- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
A Novel Protocol to Quantitatively Measure the Endocytic Trafficking of Amyloid Precursor Protein (APP) in Polarized Primary Neurons with Sub-cellular Resolution
Published: Vol 7, Iss 23, Dec 5, 2017 DOI: 10.21769/BioProtoc.2629 Views: 7524
Reviewed by: Yanjie LiAlessandro DidonnaAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Intraepidermal Nerve Fiber Quantification of the Mouse Hind Paw Footpads: A Detailed and Simplified Protocol
Anastasia Yerushkin [...] Amir Dori
Dec 5, 2025 1224 Views
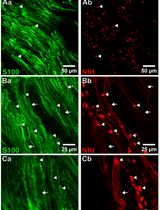
Whole-Mount Immunostaining for the Visual Separation of A- and C-Fibers in the Study of the Sciatic Nerve
Valeriia Ustymenko [...] Nana Voitenko
Dec 5, 2025 1421 Views
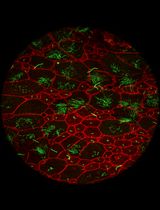
Whole-Mount Visualization of Primary Cilia in the Developing Mouse Brain
Oscar Torres Gutierrez [...] Xuecai Ge
Dec 20, 2025 1003 Views
Abstract
Alzheimer’s disease’s established primary trigger is β-amyloid (Aβ) (Mucke and Selkoe, 2012). The amyloid precursor protein (APP) endocytosis is required for Aβ generation at early endosomes (Rajendran and Annaert, 2012). APP retention at endosomes depends on its sorting for degradation in lysosomes (Haass et al., 1992; Morel et al., 2013; Edgar et al., 2015; Ubelmann et al., 2017). The following endocytosis assay has been optimized to assess the amyloid precursor protein (APP) endocytosis and degradation by live murine cortical primary neurons (Ubelmann et al., 2017).
Keywords: APPBackground
Aβ42 accumulation is a primary trigger of Alzheimer’s disease. APP endocytosis is required for Aβ42 generation (Koo and Squazzo, 1994; Grbovic et al., 2003; Cirrito et al., 2008; Rajendran et al., 2008). The endocytosis of APP has been analysed in pulse-chase kinetic experiments in bulk by classical biotinylation of surface proteins (Sannerud et al., 2011; Xiao et al., 2012; Sullivan et al., 2014), in single cells by specific labelling of surface APP using antibodies against N-terminal extracellular domain of APP (Yamazaki et al., 1995; Xiao et al., 2012). The majority of these studies used non-neuronal cells (Yamazaki et al., 1996; Lee et al., 2008; Sullivan et al., 2014), and neuronal-like cell lines (Xiao et al., 2012), few used primary neurons (Yamazaki et al., 1995; Sullivan et al., 2014). Primary neurons differentiate like in vivo axons and dendrites, with their specialized presynaptic terminals and post-synaptic compartments. However, careful measurements and distinction between these neuronal compartments are lacking in these reports. We developed a method of analysing APP endocytosis in the different neuronal compartments, the soma or cell body, dendrites and axons that we describe in this bio-protocol. Our protocol details the procedure for following and measuring APP endocytosis in polarized neurons using classical immunofluorescence and semi-quantitative cell biology analysis methods.
We believe our method will allow the field to move forward by reliably measuring semi-quantitatively the compartmentalized endocytosis of APP specific to polarized neurons.
