- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Automatic Leaf Epidermis Assessment Using Fourier Descriptors in Texture Images
Published: Vol 7, Iss 23, Dec 5, 2017 DOI: 10.21769/BioProtoc.2630 Views: 7155
Reviewed by: Trinadh Venkata Satish TammanaAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
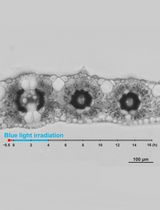
Live Leaf-Section Imaging for Visualizing Intracellular Chloroplast Movement and Analyzing Cell–Cell Interactions
Yuta Kato [...] Mitsutaka Taniguchi
Aug 5, 2025 2312 Views

Live-Cell Monitoring of Piecemeal Chloroplast Autophagy
Masanori Izumi [...] Shinya Hagihara
Nov 5, 2025 1688 Views
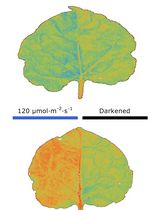
Chloroplast Movement Imaging Under Different Light Regimes With a Hyperspectral Camera
Paweł Hermanowicz [...] Justyna Łabuz
Dec 20, 2025 749 Views
Abstract
The identification of plant species is not a trivial task, since it is carried out by experts and depends on the presence of fruits, flowers and leaves. However, fruits and flowers are not available throughout the year, while leaves are accessible most of the year. In order to assist the specialized work of species identification, methods of texture image analysis are used to extract characteristics from samples of imaged leaves and thus predict the species. Texture image analysis is a versatile and powerful technique able to extract measurements from patterns in the images. Using this technique, recent research has found a close relationship between texture and plant species (da Silva et al., 2015 and 2016). Here, we describe the procedure to extract texture features from microscopic images of leaves using Fourier (Cosgriff, 1960; Azencott, 1997; Gonzalez and Woods, 2006). It is important to highlight that other methods for texture extraction can be used as well.
This protocol is split into two parts: (A) leaf epidermal dissociation and (B) automatic method for leaf epidermal image analysis.
Background
The protocol for plant material dissociation was described by Franklin (1945) for wood; botanical anatomists have adapted the method for leaf epidermal dissociation. Dissociation methods allow the visualization of the epidermal cells as well as their attachments in a clearer way than diaphanization and epidermal imprints methods. In addition to the dissociation method described herein, there are others using Jeffrey’s solution (1:1 solution of chromic acid and nitric acid, each at 10%) or only nitric acid (Johansen, 1940), but these methods use strong acids and more hazardous to human health and the environment. Moreover, in our experience, epidermal dissociation with Franklin solution is more effective for a greater number of species.
As leaf epidermal traits have a great taxonomic value, texture analysis from epidermal images can be a suitable method for plant identification, once texture analysis is a powerful technique that allows the extraction of measurements from patterns in the images. In the study realized by (da Silva et al., 2016), three different methods were used in the process of texture feature extraction from leaf epidermal images: Fourier descriptors, corrosion-inspired texture analysis and local binary patterns; among them, Fourier descriptors were the best option, achieving higher success rate in plant species identification. A brief description of the methods for epidermal dissociation and texture feature extraction from microscopic epidermal images using Fourier descriptors has been reported in (da Silva et al., 2016).
Materials and Reagents
- Razor blade
- Glass vial
- Petri dish
- Needles and brush
- Glass slide (Perfecta, catalog number: 7105 )
- Cover glass (Perfecta, catalog number: 7004 )
- Gloves
- Leaf samples of different plant species (see species studied in Supplementary Table 1)
- Glacial acetic acid (Dinâmica Química, catalog number: 1242 )
- Hydrogen peroxide (30 volumes) (Dinâmica Química, catalog number: 2138 )
- Glycerin (Chem-Impex International, catalog number: 00599 )
- Safranin (Vetec, catalog number: 379 )
- 1% aqueous Safranin solution (see Recipes)
Equipment
- Analytical balance (Shimadzu, model: AUY220 )
- Oven (Fanem, model: 315 SE )
- Light microscope (ZEISS, model: Axio Scope.A1 ) with equipment for photomicrography (ZEISS, model: Axiocam ICc3 )
Software
- MATLAB R2014a (Version 8.3) (MATLAB and Statistics Toolbox Release, 2014a)
- Weka 3 (Hall et al., 2009), a data mining software in Java
Procedure
- Leaf epidermal dissociation
- With the aid of a razor blade, remove from each leaf two or three samples of approximately 1 cm2 but less from its middle region, including the midrib (Figure 1A). Fresh or dry leaves can be used. Put the leaf fragments in a glass vial containing 1:1 glacial acetic acid and hydrogen peroxide (Franklin solution) in a drying oven at 60 °C (add enough volume so that the vial is left with about 1 cm of solution). Generally, this procedure is completed when the epidermis is beginning to peel off from the leaf fragments or when the leaf fragments become transparent and with air bubbles inside (Figures 1B-1E). The procedure can last from 12 h to a few days, depending on the morphological characteristics of each leaf. Some plants with thicker and sclerified leaves will require higher temperatures (up to 90 °C). Thus, if a leaf fragment incubated at 60 °C fails to produce an isolated epidermis, it must be heated up to 80-90 °C (it is not necessary to change the dissociation solution, however, it may be necessary to add more solution as it evaporates more at higher temperatures).
Notes:- For example, the species Clusia criuva, Duguetia furfuracea and Roupala montana required a temperature of 90 °C for leaf epidermal dissociation.
- We recommend that the fragments be removed from the middle region of the leaves by standardization; this region is usually used by botanists in leaf anatomy studies. It is possible to have differences in the size and density of the stomata according to the sampled region of the leaf.
