10 µl SSC buffer (20x, RT)
20 µl SDS (10%, RT)
Nuclease-free water up to 2 ml - 10% Boehringer blocking reagent
1 g blocking reagent powder in a bottle
Filtered maleic acid buffer, pH 7.5 up to 10 ml
Autoclave to dissolve and store at 4 °C - Filtered maleic acid buffer, pH 7.5
11.61 g maleic acid (100 mM)
8.76 g NaCl (150 mM)
Adjust pH to 7.5 by NaOH
Autoclave, filtrate and store at RT - Tris-HCl 2 M, pH 9.5
- Dissolve 121.1 g of Tris-base in 400 ml of deionized water
- Adjust the pH to the desired value by adding concentrated HCl
- Allow the solution to cool to room temperature before making final adjustments to the pH
- Adjust the volume of the solution to 500 ml with deionized water
- Dispense into 50 ml aliquots and sterilize by autoclaving
- Dissolve 121.1 g of Tris-base in 400 ml of deionized water
- Alkaline phosphatase detection buffer pH 9.5 (APB)
50 ml Tris-HCl 2 M pH 9.5 (100 mM final)
5.85 g NaCl (100 mM final)
10.17 g MgCl2·7H2O (50 mM final)
Nuclease-free water up to 1 L
Aliquot, autoclave and store at room temperature
Acknowledgments
This protocol has been adapted from de Boer et al. (1998) and Rosso et al. (1999). It has made possible the study of the expression profile of several dozens of genes (Jaubert et al., 2002a and 2002b; Neveu et al., 2003; Jaubert et al., 2004; Ledger et al., 2006; Dubreuil et al., 2007; Jaouannet et al., 2012; Jaouannet et al., 2013; Danchin et al., 2013; Nguyen et al., 2017). These works were funded by INRA, CNRS, University of Nice Sophia Antipolis, the Region Provence-Alpes-Côte d’Azur, the French Government, the European Union and NATO Collaborative Research Grant. C-N.N. was supported by USTH fellowships, 911-USTH program, from the Ministry of Education and Training of The Socialist Republic of Vietnam. The authors have no conflict of interest or competing interests to declare.
References
- Caillaud, M. C. and Favery, B. (2016). In vivo imaging of microtubule organization in dividing giant cell. Methods Mol Biol 1370: 137-144.
- Danchin, E. G. J., Arguel, M.-J., Campan-Fournier, A., Perfus-Barbeoch, L., Magliano, M., Rosso, M.-N., Da Rocha, M., Da Silva, C., Nottet, N., Labadie, K., Guy, J., Artiguenave, F. Abad, P. (2013). Identification of novel target genes for safer and more specific control of root-knot nematodes from a pan-genome mining. PLoS Pathog 9(10): e1003745.
- de Boer, J. M., Yan, Y., Smant, G., Davis, E. L. and Baum, T. J. (1998). In-situ hybridization to messenger RNA in Heterodera glycines. J Nematol 30(3): 309-312.
- Dubreuil, G., Magliano, M., Deleury, E., Abad, P. and Rosso, M. N. (2007). Transcriptome analysis of root-knot nematode functions induced in the early stages of parasitism. New Phytol 176(2): 426-436.
- Jaouannet, M., Magliano, M., Arguel, M. J., Gourgues, M., Evangelisti, E., Abad, P. and Rosso, M. N. (2013). The root-knot nematode calreticulin Mi-CRT is a key effector in plant defense suppression. Mol Plant Microbe Interact 26(1): 97-105.
- Jaouannet, M., Perfus-Barbeoch, L., Deleury, E., Magliano, M., Engler, G., Vieira, P., Danchin, E. G., Da Rocha, M., Coquillard, P., Abad, P. and Rosso, M. N. (2012). A root-knot nematode-secreted protein is injected into giant cells and targeted to the nuclei. New Phytol 194(4): 924-931.
- Jaubert, S., Laffaire, J. B., Abad, P. and Rosso, M. N. (2002a). A polygalacturonase of animal origin isolated from the root-knot nematode Meloidogyne incognita. FEBS Lett 522(1-3): 109-112.
- Jaubert, S., Laffaire, J. B., Ledger, T. N., Escoubas, P., Amri, E. Z., Abad, P. and Rosso, M. N. (2004). Comparative analysis of two 14-3-3 homologues and their expression pattern in the root-knot nematode Meloidogyne incognita. Int J Parasitol 34(7): 873-880.
- Jaubert, S., Ledger, T. N., Laffaire, J. B., Piotte, C., Abad, P. and Rosso, M. N. (2002b). Direct identification of stylet secreted proteins from root-knot nematodes by a proteomic approach. Mol Biochem Parasitol 121(2): 205-211.
- Ledger, T. N., Jaubert, S., Bosselut, N., Abad, P. and Rosso, M. N. (2006). Characterization of a new β-1,4-endoglucanase gene from the root-knot nematode Meloidogyne incognita and evolutionary scheme for phytonematode family 5 glycosyl hydrolases. Gene 382: 121-128.
- Liao, Y., Fung, T. S., Huang, M., Fang, S., Zhong, Y. and Liu, D. (2014). RNA isolation and Northern blot analysis. Bio-protocol 4(6): e1077.
- Neveu, C., Jaubert, S., Abad, P. and Castagnone-Sereno, P. (2003). A set of genes differentially expressed between avirulent and virulent Meloidogyne incognita near-isogenic lines encode secreted proteins. Mol Plant Microbe Interact 16(12): 1077-1084.
- Nguyen, C. N., Perfus-Barbeoch, L., Quentin, M., Zhao, J., Magliano, M., Marteu, N., Da Rocha, M., Nottet, N., Abad, P. and Favery, B. (2017). A root-knot nematode small glycine and cysteine-rich secreted effector, MiSGCR1, is involved in plant parasitism. New Phytol.
- Rosso, M. N., Favery, B., Piotte, C., Arthaud, L., De Boer, J. M., Hussey, R. S., Bakker, J., Baum, T. J. and Abad, P. (1999). Isolation of a cDNA encoding a β-1,4-endoglucanase in the root-knot nematode Meloidogyne incognita and expression analysis during plant parasitism. Mol Plant Microbe Interact 12(7): 585-591.
Article Information
Copyright
© 2018 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Jaouannet, M., Nguyen, C., Quentin, M., Jaubert-Possamai, S., Rosso, M. and Favery, B. (2018). In situ Hybridization (ISH) in Preparasitic and Parasitic Stages of the Plant-parasitic Nematode Meloidogyne spp.. Bio-protocol 8(6): e2766. DOI: 10.21769/BioProtoc.2766.
Category
Microbiology > Microbe-host interactions > Nematode
Plant Science > Plant immunity > Host-microbe interactions
Cell Biology > Cell staining > Nucleic acid
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link