Materials and Reagents
- 24-well dishes (SARSTEDT, catalog number: 83.1836 ) for mammalian cell culture
- Circular glass coverslips, 13 mm (VWR, Marienfeld, catalog number: 630-1597 )
Note: Autoclaved, pre-washed with 40% ethanol/60% HCl for 1 h at RT and washed 4 times, 15 min each, with Milli-Q water at RT; coated overnight with 200 µl 0.1% (w/v) poly-D-lysine at 37 °C in a 5% CO2 and 20% O2 humidified incubator and washed 3 x with sterile Milli-Q water. - Superfrost glass slides (MENZEL GERHARD, catalog number: 2586E )
- Plastic Pasteur pipette (SARSTEDT, catalog number: 86.1171 )
- Parafilm (Fisher Scientific, catalog number: 11782644)
Manufacturer: Bemis, Parafilm, catalog number: PM999 . - Wild-type females and males mouse embryos (Balbc, embryonic day 16; Charles River)
- APP-RFP plasmid (Szodorai et al., 2009) (S. Kins, University of Kaiserslautern)
- 0.1% (w/v) poly-D-lysine (Sigma-Aldrich, catalog number: P1149 )
- Plating medium:
- DMEM, high glucose, pyruvate (Thermo Fisher Scientific, GibcoTM, catalog number: 11995065 )
- 10% fetal bovine serum (FBS), qualified, heat inactivated, US origin (Thermo Fisher Scientific, GibcoTM, catalog number: 16140071 )
- 1% penicillin-streptomycin (10,000 U/ml) (Thermo Fisher Scientific, GibcoTM, catalog number: 15140122 )
- DMEM, high glucose, pyruvate (Thermo Fisher Scientific, GibcoTM, catalog number: 11995065 )
- Neurobasal medium:
- Trypsin (2.5%), no phenol red (Thermo Fisher Scientific, GibcoTM, catalog number: 15090046 )
- Lipofectamine 2000 (Thermo Fisher Scientific, InvitrogenTM, catalog number: 11668019 )
- Opti-MEM (Thermo Fisher Scientific, catalog number: 31985062 )
- Murine anti-APP N-terminal monoclonal (22C11) (Merck, catalog number: MAB348 )
- HEPES (1 M) (Thermo Fisher Scientific, GibcoTM, catalog number: 15630080 )
- Phosphate buffer saline (PBS) (Thermo Fisher Scientific, GibcoTM, catalog number: 10010031 )
- Paraformaldehyde (Sigma-Aldrich, catalog number: P6148 )
- Sucrose (NZYTech, catalog number: MB18601 )
- Saponin (Sigma-Aldrich, catalog number: 47036 )
- Donkey anti-mouse Alexa 488 (Thermo Fisher Scientific, catalog number: A-21202 )
- Coverslip-Slide Mounting solution (FluoroMount-G) (SouthernBiotech, catalog number: 0100-01 )
- DAPI (Sigma-Aldrich, catalog number: D9542 )
- HBSS (GE Healthcare, HycloneTM, catalog number: SH30031.03 )
- 50% glucose in sterile water (NZYTech, catalog number: MB16801 )
- Bovine serum albumin fraction V (BSA) (NZYTech, catalog number: MB04602 )
Equipment
- CO2 incubator for primary cell culture (BINDER, model: CB 160 )
- Counting chamber (Belden, Hirschmann, catalog number: 8100103 )
- Epifluorescence upright microscope Z2 (Carl Zeiss, model: Axio Imager Z2 ) equipped a 60x NA-1.4 oil immersion objective and an AxioCam MRm CCD camera (Carl Zeiss)
Software
- ImageJ software (free download from http://rsb.info.nih.gov/ij/)
- GraphPad Prism 6 (https://www.graphpad.com/scientific-software/prism/)
Procedure
Note: The APP endocytosis assay is performed on neurons cultivated for 9 days in vitro (9 DIV).
- Cell culture and transfection (Day 1, Day 6, Day 8)
- Day 1: Prepare primary cortical neurons from cortices and hippocampi from wild-type females and males mouse embryos (E16) as previously described (Almeida et al., 2005 and 2006; Ubelmann et al., 2017). Briefly, dissociated neurons are plated on 24-well plates containing poly-D-lysine coated glass coverslips (5-10 x 104 cells/cm2) in plating medium (DIV 0). After 3 h or overnight, replace the plating medium with Neurobasal medium allowing only neurons to grow and differentiate until use. (Almeida et al., 2005 and 2006; Ubelmann et al., 2017)
- Day 8: For expression of APP-RFP cDNA, 8 DIV primary cortical neurons are transiently transfected with Lipofectamine 2000 according to manufacturer’s protocol, using 0.5 µg cDNA: 0.5 µl Lipofectamine mix in 25 µl Opti-MEM per well of 24-well plate with 250 µl fresh antibiotics-free Neurobasal medium and incubated overnight until assaying APP endocytosis .
Note: Expect on average 10 transfected healthy neurons at 8 DIV at stage 5 of differentiation (Dotti et al., 1988) and the neurites should not show bead-like structures that indicate degeneration. This efficiency can be achieved using freshly prepared cDNA with a regular midi-prep kit (we use a local NZYTECH brand). - (Optional) Day 6: For knockdown analysis, primary neurons at 6 days in vitro (6 DIV) were transfected with siRNA using Lipofectamine RNAiMax according to manufacturer’s protocol, after substituting culture media with fresh antibiotics-free Neurobasal medium.