- For example, the species Clusia criuva, Duguetia furfuracea and Roupala montana required a temperature of 90 °C for leaf epidermal dissociation.
- After this procedure, pour a little of the solution along with the leaf fragments in a Petri dish containing distilled water. Separate the adaxial and abaxial faces of the epidermis from the leaf fragment with a brush or needles (Figure 1F). In Petri dish, never remove the epidermis from the aqueous medium to maintain it stretched. Put the abaxial epidermis on a slide with a needle; the outer side of the epidermis must be placed upwards on the slide (Figure 1G). Whenever you deal with the epidermis on the slide, the epidermis must be stretched.
Note: The inclusion of the midrib in the leaf fragment helps to distinguish between the adaxial and abaxial surfaces of the epidermis. - To remove residues of glacial acetic acid and hydrogen peroxide solution, wash the fragment with distilled water holding the epidermis with a needle. Remove water excess from the slide with soft paper. Put one drop of 1% aqueous Safranin (see Recipes) on the epidermis (Figure 1H), stain for 10 min. Then, wash the epidermis with distilled water to remove the stain excess. Dry the slide with soft paper and, for mounting, put one to two drops of glycerin on the epidermis and carefully put the cover glass (Figure 1I).
Note: Use gloves and protection glasses whenever you deal with Franklin solution or its components. Use gloves whenever you deal with Safranin. - The epidermis is observed and photographed under an optical microscope. Since the plant epidermis has distinct characteristics, mainly of thickness, each image must be taken with adequate light intensity so that it does not become dark or too clear. All the images were taken at 200x magnification.
- The synthesis of the procedure is presented in Figure 1.
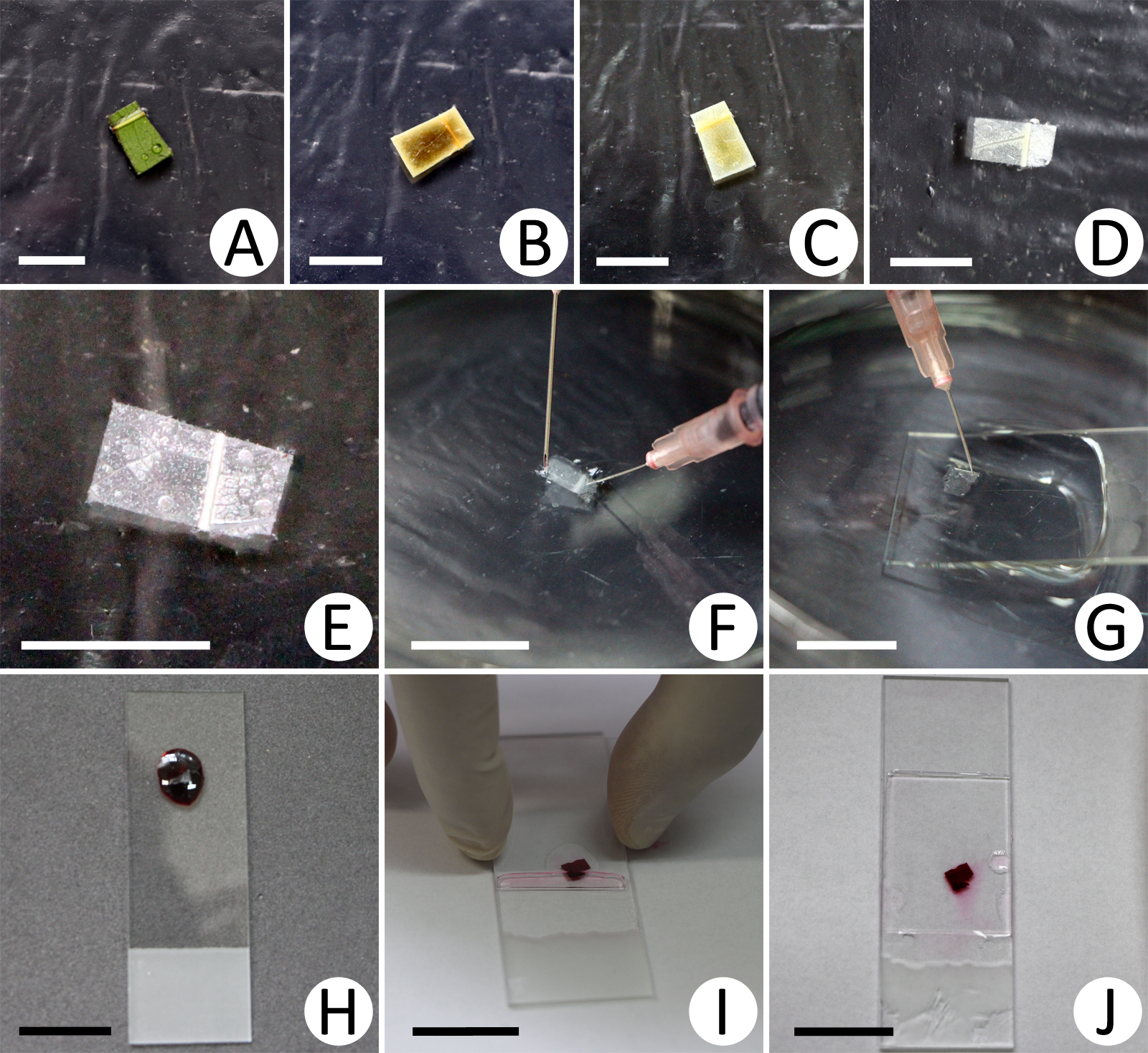

Figure 1. Procedure for epidermal decoupling. A. Fresh leaf fragment; B-D. Stages of mesophyll digestion by the solution of glacial acetic acid and hydrogen peroxide; E. Leaf fragment ready for epidermal separation; F. Faces of the epidermis being dissociated from leaf fragment; G. Epidermis being placed on the slide; H. Staining; I. Mounting; J. Ready slide. Scale bars: A-E = 1 cm; F-J = 2 cm.
