- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Bioluminescence Monitoring of Neuronal Activity in Freely Moving Zebrafish Larvae
Published: Vol 7, Iss 18, Sep 20, 2017 DOI: 10.21769/BioProtoc.2550 Views: 10308
Reviewed by: Geoff LauAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
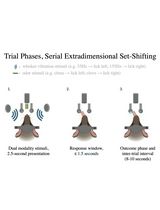
Training Mice to Perform Attentional Set-Shifting Under Head Restraint
Katarina Kalajzic [...] Timothy Spellman
Sep 5, 2025 1470 Views
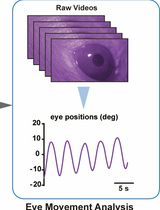
Mouse Vestibulo-Ocular Reflex Testing for Otolith Organs and Horizontal Semicircular Canal
Tong Zhao [...] Fangyi Chen
Nov 20, 2025 1199 Views

Station Holding During Rheotaxis: A Sensitive Assay of Lateral Line Function in Larval Zebrafish
Sophie Cohen-Bodénès [...] Lavinia Sheets
Dec 20, 2025 762 Views
Abstract
The proof of concept for bioluminescence monitoring of neural activity in zebrafish with the genetically encoded calcium indicator GFP-aequorin has been previously described (Naumann et al., 2010) but challenges remain. First, bioluminescence signals originating from a single muscle fiber can constitute a major pitfall. Second, bioluminescence signals emanating from neurons only are very small. To improve signals while verifying specificity, we provide an optimized 4 steps protocol achieving: 1) selective expression of a zebrafish codon-optimized GFP-aequorin, 2) efficient soaking of larvae in GFP-aequorin substrate coelenterazine, 3) bioluminescence monitoring of neural activity from motor neurons in free-tailed moving animals performing acoustic escapes and 4) verification of the absence of muscle expression using immunohistochemistry.
Keywords: GFP-Aequorin-optBackground
Unlike fluorescent genetically encoded calcium indicators (GECIs) (Grienberger and Konnerth, 2012), such as the GCaMP family, the bioluminescent indicator GFP-aequorin (Shimomura et al., 1962) does not require light excitation and therefore opens new avenues for monitoring neural activity in moving animals, including flies (Martin et al., 2007), mice (Rogers et al., 2007) and zebrafish larvae (Naumann et al., 2010). However, efficient use of GFP-aequorin remains challenging to achieve in zebrafish larvae, restricting its widespread use as a calcium indicator. The limitation lies in the fact that bioluminescence signals originating from a single muscle fiber are so large they constitute a major pitfall. Once absence of muscle expression is verified for a given transgenic line, bioluminescence signals emanating from neurons only are very small. To overcome these limitations, we developed a codon-optimized variant of GFP-aequorin for zebrafish larvae, achieved selective expression in motor and sensory neurons using existing transgenic lines, modified coelenterazine soaking protocol in order to conduct experiments at 4 days post-fertilization, created a behavioral bioluminescence assay for monitoring neuronal activity during acoustic evoked stereotyped escape responses in zebrafish larvae.
Materials and Reagents
- Microscope slide 76 x 26 x 1.1 mm (VINCENT LEERMIDDELEN SCIENTIFIC, catalog number: 29201316 )
- Cover glass Knittel Glass 24 x 60 mm
- Zebrafish larvae Adult AB and and Tüpfel long fin (TL) strains of Danio rerio aged between 0 and 4 dpf (day post fertilization) were used for this study; transgenic lines used in this protocol (available on request): Tg(mnx1:gal4)icm11;Tg(UAS:GFP-aequorin-opt)icm09
- Mammalian GFP-aequorin sequence (provided by Dr. Ludovic Tricoire, Université Pierre et Marie Curie, Paris, France, see text file in Supplement file 1)
- PT2 14xUAS plasmid (provided by Pr. Koichi Kawakami, National Institute of Genetics, Mishima, Japan, map can be downloaded at http://kawakami.lab.nig.ac.jp/trans.html)
- Instant Ocean® salts
- Methylene blue
- Fixation solution (4% PFA) paraformaldehyde, powder 95% (Sigma-Aldrich, catalog number: 158127 )
- NGS (Abcam, catalog number: ab7481 )
- DMSO (Sigma-Aldrich, catalog number: D8418 )
- Triton X-100 (Sigma-Aldrich, catalog number: T9284 )
- Phosphate-buffered saline (PBS) (Thermo Fisher Scientific, GibcoTM, catalog number: 18912014 )
- Chicken anti-GFP primary antibody (Abcam, catalog number: ab13970 )
- Alexa Fluor 488 goat anti-chicken IgG (Thermo Fisher Scientific, InvitrogenTM, catalog number: A-11039 )