- Day 1: Prepare primary cortical neurons from cortices and hippocampi from wild-type females and males mouse embryos (E16) as previously described (Almeida et al., 2005 and 2006; Ubelmann et al., 2017). Briefly, dissociated neurons are plated on 24-well plates containing poly-D-lysine coated glass coverslips (5-10 x 104 cells/cm2) in plating medium (DIV 0). After 3 h or overnight, replace the plating medium with Neurobasal medium allowing only neurons to grow and differentiate until use. (Almeida et al., 2005 and 2006; Ubelmann et al., 2017)
- APP endocytosis assay of 9 DIV primary neurons expressing APP-RFP (Figure 1) by the following steps (Day 9)

Figure 1. Schematic of APP endocytosis monitored using anti-APP antibody incubation for 10 min in axon, dendrite and cell body. EE: Early endosome; Lys: Lysosome.- Remove cell culture media with a plastic Pasteur pipette, add 400 µl B27-free Neurobasal medium for 30 min at 37 °C in cell culture incubator. All pipetting should be done slowly against the wall of the well without touching the cells.
- Dilute 0.25 µl of anti-APP antibody (22C11; 0.25 µg/µl stock concentration) into 25 µl complete Neurobasal medium with 10 mM HEPES (0.25 µl of stock solution at 1 M, pH 7.2-7.5) per coverslip.
- Place one 25 µl droplet of diluted anti-APP antibody per glass coverslip onto Parafilm stretched on a 24 wells plate lid.
- Place the coverslips, carefully and as fast as possible, over the droplets with cells facing the antibody solution, cover the reaction in a container to avoid evaporation and incubate at 37 °C for 10 min in a cell culture incubator. This step allows for anti-APP antibody binding to cell surface APP and subsequent endocytosis.
(Optional)- To monitor endocytosed APP lysosomal degradation, further incubate cells for 60 min upon washing by dipping coverslips once for 4 s in 500 µl pre-warmed PBS.
- To detect APP at the plasma membrane, incubate cells with anti-APP antibody for only 4 min at 37 °C in a cell culture incubator. This optional step allows labelling anti-APP antibody bound to cell surface APP without significant detection of endocytosis.
- To monitor endocytosed APP lysosomal degradation, further incubate cells for 60 min upon washing by dipping coverslips once for 4 s in 500 µl pre-warmed PBS.
- Wash each coverslip by dipping once in 500 µl pre-warmed PBS for 4 s.
- Fix cells by placing coverslips back in a 24-well plate and add 500 µl 4% paraformaldehyde/4% sucrose for 20 min at room temperature (RT). Replace fixative solution with 500 µl PBS, and after 3 washes proceed for detection of anti-APP.
- Remove cell culture media with a plastic Pasteur pipette, add 400 µl B27-free Neurobasal medium for 30 min at 37 °C in cell culture incubator. All pipetting should be done slowly against the wall of the well without touching the cells.
- Anti-APP antibody detection (Day 9)
- For detection of endocytosed APP bound to anti-APP antibody, permeabilize fixed cells with 500 µl 0.1% saponin/PBS (permeabilization buffer) for 60 min at RT. Remove the permeabilization buffer and wash each coverslip with PBS.
Optional: For detection of anti-APP bound to APP at the cell surface (4 min), no permeabilization is required. Do not include saponin in steps C2 and C3. - Block non-specific binding of the antibody to serum proteins with 500 µl of blocking buffer (3% FBS/0.1% saponin/PBS) for 60 min at RT.
- Place each coverslip (cells facing down) onto a 50 µl droplet of diluted donkey anti-mouse Alexa 488 (1:250) in 3% FBS/0.1% saponin/PBS.
Optional: Include DAPI in the antibody solution (1:10,000 of a stock of 1 mg/ml) to counterstain cell nuclei. - Cover the reaction and incubate it for 60 min at RT, in the dark.
- Place coverslips back on a 24-well plate and wash 3 times with PBS at RT.
- Mount coverslips with cells facing down onto a 25 µl droplet of Fluoromount G at RT on microscope slides and let dry overnight in the dark to preserve the fluorescence signal.
Note: Pipette Fluoromount G quickly to prevent it from drying but gently to avoid bubbles.