- With the aid of a razor blade, remove from each leaf two or three samples of approximately 1 cm2 but less from its middle region, including the midrib (Figure 1A). Fresh or dry leaves can be used. Put the leaf fragments in a glass vial containing 1:1 glacial acetic acid and hydrogen peroxide (Franklin solution) in a drying oven at 60 °C (add enough volume so that the vial is left with about 1 cm of solution). Generally, this procedure is completed when the epidermis is beginning to peel off from the leaf fragments or when the leaf fragments become transparent and with air bubbles inside (Figures 1B-1E). The procedure can last from 12 h to a few days, depending on the morphological characteristics of each leaf. Some plants with thicker and sclerified leaves will require higher temperatures (up to 90 °C). Thus, if a leaf fragment incubated at 60 °C fails to produce an isolated epidermis, it must be heated up to 80-90 °C (it is not necessary to change the dissociation solution, however, it may be necessary to add more solution as it evaporates more at higher temperatures).
- Leaf epidermis image analysis
- The computational and statistical methods have been implemented using the software MATLAB R2014a (Version 8.3) (MATLAB and Statistics Toolbox Release, 2014a) and the classification procedures have been performed using Weka 3 (Hall et al., 2009), a data mining software in Java.
- Pre-processing method
- The first step is to convert the color image into a gray scale image. The pre-processing procedure was conducted by performing a histogram stretching to increase the image contrast. Let an image f (x, y), g(x, y) be the enhanced image of f (x, y) given by
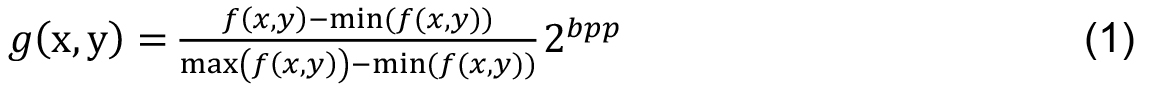
where, bpp is the number of bits per pixel of the image f (x, y). In one image with 256 gray levels, bpp = 8. In the enhanced image g(x, y), 1% of data is saturated at low and high intensities of the original image. - Next frame shows the code in MATLAB to obtain the stretching of the input image.

- Figure 2 shows two examples, using Baccharis linearifolia and Esenbeckia pulmila, of the pre-processing stage.
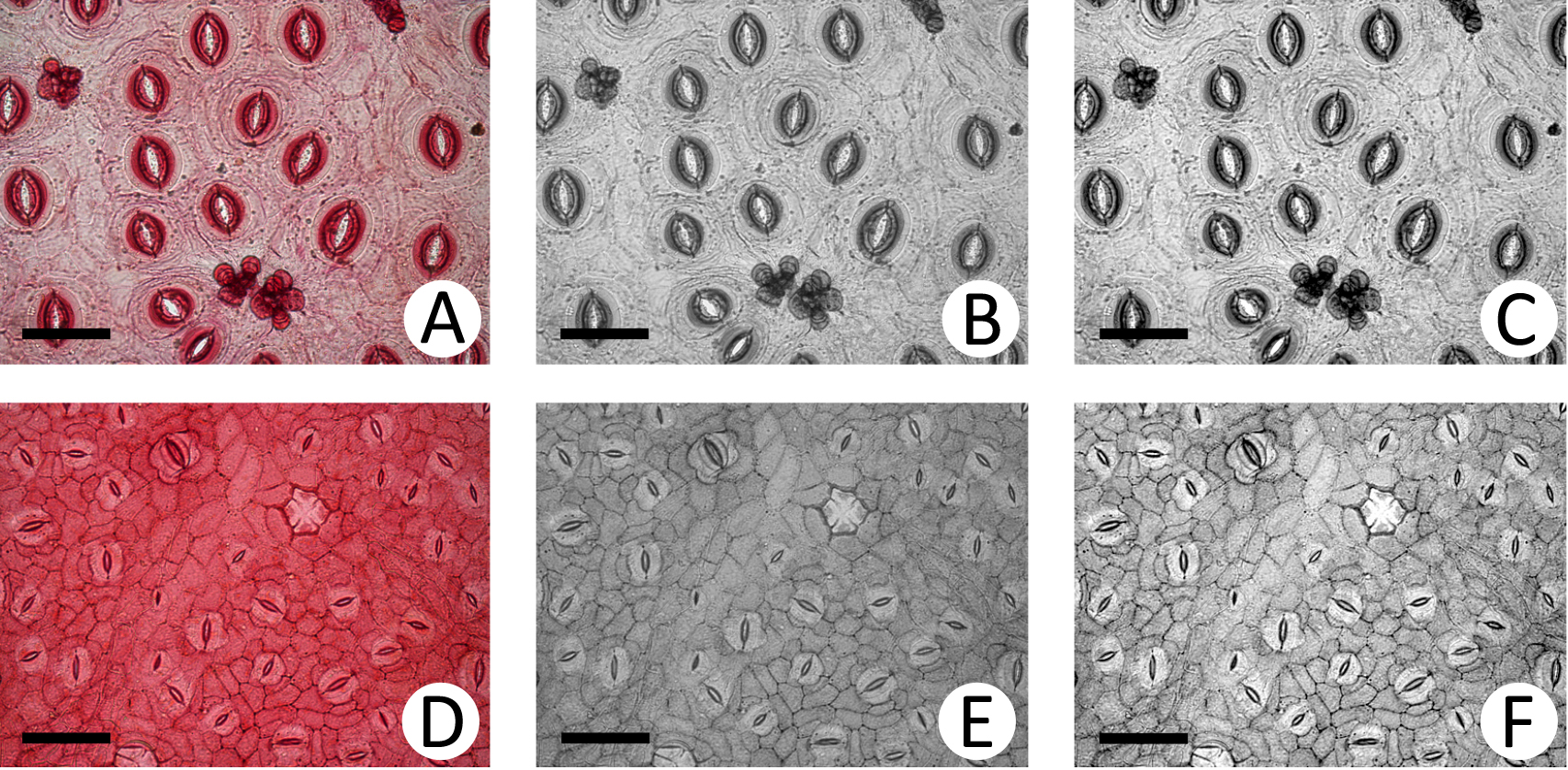
Figure 2. Pre-processing. Images of Baccharis linearifolia (A-C) and Esenbeckia pulmila (D-F). A and D: Original images; B and E: Gray images of the first column; C and F: Stretching of the images in the second column. Images are at magnification of 200x (scale bars = 50 µm).