- Mounting Medium for fluorescence (Vector Laboratories, catalog number: H-1000 )
- Coelenterazine-h (Biotium, catalog number: 10111 )
- Propylene glycol
- Cyclodextrine
- Agarose
- ‘Blue water’ (see Recipes)
- Blocking solution (see Recipes)
- Incubation solution (see Recipes)
- Washing solution (see Recipes)
Equipment
- Wave generator (Agilent Technologies, catalog number: 33210A )
- Audio amplifier (Lepai, catalog number: LP-2020A )
- Upright confocal microscope (Olympus, model: FV1000 )
- 850 nm LED (Effisharp, Effilux, France, EFFI-sharp_CM_850_2)
- Long-pass 780 filter (Asahi Spectra USA, catalog number: ZIL0780 )
- Long-pass 810 filter (Asahi Spectra USA, catalog number: XIL0810 )
- Diffuser (Thorlabs, catalog number: DG10-120-B )
- High-speed infrared sensitive camera (Mikrotron, catalog number: Eosens MC1362 )
- Objective Nikkor 50 mm f/1.8D (Nikon, Japan)
- Photomultiplier tube (PMT) (Hamamatsu Photonics, catalog number: H7360-02 )
- Acquisition card (National Instruments, catalog number: PCI 6602 )
- Band-pass filter (525 nm/50 nm) (ZEISS, catalog number: 489038-8002-000 )
- Short-pass filter (670 nm) (Asahi Spectra USA, catalog number: XVS0670 )
- 2-Ohm speaker
- TTL chronogram generator (RD Vision, France, EG Chrono)
- Black cardboard box protecting the setup from residual photons in the room
- Standard equipment for molecular biology
- Sonicator (Emerson Electric, Branson, model: B1510-DTH )
Software
- Hiris video software (RD Vision, France)
- MATLAB (The MathWorks, catalog number: R2012b)
Procedure
- Generation of the codon-optimized Tg(UAS:GFP-aequorin-opt) transgenic line
- Generate a codon-optimized sequence, GFP-aequorin-opt, for expression in zebrafish from original sequence of mammalian GFP-aequorin (we used to online tool freely available made by Integrated DNA Technology: https://eu.idtdna.com/CodonOpt, original and optimized DNA sequences provided in Supplement file 2).
- Subclone GFP-aequorin-opt into a PT2 14xUAS plasmid.
- Inject the UAS:GFP-aequorin-opt construct in the Tg(mnx1:gal4)icm11 embryos to generate the Tg(mnx1:gal4;UAS:GFP-aequorin-opt)icm09 double transgenic line. Injection mix is composed as follows: 2 µl ADN (120 ng/µl) + 2 µl ARN transposase (175 ng/µl) + 1 µl KCl 2 M +1 µl 2% phenol red, complete to 10 µl with MilliQ H2O.
- Maintain adult AB and Tüpfel long fin (TL) strains of Danio rerio on a 14/10 h light cycle and water is maintained at 28.5 °C, conductivity at 500 μS and pH at 7.4.
- Raise embryos in ‘blue water’ (3 g of Instant Ocean® salts and 2 ml of methylene blue at 1% in 10 L of osmosed water, see Recipes) at 28.5 °C during the first 24 h before screening for GFP expression.
- Characterization of GFP-aequorin expression with immunohistochemistry
- Fix 4 dpf larvae in 4% PFA for 4 h at 4 °C followed by 3 x 5 min washes in PBS.
- Block larvae for 1 h in blocking solution (see Recipes) (agitation required).
- Incubate larvae with the primary antibody (anti-GFP, dilution 1:500) over night at 4 °C in incubation solution (see Recipes) (agitation required).
- Wash three times for 5 min in washing solution (see Recipes), then incubate larvae in the dark with the secondary antibody (Alexa Fluor 488 goat anti-chicken IgG, dilution1:1,000) in PBST (agitation required) for 2 h at RT.
- Wash three times for 5 min in PBST, then mount larvae on a slide with mounting medium and image on a standard upright confocal microscope (Olympus FV-1000).
- Perform negative IHC controls by omitting the primary antibody.
- Image the entire immunostained Tg(mnx1:gal4;UAS:GFP-aequorin-opt)icm09 larvae to confirm selective expression of GFP-aequorin-opt in spinal motor neuron populations and absence from muscle fibers. We noted more prominently primary dorsal motor neurons but also intermediate and ventral secondary motor neurons (Figure 1) without any expression in the muscles and only very limited expression in the brain and hindbrain.

Figure 1. Expression pattern of GFP-aequorin-opt in motor neurons. Fluorescent image (upper panel) and immunohistochemistry for GFP (lower panel) in a 4 dpf Tg(mnx1:gal4;UAS:GFP-aequorin-opt) double transgenic zebrafish larva showing selective expression in spinal motor neurons (arrowhead: dorsal primary, arrow: ventral secondary motor neurons), and strictly no expression in muscle fibers (white arrow in the upper panel).