- For detection of endocytosed APP bound to anti-APP antibody, permeabilize fixed cells with 500 µl 0.1% saponin/PBS (permeabilization buffer) for 60 min at RT. Remove the permeabilization buffer and wash each coverslip with PBS.
- Image acquisition (Day 10)
Image acquisition is done using epifluorescence microscopy (such as ZEISS Z2) with a 60x NA 1.4 oil immersion objective and a CCD camera (see Equipment for details on microscope used). See Figure 2 for a representative image.- Neuronal cell body, portions of dendrites and axons must be in focus. Not well-developed neurons or neurons expressing low or high levels of APP-RFP should be excluded. Exposure times should be determined based on the sample with the brightest expected signal, to use as much of the dynamic range of the camera (ideally use a camera with 16-bit binary range) without saturating any of the pixels.
- Acquire 10-20 neurons per condition to have sufficient data for statistical analysis. To image the whole neuron, it may be necessary to acquire multiple fields.

Figure 2. Representative image of APP endocytosis by primary neurons expressing APP-RFP incubated for 10 min with anti-APP (22C11) detected with anti-mouse Alexa 488. Dd: dendrite; Ax: axon; scale bars = 10 µm (adapted from Ubelmann et al., 2017).
- Neuronal cell body, portions of dendrites and axons must be in focus. Not well-developed neurons or neurons expressing low or high levels of APP-RFP should be excluded. Exposure times should be determined based on the sample with the brightest expected signal, to use as much of the dynamic range of the camera (ideally use a camera with 16-bit binary range) without saturating any of the pixels.
- Image analysis & APP endocytosis quantification in cell body, dendrite and axon (Figure 3) (Day 10-11)
Note: Quantification of endocytosed fluorescent signal acquired with ImageJ/Fiji.- Identify the cell body, a dendrite and the axon based on neuronal morphology and on RFP signal and roughly outline composite selections (non-contiguous ROIs) corresponding to cell body, two to five ~20 µm segments in dendrites and one in the axon (usually thinner, longer and with branches at 90° angle) using ‘polygon selection’ while pressing ‘shift’ (see Figure 3). Axons and dendrites can alternatively be counterstained with Ankyrin-G (Santa Cruz Biotechnology) or MAP2 (Sigma-Aldrich).

Figure 3. Image analysis of APP endocytosis: Step 1 - Duplicate image (shift + D) containing composite selections and remove unselected pixels by using ‘Clear outside’ (see Figure 4).

Figure 4. Image analysis of APP endocytosis: Step 2 - To refine the ROI to the cell boundary: auto ‘Threshold’ the APP-RFP signal in the cell body; create a new ROI by clicking on the thresholded neurite with the ‘Magic Wand’; add ROI to ‘ROI manager’ (shift + t) (see Figure 5).

Figure 5. Image analysis of APP endocytosis: Step 3 - To refine the ROI to the dendrites and axons: adjust the auto ‘Threshold’ to the APP-RFP signal in the dendrites and in the axon if necessary; create new ROIs by clicking on the thresholded neurites with the ‘Magic Wand’; add ROIs to ‘ROI manager’ (shift + t) (see Figure 6).

Figure 6. Image analysis of APP endocytosis: Step 4 - Transfer all regions to endocytosed Alexa488-anti-APP channel, draw a background ROI in a cell-free region using the tool ‘polygon selection’ and add it to the ‘ROI manager’. In the ROI manager, press ‘measure’ to obtain the mean Alexa488-anti-APP fluorescence intensities per ROI (background, cell body, dendrites and axon). Select, copy and export measurements to Microsoft Excel (see Figure 7).

Figure 7. Image analysis of APP endocytosis: Step 5 - Transfer all regions back to APP-RFP channel, and in the ROI manager, press ‘measure’ to obtain the mean APP-RFP fluorescence intensities per ROI (background, cell body, dendrites and axon). Select, copy and export measurements to Microsoft Excel (see Figure 8).

Figure 8. Image analysis of APP endocytosis: Step 6
- Identify the cell body, a dendrite and the axon based on neuronal morphology and on RFP signal and roughly outline composite selections (non-contiguous ROIs) corresponding to cell body, two to five ~20 µm segments in dendrites and one in the axon (usually thinner, longer and with branches at 90° angle) using ‘polygon selection’ while pressing ‘shift’ (see Figure 3). Axons and dendrites can alternatively be counterstained with Ankyrin-G (Santa Cruz Biotechnology) or MAP2 (Sigma-Aldrich).