- The first step is to convert the color image into a gray scale image. The pre-processing procedure was conducted by performing a histogram stretching to increase the image contrast. Let an image f (x, y), g(x, y) be the enhanced image of f (x, y) given by
- Image texture characterization
As an image is represented as numerical matrix, a variety of methods can be used to extract patterns of texture in images. In this protocol, Fourier descriptors will be described.
Fourier Descriptors- Proposed by Cosgriff (1960), Fourier descriptors represent periodic functions given by the sum of sines and cosines of a periodic signal obtained from the discrete Fourier Transform (DFT). Discrete Fourier transform of a function 1D f (x) is given by:
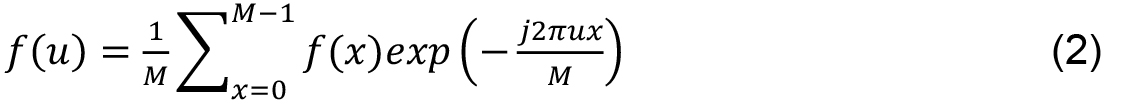
M is the length of the signal, u, x ∈ [0, 1..., M - 1] and j is a complex number. Inverse transform, from F (u) to f (x) is given by: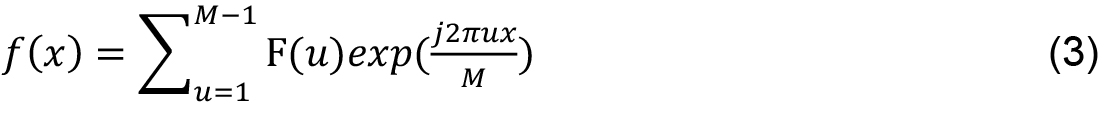
For two-dimensional functions, as images, the Fourier transform is given by: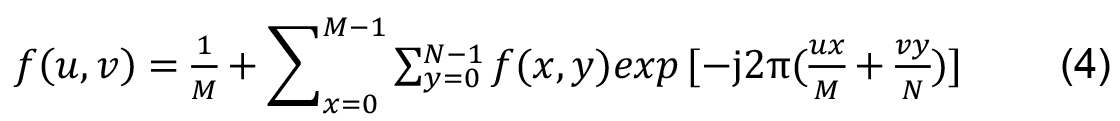
M and N are the signal dimensions. Inverse transform of a 2D signal is: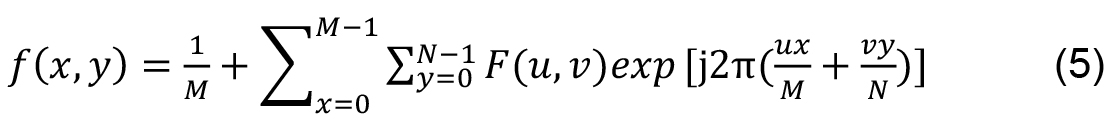
- When the Fourier Transform is applied to the image, lower frequency coefficients remain at the extremities of the spectrum. The Figure 3A shows the original image and the Figure 3B, the spectrum of the Fourier transformation. Then, a shift operation is performed on the resultant image moving the origin of the Fourier Transform to the central coordinates ([M/2] for 1D signals and [M/2, N/2] for 2D signals) at frequency domain, as shown in Figure 3C.