- Fix 4 dpf larvae in 4% PFA for 4 h at 4 °C followed by 3 x 5 min washes in PBS.
- Soaking of larvae in coelenterazine solution
- Prepare 10 mM stock solution from lyophilized coelenterazine-h: e.g.,
- For 250 µg of coelenterazine-h, final volume is 60 µl (M = 407.5 g/M).
- Add propylene glycol to (25% of final volume, e.g., 15.4 µl).
- Sonicate cyclodextrine at 45% (4.5 g in 10 ml).
- Add cyclodextrine (75% of final volume, e.g., 46 µl).
- Prepare 60 µM coelenterazine-h soaking solution from stock:
Dilute stock solution in ‘blue water’ (e.g., 6 µl in 1 ml for 10 embryos).
- Dechorionate embryos at 1 day post-fertilization under optical magnification.
- Soak dechorionated embryos overnight at 26 °C (e.g., 100 µl for each embryo, use 48-well plate sealed with paraffin).
- Renew 60 µM soaking solution at 2 days post-fertilization. Embryos are maintained in the dark.
- Perform behavioral experiments at 4 days post-fertilization (total soaking time is 72 h).
- Monitoring neuronal activity with bioluminescence
- Build a lightproof setup for bioluminescence assay (Figure 2)

Figure 2. Bioluminescence setup for escapes. A. Signals emitted from spinal motor neurons in Tg(mnx1:gal4;UAS:GFP-aequorin-opt) double transgenic zebrafish larvae at 4 dpf were recorded using a photomultiplier tube under infrared illumination during active behaviors elicited by an acoustic stimulus. B. Picture of the setup showing every component: 1: high-speed camera; 2: 50 mm objective; 3: IR 850 nm LED; 4: Diffuser; 5: Long-pass filters; 6: photomultiplier tube.
- Using black boards, create a 1 m square lightproof box.
- Infrared light illumination is provided by an 850 nm LED mounted with 2 long-pass 780 and 810 filters and a diffuser.
- Video acquisition is performed at 1,000 Hz using a high-speed infrared sensitive camera at 320 x 320 pixels resolution controlled by the video software (Hiris®).
- Photons are counted with a photomultiplier tube located under the larva arena and sent to an acquisition card. A band-pass filter (525 nm/50 nm) and a short-pass filter (670 nm) are placed between the larva and the PMT.
- A custom application-programming interface synchronizes the video acquisition with the photon count and the stimulus delivery using a 30 trials batched TTL chronogram.
- Using black boards, create a 1 m square lightproof box.
- Run the bioluminescence assay one larva at a time
- Place larva in a circular (2 cm diameter) 3D-printed arena (larva can also be head-embedded in 1.5% low-melting point agarose with the tail free to move).
- Place the larva in the arena and attach the arena to a small 2-Ohm speaker.
- Deliver sinusoidal stimuli (5 cycles, 500 Hz) produced by the waveform generator and audio amplifier through a 2-Ohm speaker attached to the larva arena.
- Adjust intensity to the lowest value reliably eliciting an escape response (between 0.5 and 5 V usually).
- Each trial consists in a 500 msec baseline followed by a 10 msec acoustic stimulus and 1,990 msec subsequent recording.
- Assays consist of 30 trials with 1-min inter-trial intervals to reduce habituatio.
- Place larva in a circular (2 cm diameter) 3D-printed arena (larva can also be head-embedded in 1.5% low-melting point agarose with the tail free to move).
- Build a lightproof setup for bioluminescence assay (Figure 2)
Data analysis
- Kinematics analysis (blue trace in Figure 3, right panel)
- Using a custom MATLAB algorithm (see Supplement file 3), manually locate the base and tip of the tail. The tail is subsequently automatically tracked.
- The tail angle is computed for each frame and filtered using median filtering (window size = 10).
- The start of the movement is determined as the first frame followed by 3 with a differential tail angle value above 0.08 degree in our conditions.
- The end of the movement is determined as the last of 20 frames with a differential tail angle value below 0.1 degree in our conditions.
- Local minimal and maximal values of the tail angle occur at least 2 msec apart and 1° above the 5 msec preceding value.
- Automated movement categorization is determined as follows: ‘escapes’ for all movements with maximum values of tail angle > 45° and number of cycles > 1; ‘slow swims’ for all movement with maximum values of tail angle < 25° and number of cycles > 1.
- Bioluminescence recording and analysis (green trace in Figure 3B)
- Photons are counted at 1 kHz (temporal resolution of 1 msec) and then binned every 10 msec.
- The signal is filtered using a running average with a window size of 10, giving a typical signal-to-noise ratio (SNR) for active movements of 50 to 1.
- Noise is extrapolated from a linear fit of the cumulative photon count before the stimulus and subtracted from the signal.