Data analysis
Note: Data analysis is done using Microsoft Excel.
- Subtract the background mean fluorescence intensity from all measurements, per single neuron.
- Normalize the Alexa488-anti-APP fluorescence intensities (from the 3 neuronal compartments) by the APP-RFP fluorescence intensity in the cell body to control for the different expression level of APP-RFP, per single neuron.
- Different conditions can be compared using a classical t-test provided the data follow a normal distribution.
- The sample size is about 20 cells per condition in each independent experiment, based on previous studies. Statistical significance for at least three independent experiments is determined on normal data (D’Agostino-Pearson omnibus normality test) by two-tailed Student’s t-test and for multiple comparisons one-way ANOVA with Tukey’s test using GraphPad Prism 6.
- Statistical significance for nonparametric data was tested by Mann-Whitney test.
Notes
- Variability is often due to the culture of primary neurons; the level of differentiation should be kept constant between independent experiments.
- In our hands, APP endocytosis is robust with the average results very reproducible. However, it is expected variability between different neurons in the same experiment.
- For optimal APP transfection and thus experimental conditions, good quality DNA plasmid and healthy and well developed primary neurons are paramount.
Acknowledgments
We thank for the gift of APP-RFP plasmids from Dr. S. Kins (University of Kaiserslautern). This work has been supported by CEDOC and by a Marie Curie Integration Grant (334366 TrafficInAD FP7-PEOPLE-2012-CIG; Marie Curie Actions, EC). iNOVA4Health–UID/Multi/04462/2013, a program financially supported by Fundação para a Ciência e Tecnologia (FCT)/Ministério da Educação e Ciência, through national funds and co-funded by FEDER under the PT2020 Partnership Agreement is acknowledged. CGA is Investigator FCT (IF/00998/2012, FCT). FU was recipient of an FRM postdoctoral fellowship (SPE20130326599) and an FCT post-doctoral fellowship (SFRH/BPD/94186/2013). This protocol was adapted from our recent publication in EMBO reports (Ubelmann et al., 2017). The authors declare no conflict of interest.
References
- Almeida, C. G., Takahashi, R. H. and Gouras, G. K. (2006). β-amyloid accumulation impairs multivesicular body sorting by inhibiting the ubiquitin-proteasome system. J Neurosci 26(16): 4277-4288.
- Almeida, C. G., Tampellini, D., Takahashi, R. H., Greengard, P., Lin, M. T., Snyder, E. M. and Gouras, G. K. (2005). β-amyloid accumulation in APP mutant neurons reduces PSD-95 and GluR1 in synapses. Neurobiol Dis 20(2): 187-198.
- Cirrito, J. R., Kang, J. E., Lee, J., Stewart, F. R., Verges, D. K., Silverio, L. M., Bu, G., Mennerick, S. and Holtzman, D. M. (2008). Endocytosis is required for synaptic activity-dependent release of amyloid-beta in vivo. Neuron 58(1): 42-51.
- Dotti, C. G., Sullivan, C. A. and Banker, G. A. (1988). The establishment of polarity by hippocampal neurons in culture. J Neurosci 8: 1454-1468.
- Edgar, J. R., Willen, K., Gouras, G. K. and Futter, C. E. (2015). ESCRTs regulate amyloid precursor protein sorting in multivesicular bodies and intracellular amyloid-beta accumulation. J Cell Sci 128(14): 2520-2528.
- Grbovic, O. M., Mathews, P. M., Jiang, Y., Schmidt, S. D., Dinakar, R., Summers-Terio, N. B., Ceresa, B. P., Nixon, R. A. and Cataldo, A. M. (2003). Rab5-stimulated up-regulation of the endocytic pathway increases intracellular β-cleaved amyloid precursor protein carboxyl-terminal fragment levels and Aβ production. J Biol Chem 278(33): 31261-31268.
- Haass, C., Koo, E. H., Mellon, A., Hung, A. Y. and Selkoe, D. J. (1992). Targeting of cell-surface beta-amyloid precursor protein to lysosomes: alternative processing into amyloid-bearing fragments. Nature 357(6378): 500-503.
- Koo, E. H. and Squazzo, S. L. (1994). Evidence that production and release of amyloid β-protein involves the endocytic pathway. J Biol Chem 269(26): 17386-17389.