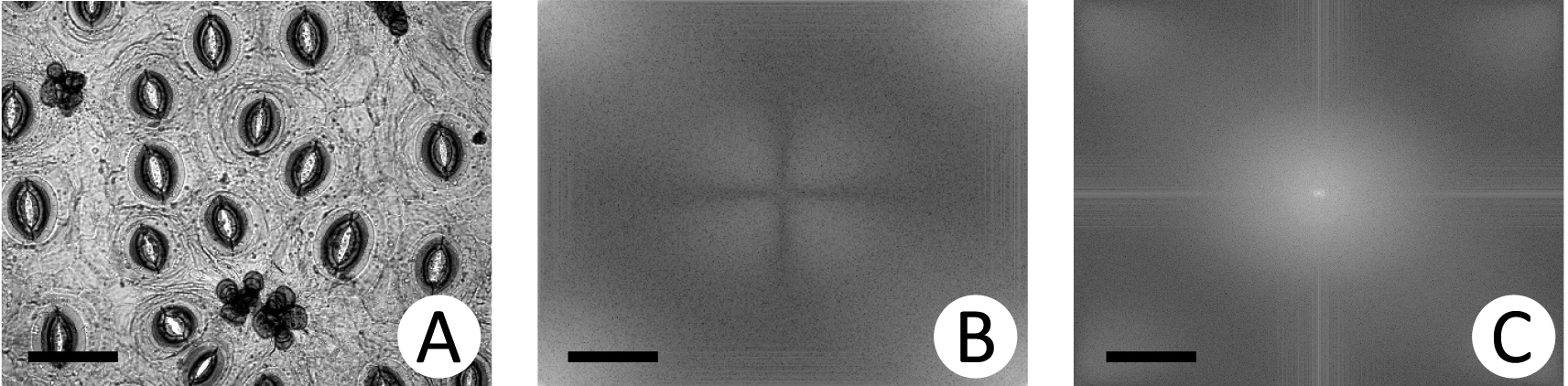

Figure 3. Fourier Transform. A. Original image; B. Fourier transform of (A); C. Shift of (B). Images are at magnification of 200x (scale bars = 50 µm). - Low frequency components, found at the center of the shifted Fourier Transform, contain the most relevant information of the behavior of a signal. High frequency components are found at the extremities and represent abrupt changes and noise. These complex values can be used as characteristics of texture (Azencott et al., 1997; Gonzalez, and Woods, 2006). Here, the central values are prioritized as can be seen in the following. Two approaches are used to obtain the characteristics, also called descriptors, of the image texture.
- Fourier Circular: After shift operation, G circular rings are used to obtain the sum of all spectrum absolute values from the origin to each circular ring. G is given by min ([M/2J], [N/2J]), where the image has M x N pixels size. So, radius equal to 1, 2..., G providing G descriptors. The circular rings can be seen in Figure 4A.
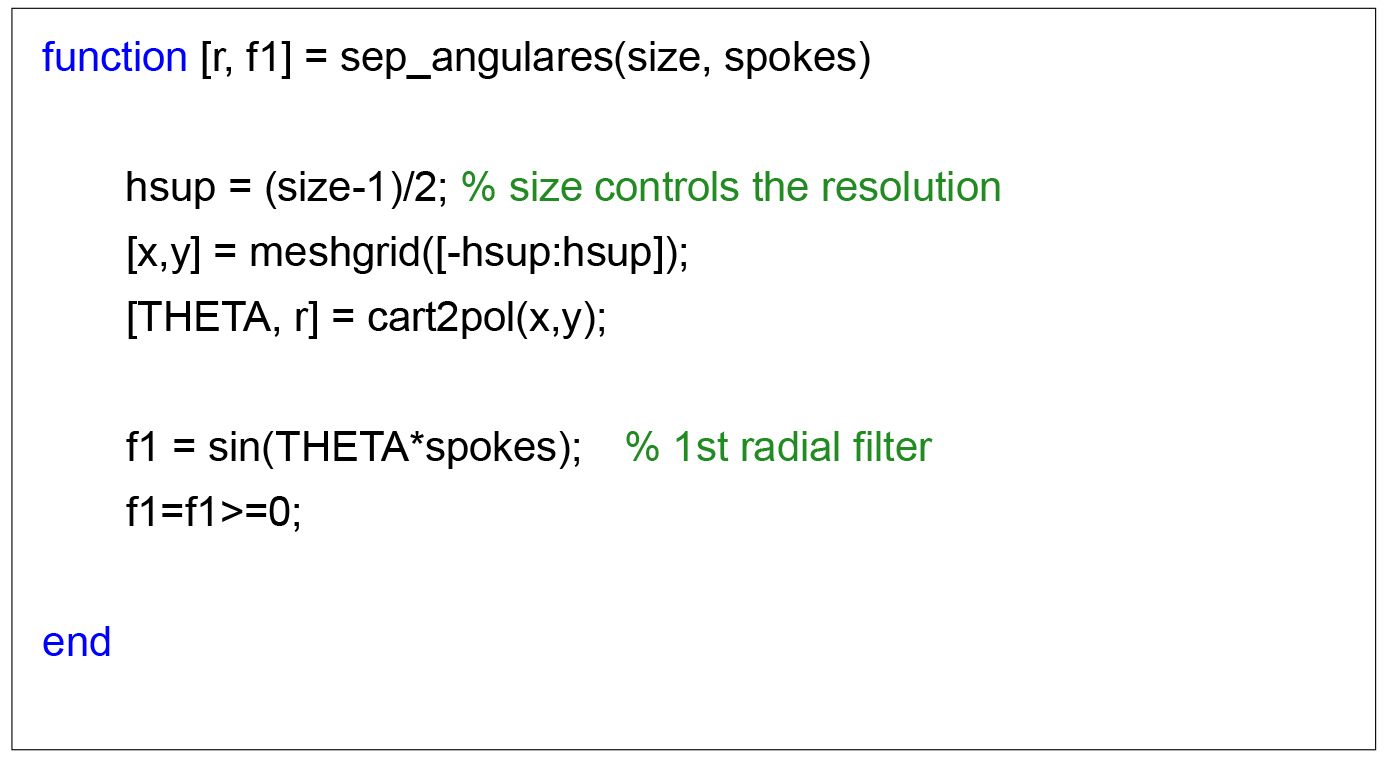
Fourier Circular-Angular Complementing the circular rings, F (u, v) is partitioned into eight angles equally spaced over the image, as in Figure 4B, resulting in Figure 4C. Eight circular rings have been used with radius equal to 3, 6, 9, 12, 15, 18, 21 and 24 pixels of distance. Thus 64 sectors are obtained, totaling 64 descriptors by summing the spectrum absolute values of each sector. The order in which the descriptors vector is composed does not interfere with the result. In this implementation, check the code list, the radius loop is inside the quadrant loop.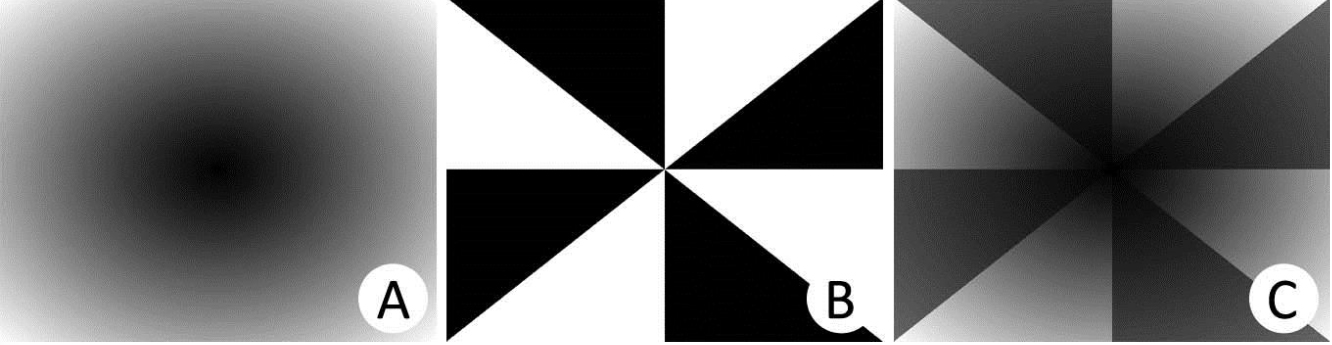
Figure 4. Circular and angular rings to obtain the Fourier descriptors. A. Circular; B. Angular; C. Circular-Angular. - The next three frames show the complete code in MATLAB to obtain the Fourier Circular-Angular descriptors.