- The start and end of the bioluminescent signal are computed respectively as the first time point followed by 3 points with a value above 0.4 photons/10 msec and below 0.2 photons/10 msec from this first point bioluminescence value.
- The time-to-peak is calculated between the start and the peak of the bioluminescent signal while the decay coefficient is derived from the one-term exponential fit between the peak and the end of the signal.

Figure 3. Example of kinematics and bioluminescence data for an escape response. A. 4 dpf larva in the head-embedded setup with the tail tracked (red dots) in transmitted IR light. B. Superimposed, and color-coded according to delay from stimulus, behaviors elicited by an acoustic stimulus; C. Example traces of typical bioluminescence signals and tail angle observed for an escape response; D. Example of a color-coded slow swim and (E) corresponding bioluminescence signal and tail angle traces for comparison with escapes.
Notes
- Soaking in coelenterazine is key to obtaining a good signal-to-noise ratio for bioluminescence: pay attention to soaking conditions (incubation temperature, hour of coelenterazine renewal, keep the wells of the plate air-tight).
- Noise due to ambient light must be tested before running the behavioral experiment; make sure the box is completely lightproof.
- The Tg(UAS:GFP-aequorin-opt) zebrafish line is freely available to all users that request it.
Recipes
- ‘Blue water’
3 g of Instant Ocean® salts and 2 ml of 1% methylene blue in 10 L of ultrapure water
- Blocking solution
10% NGS
1% DMSO
0.5% Triton X-100
in 0.1 M PBS
- Incubation solution
1% NGS
1% DMSO
0.5% Triton X-100
in 0.1% PBS
- Washing solution
0.1 M PBST with 0.5% Triton X-100
Acknowledgments
We would like to thank Dr. Ludovic Tricoire (University Pierre et Marie Curie, France) for providing the original GFP-aequorin sequence, Dr. Tom Auer and Dr. Filippo Del Bene (Institut Curie, France) for providing the mnx1 construct and Prof. Koichi Kawakami (NIG, Japan) for providing the PT2 14xUAS plasmid. We are indebted to RD Vision for developing the custom API to synchronize photon collection and video acquisition. We thank Bogdan Buzurin and Natalia Maties for fish care. SK received a PhD fellowship from Inserm and Assistance Publique–Hôpitaux de Paris. This work received financial support from the Institut du Cerveau et de la Moelle épinière with the French program ‘Investissements d’avenir’ ANR-10-IAIHU-06, the ENP Chair d’excellence, the Fondation Bettencourt Schueller (FBS), the City of Paris Emergence program, the ATIP/Avenir junior program from INSERM and CNRS, the NIH Brain Initiative grant no. 5U01NS090501 and the European Research Council (ERC) starter grant ‘OptoLoco’ #311673.
References
- Grienberger, C. and Konnerth, A. (2012). Imaging calcium in neurons. Neuron 73(5): 862-885.
- Martin, J. R., Rogers, K. L., Chagneau, C. and Brulet, P. (2007). In vivo bioluminescence imaging of Ca2+ signalling in the brain of Drosophila. PLoS One 2(3): e275.
- Naumann, E. A., Kampff, A. R., Prober, D. A., Schier, A. F. and Engert, F. (2010). Monitoring neural activity with bioluminescence during natural behavior. Nat Neurosci 13(4): 513-520.
- Rogers, K. L., Picaud, S., Roncali, E., Boisgard, R., Colasante, C., Stinnakre, J., Tavitian, B. and Brulet, P. (2007). Non-invasive in vivo imaging of calcium signaling in mice. PLoS One 2(10): e974.
- Shimomura, O., Johnson, F. H. and Saiga, Y. (1962). Extraction, purification and properties of aequorin, a bioluminescent protein from the luminous hydromedusan, Aequorea. J Cell Comp Physiol 59: 223-239.
Article Information
Copyright
Knafo et al. This article is distributed under the terms of the Creative Commons Attribution License (CC BY 4.0).
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Knafo, S., Prendergast, A., Thouvenin, O., Figueiredo, S. N. and Wyart, C. (2017). Bioluminescence Monitoring of Neuronal Activity in Freely Moving Zebrafish Larvae. Bio-protocol 7(18): e2550. DOI: 10.21769/BioProtoc.2550.
- Knafo, S., Fidelin, K., Prendergast, A., Tseng, P. B., Parrin, A., Dickey, C., Bohm, U. L., Figueiredo, S. N., Thouvenin, O., Pascal-Moussellard, H. and Wyart, C. (2017). Mechanosensory neurons control the timing of spinal microcircuit selection during locomotion. Elife 6.
Category
Neuroscience > Behavioral neuroscience > Sensorimotor response
Neuroscience > Neuroanatomy and circuitry > Live-cell imaging
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