- Lee, J., Retamal, C., Cuitino, L., Caruano-Yzermans, A., Shin, J. E., van Kerkhof, P., Marzolo, M. P. and Bu, G. (2008). Adaptor protein sorting nexin 17 regulates amyloid precursor protein trafficking and processing in the early endosomes. J Biol Chem 283(17): 11501-11508.
- Morel, E., Chamoun, Z., Lasiecka, Z. M., Chan, R. B., Williamson, R. L., Vetanovetz, C., Dall'Armi, C., Simoes, S., Point Du Jour, K. S., McCabe, B. D., Small, S. A. and Di Paolo, G. (2013). Phosphatidylinositol-3-phosphate regulates sorting and processing of amyloid precursor protein through the endosomal system. Nat Commun 4: 2250.
- Mucke, L. and Selkoe, D. J. (2012). Neurotoxicity of amyloid β-protein: synaptic and network dysfunction. Cold Spring Harb Perspect Med 2(7): a006338.
- Rajendran, L. and Annaert, W. (2012). Membrane trafficking pathways in Alzheimer's disease. Traffic 13(6): 759-770.
- Rajendran, L., Schneider, A., Schlechtingen, G., Weidlich, S., Ries, J., Braxmeier, T., Schwille, P., Schulz, J. B., Schroeder, C., Simons, M., Jennings, G., Knolker, H. J. and Simons, K. (2008). Efficient inhibition of the Alzheimer's disease β-secretase by membrane targeting. Science 320(5875): 520-523.
- Sannerud, R., Declerck, I., Peric, A., Raemaekers, T., Menendez, G., Zhou, L., Veerle, B., Coen, K., Munck, S., De Strooper, B., Schiavo, G. and Annaert, W. (2011). ADP ribosylation factor 6 (ARF6) controls amyloid precursor protein (APP) processing by mediating the endosomal sorting of BACE1. Proc Natl Acad Sci U S A 108(34): E559-568.
- Sullivan, S. E., Dillon, G. M., Sullivan, J. M. and Ho, A. (2014). Mint proteins are required for synaptic activity-dependent amyloid precursor protein (APP) trafficking and amyloid β generation. J Biol Chem 289(22): 15374-15383.
- Szodorai, A., Kuan, Y. H., Hunzelmann, S., Engel, U., Sakane, A., Sasaki, T., Takai, Y., Kirsch, J., Muller, U., Beyreuther, K., Brady, S., Morfini, G. and Kins, S. (2009). APP anterograde transport requires Rab3A GTPase activity for assembly of the transport vesicle. J Neurosci 29(46): 14534-14544.
- Ubelmann, F., Burrinha, T., Salavessa, L., Gomes, R., Ferreira, C., Moreno, N. and Guimas Almeida, C. (2017). Bin1 and CD2AP polarise the endocytic generation of beta-amyloid. EMBO Rep 18(1): 102-122.
- Xiao, Q., Gil, S. C., Yan, P., Wang, Y., Han, S., Gonzales, E., Perez, R., Cirrito, J. R. and Lee, J. M. (2012). Role of phosphatidylinositol clathrin assembly lymphoid-myeloid leukemia (PICALM) in intracellular amyloid precursor protein (APP) processing and amyloid plaque pathogenesis. J Biol Chem 287(25): 21279-21289.
- Yamazaki, T., Koo, E. H. and Selkoe, D. J. (1996). Trafficking of cell-surface amyloid beta-protein precursor. II. Endocytosis, recycling and lysosomal targeting detected by immunolocalization. J Cell Sci 109 (Pt 5): 999-1008.
- Yamazaki, T., Selkoe, D. J. and Koo, E. H. (1995). Trafficking of cell surface beta-amyloid precursor protein: retrograde and transcytotic transport in cultured neurons. J Cell Biol 129(2): 431-442.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Ubelmann, F., Burrinha, T. and Guimas Almeida, C. (2017). A Novel Protocol to Quantitatively Measure the Endocytic Trafficking of Amyloid Precursor Protein (APP) in Polarized Primary Neurons with Sub-cellular Resolution. Bio-protocol 7(23): e2629. DOI: 10.21769/BioProtoc.2629.
Category
Neuroscience > Neuroanatomy and circuitry > Immunofluorescence
Cell Biology > Cell imaging > Fluorescence
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