- Proposed by Cosgriff (1960), Fourier descriptors represent periodic functions given by the sum of sines and cosines of a periodic signal obtained from the discrete Fourier Transform (DFT). Discrete Fourier transform of a function 1D f (x) is given by:
- The computational and statistical methods have been implemented using the software MATLAB R2014a (Version 8.3) (MATLAB and Statistics Toolbox Release, 2014a) and the classification procedures have been performed using Weka 3 (Hall et al., 2009), a data mining software in Java.
Data analysis
- For each image of the leaf, a set of characteristics is extracted to represent it. Then, these characteristics are compared to identify to which species of plant this leaf belongs. To proceed with the experiments, the species of the samples in the training set are known and the goal is to predict the species of the test set that are unknown. The descriptors of one image belonging to the test set are compared to all samples of the training set using the Nearest Neighbor algorithm (Aha et al., 1991) to predict the species. The species is assigned to a given sample according to the closest sample in the training set.
- The strategy to distribute the samples in a training and test set is the k-fold cross-validation scheme (Hastie et al., 2001), represented in Figure 5. In this approach, the set of samples is equally divided into k folds. Then, k-1 folds are joined forming the training set while the remaining fold is assigned to the test set. This procedure is performed k times varying the test fold. At the end, all the samples have been classified generating the success rate by calculating the ratio between the number of samples correctly classified by the number of samples.
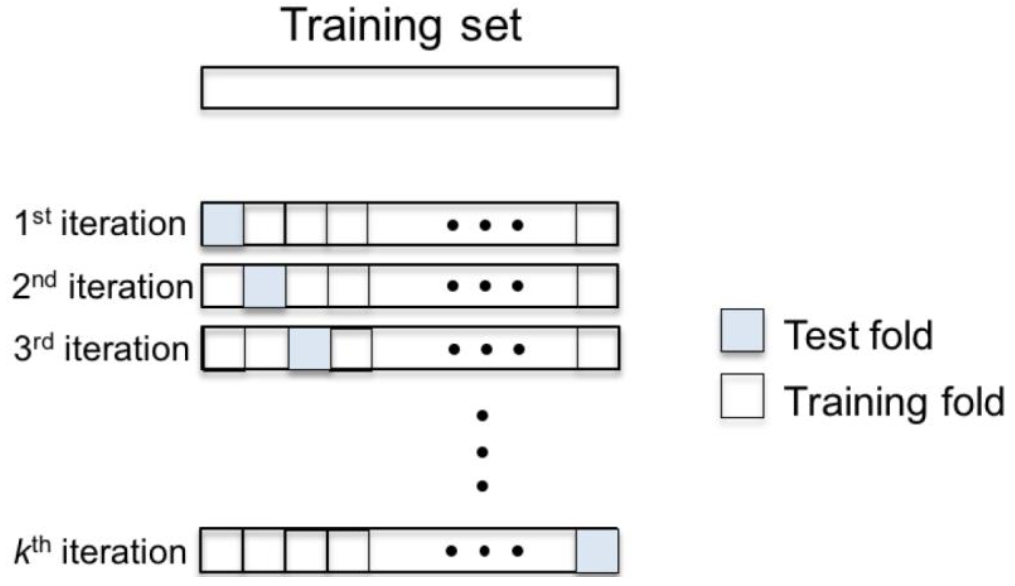
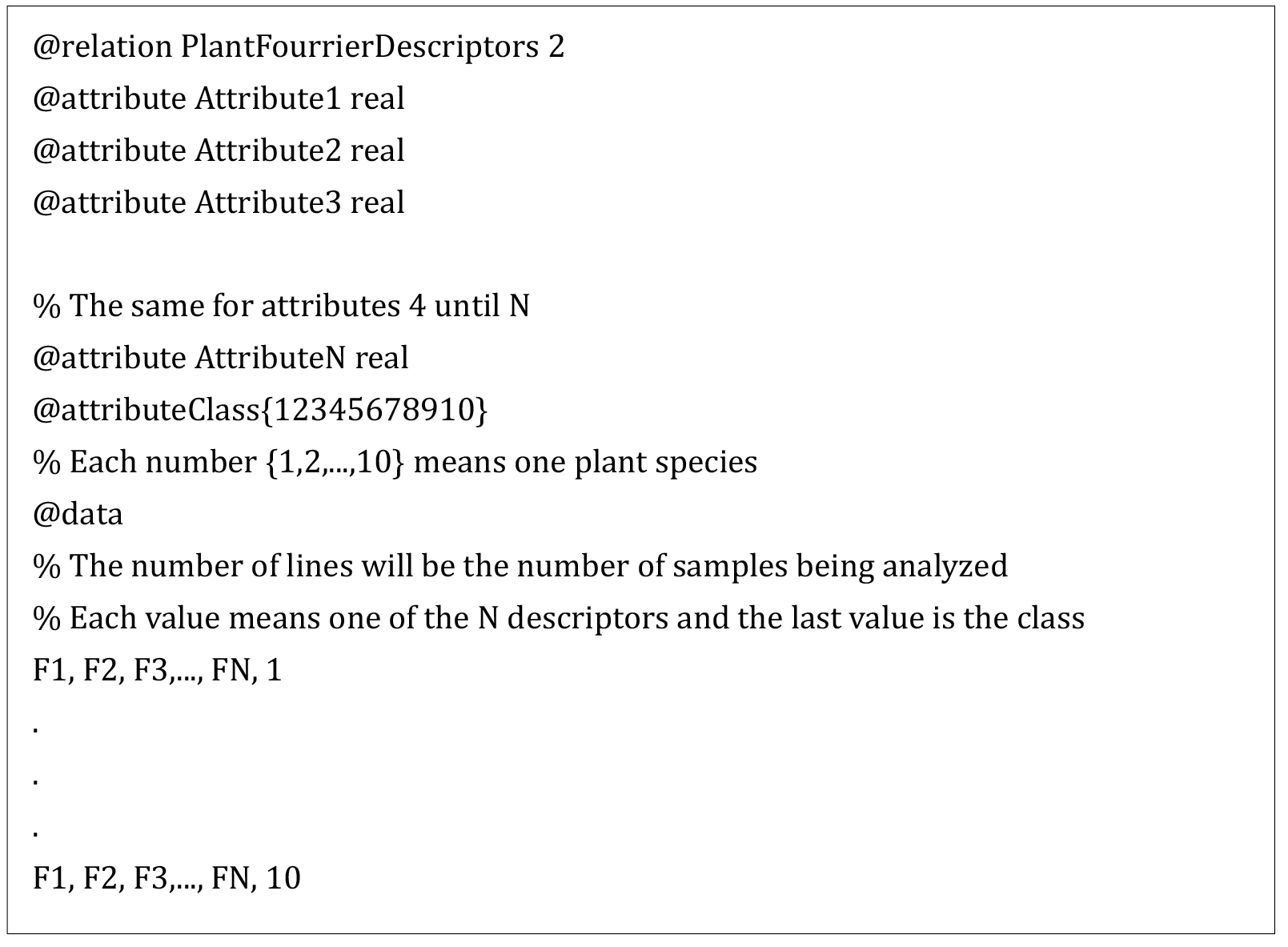
Figure 5. k-fold cross-validation scheme - The data analysis procedure can be summarized by creating a file with all the characteristics of each sample to be read by Weka and perform the classification. Next frame shows the template to construct the file to be read by Weka to perform the classification of the samples.

- Synthesis of the computer procedure
- Pre-process all the images to enhance the contrast of the image.
- Extract characteristics using Fourier descriptors from each image, as shown in Figure 6.
- Classify all the samples using Nearest Neighbor in a k-fold cross validation scheme and check the success rate.
- Training the system and apply it to the leaf identification or analysis.
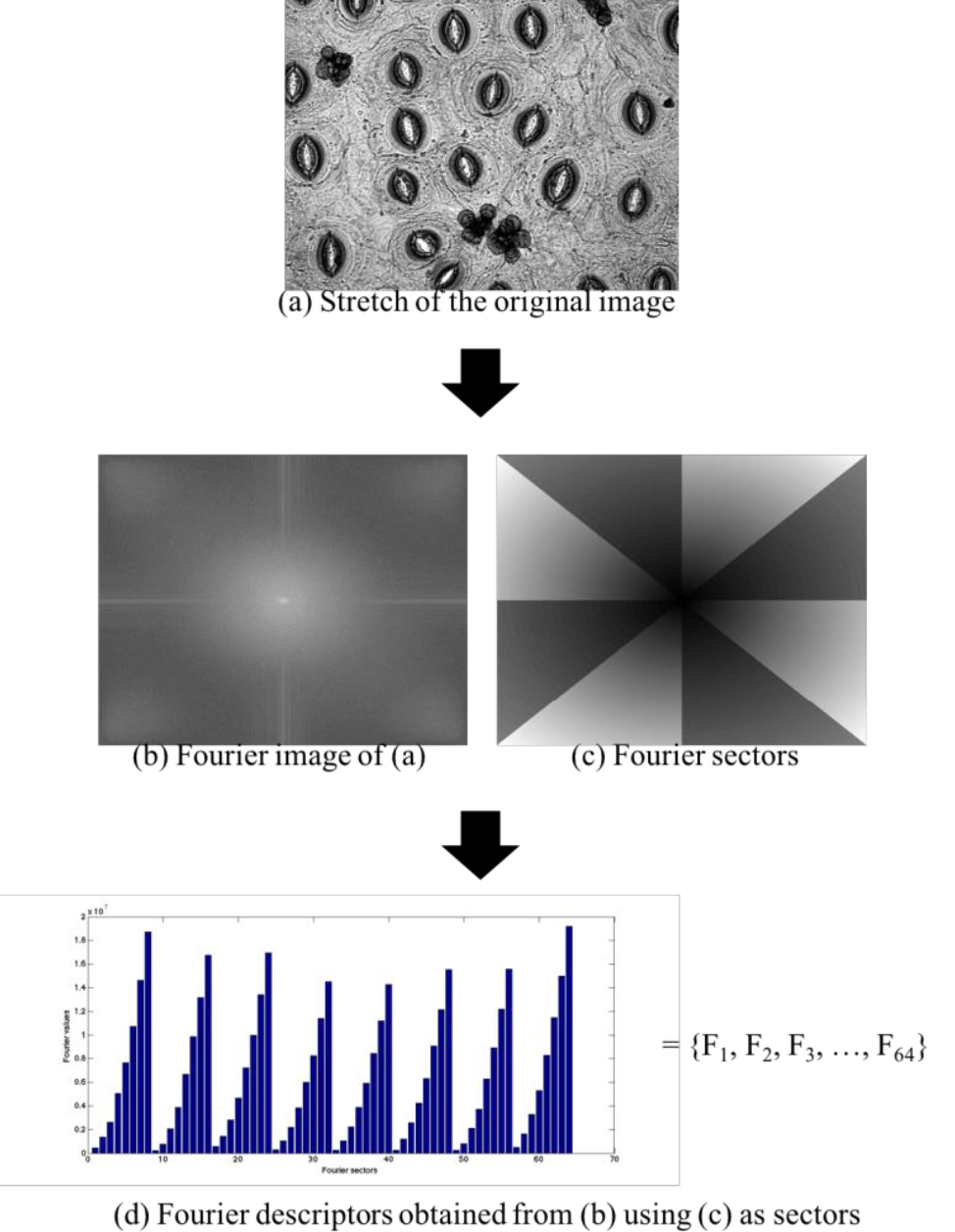
Figure 6. Extraction of characteristics using Fourier descriptors
- Pre-process all the images to enhance the contrast of the image.
Notes
The success in dissociation of the epidermis in different species depends on the level of individual training. Each species behaves differently during the procedure. As necessary, oven temperature can be increased to speed up and to facilitate the process. Thus, the temperature can influence the result; thicker leaves usually require higher temperatures (up to 90 °C) than thinner ones (60 °C).
Recipes
- 1% aqueous Safranin solution
Add 1 g Safranin in 100 ml distilled water in a bottle
Mix well and store in the dark in a refrigerator (can be kept for several months under these conditions)
Acknowledgments
This protocol was adapted from da Silva et al. (2016). The authors gratefully thank the financial support of São Paulo Research Foundation (FAPESP), with grant Nos.: 2011/01523-1, 2011/23112-3 and 2011/21467-9, National Council for Scientific and Technological Development (CNPq) with grant Nos.: 307797/2014-7 and 484312/2013-8 and PROPE/UNESP (14/2012/Renove), and Coordination for the Improvement of Higher Education Personnel (CAPES). The authors declare that there are no conflicts of interest or competing interest.
References
- Aha, D. W., Kibler, D. and Albert, M. K. (1991). Instance-based learning algorithms. Mach Learn 6(1): 37-66.
- Azencott, R. Wang, J. P. and Younes, L. (1997). Texture classification using windowed Fourier filters. IEEE T Pattern Anal 19(2):148-153.
- Cosgriff, R. L. (1960). Identification of shape. Ohio State University Research Foundation, Columbus. ASTIA AD 254: 792.
- da Silva, N. R., da Silva Oliveira M. W., Almeida Filho, H. A., Pinheiro, L. F. S., Rossatto, D. R. Kolb, R. M. and Bruno, O. M. (2016). Leaf epidermal images for robust identification of plants. Sci Rep 6:25994.
- da Silva, N. R., Florindo, J. B., Gómez, M. C., Rossatto, D. R., Kolb, R. M. and Bruno, O. M. (2015). Plant identification based on leaf midrib cross-section images using fractal descriptors. PLoS One 10(6): e0130014.
- Franklin, G. L. (1945). Preparation of thin sections of synthetic resins and wood-resin com- posites, and a new macerating method for wood. Nature 155(3924):51.
- Gonzalez, R. C. and Woods, R. E. (2006). Digital Image Processing (3rd Edition). Prentice-Hall.
- Hall, M., Frank, E. Holmes, G. Pfahringer, B. Reutemann, P. and Witten, I. H. (2009). The weka data mining software: An update. ACM SIGKDD Explorations Newsletter 11(1): 10-18.
- Hastie, T. Tibshirani, R. and Friedman, J. (2001). The Elements of Statistical Learning. Springer.
- Johansen, D. A. (1940). Plant microtechnique. McGraw-Hill.
- MATLAB and Statistics Toolbox Release. (2014a). The MathWorks, Inc., Natick, Mas-sachusetts, United States.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
da Silva, N. R., Oliveira, M. W. D. S., Filho, H. A. D. A., Pinheiro, L. F. S., Kolb, R. M. and Bruno, O. M. (2017). Automatic Leaf Epidermis Assessment Using Fourier Descriptors in Texture Images. Bio-protocol 7(23): e2630. DOI: 10.21769/BioProtoc.2630.
Category
Plant Science > Plant cell biology > Cell imaging
Cell Biology > Tissue analysis > Tissue imaging
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link








