- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Phototaxis Assay for Chlamydomonas reinhardtii
Published: Vol 7, Iss 12, Jun 20, 2017 DOI: 10.21769/BioProtoc.2356 Views: 15301
Reviewed by: Maria SinetovaAgnieszka ZienkiewiczAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Live-Cell Monitoring of Piecemeal Chloroplast Autophagy
Masanori Izumi [...] Shinya Hagihara
Nov 5, 2025 1688 Views
Chloroplast Movement Imaging Under Different Light Regimes With a Hyperspectral Camera
Paweł Hermanowicz [...] Justyna Łabuz
Dec 20, 2025 749 Views
A Simple Protocol for Periodic Live Cell Observation of Flagellate Stages in the Lichen Alga Trebouxia
Enrico Boccato [...] Mauro Tretiach
Jan 20, 2026 179 Views
Abstract
Phototaxis is a behavior in which organisms move toward or away from the light source (positive or negative phototaxis, respectively). It is crucial for phototrophic microorganisms to inhabit under proper light conditions for phototaxis. The unicellular green alga Chlamydomonas reinhardtii rapidly changes its swimming direction upon light illumination, and thus is a nice model organism for phototaxis research. Here we show two methods to assay Chlamydomonas phototaxis; one is a quick, easy and qualitative analysis, so-called the dish assay; and the other is a quantitative single-cell analysis.
Keywords: PhototaxisBackground
The unicellular green alga Chlamydomonas reinhardtii is used as a model organism in various research fields including phototaxis of microorganisms, photosynthesis, and ciliary/flagellar motility (Hegemann and Berthold, 2009). A Chlamydomonas cell perceives light at its eyespot, the photoreceptive organelle observed as an orange spot located near the cell equator. The eyespot contains the photoreceptor proteins channelrhodopsins localized in the cellular membrane and the carotenoid-rich granule layers right behind the channelrhodopsins which function as a light reflector. Because of their relative position, the eyespot undergoes highly directional photoreception, and the cell can accurately detect the direction of light illumination (Foster and Smyth, 1980; Ueki et al., 2016). Upon photoreception, two flagella change their beating balance, and the cell changes its swimming direction either toward or away from the light source.
The Chlamydomonas phototactic direction (or ‘sign’) is regulated by cellular reduction-oxidation state, which is affected by cellular metabolism such as photosynthetic and respiratory activities (Wakabayashi et al., 2011). The phototactic sign thus indirectly reflects those activities in vivo. For instance, a mutant showing fast phototactic response has been shown to have high photosynthetic activity (Kim et al., 2016). In addition, for the regulation of flagellar beating for phototactic turning of the cell, flagellar dyneins should be strictly regulated (Kamiya and Witman, 1984; Okita et al., 2005; Hegemann and Berthold, 2009). Therefore, phototaxis assay contributes to a wide variety of biological researches, such as photoreception, photosynthesis, respiration, and motor proteins.
Various methods have been developed to quantify Chlamydomonas phototaxis. Mergenhagen developed an automatic assay system for phototaxis (photoaccumulation), which detects the density of cells in the light path by a photocell (Mergenhagen, 1984). Takahashi et al. developed a computer-assisted system that automatically detects the direction of cellular movement using an infrared-sensitive video camera (Takahashi et al., 1991). Comparing to those sophisticated systems with hand-made equipment, our protocol is rather simple, and can be carried out with equipment that is commercially or freely available.
Materials and Reagents
- 50 ml tube (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 339652 )
- 4 cm Petri dish (AS ONE, catalog number: 1-8549-01 )
- Chlamydomonas strain of interest
- Tris-acetate-phosphate medium
- (Optional) Tertiary-butyl hydroperoxide (t-BOOH: final concentration, 0.2 mM) (WAKO Pure Chemical Industries, catalog number: 026-13451 )
- (Optional) N,N’-dimethylthiourea (DMTU: final concentration, 75 mM) (Sigma-Aldrich, catalog number: D188700 )
- HEPES (pH 7.4) (NACALAI TESQUE, catalog number: 17514-15 )
- EGTA (DOJINDO, catalog number: 346-01312 )
- Potassium chloride (KCl) (NACALAI TESQUE, catalog number: 28514-75 )
- Calcium chloride dihydrate (CaCl2·2H2O) (NACALAI TESQUE, catalog number: 06731-05 )
- Phototaxis assay solution (Okita et al., 2005) (see Recipes)
Equipment
- Centrifuge (Hitachi Koki, model: CR20GIII )
- Swing rotor (Hitachi Koki, model: R4SS )
- Green light-emitting diode (LED) (λ = 525 nm) (OptoSupply, model: OSPG5111A-VW )
- Red light (or white light with a red filter [λ > 600 nm])
- White sheet of paper/plastic
- Dark room or dark box
- Digital still camera (SONY, model: RX100II )
- Imaging table with camera mount (AS ONE, model: NS-CPS360N )
- Inverted microscope equipped with video camera (Olympus, model: IX70 ; Wraymer, model: 1129HMN1/3)
- Red filter (630 nm long-pass filter) (SCHOTT, model: RG630 )
- (Optional) Photometer (Apogee Instrument, model: MQ-200 )
- (Optional) Neutral density filters (HOYA, models: ND10AH and ND30AH )
Software
- Image Hyper (Science Eye, Japan) or any particle-tracking software (e.g., ImageJ with MTrack2 plugin)
- Microsoft Excel or any spreadsheet software
Procedure
- Culture cells in Tris-acetate-phosphate medium (Gorman and Levine, 1965) with aeration at 22 °C under a 12 h/12 h light/dark cycle (light: ~30 µmol photons m-2 sec-1 white light) to the mid-log phase (~3 x 106 cells/ml).
Note: Change the medium and the other culture conditions when necessary. - Harvest cells by centrifugation at 600 x g for 5 min at room temperature.
- Suspend cells in a phototaxis assay buffer at ~1 x 107 cells/ml (for dish assay) or ~1 x 106 cells/ml for quantitative assay in a 50 ml tube.
- Place the cells under red light (~40 µmol photons m-2 sec-1) for 30-60 min.
Note: This step makes cells swim actively (Sineshchekov et al., 2000). In addition, red light (λ) does not stimulate channelrhodopsin 1 (ChR1), the main photoreceptor protein for phototaxis, and thus makes cells sensitive to following green-light illumination (Berthold et al., 2008). - (For dish assay) Put 2 ml cell suspensions in a Petri dish, place it on a white sheet and take a picture before illumination (Figure 1A).
Note: When control data for positive or negative phototaxis are necessary, add final 0.2 mM t-BOOH or 75 mM DMTU to the cell suspensions, respectively. These reagents can be added either to the harvested cell suspensions in a test tube or directly the cell suspensions in a dish. t-BOOH is a kind of reactive oxygen species (ROS) which has similar effects to H2O2, and can be substituted with 25 μM H2O2. DMTU is a H2O2 scavenger, and can be substituted with 50 mM 4-hydroxy-2,2,6,6-tetramethylpiperidine 1-oxyl (TEMPOL) (Wakabayashi et al., 2011). Another option to derive negative phototaxis control is to use the strain CC-124 (agg1). It is regarded as a ‘wild type’, but its mutation in the agg1 locus causes strong negative phototaxis (Ide et al., 2016).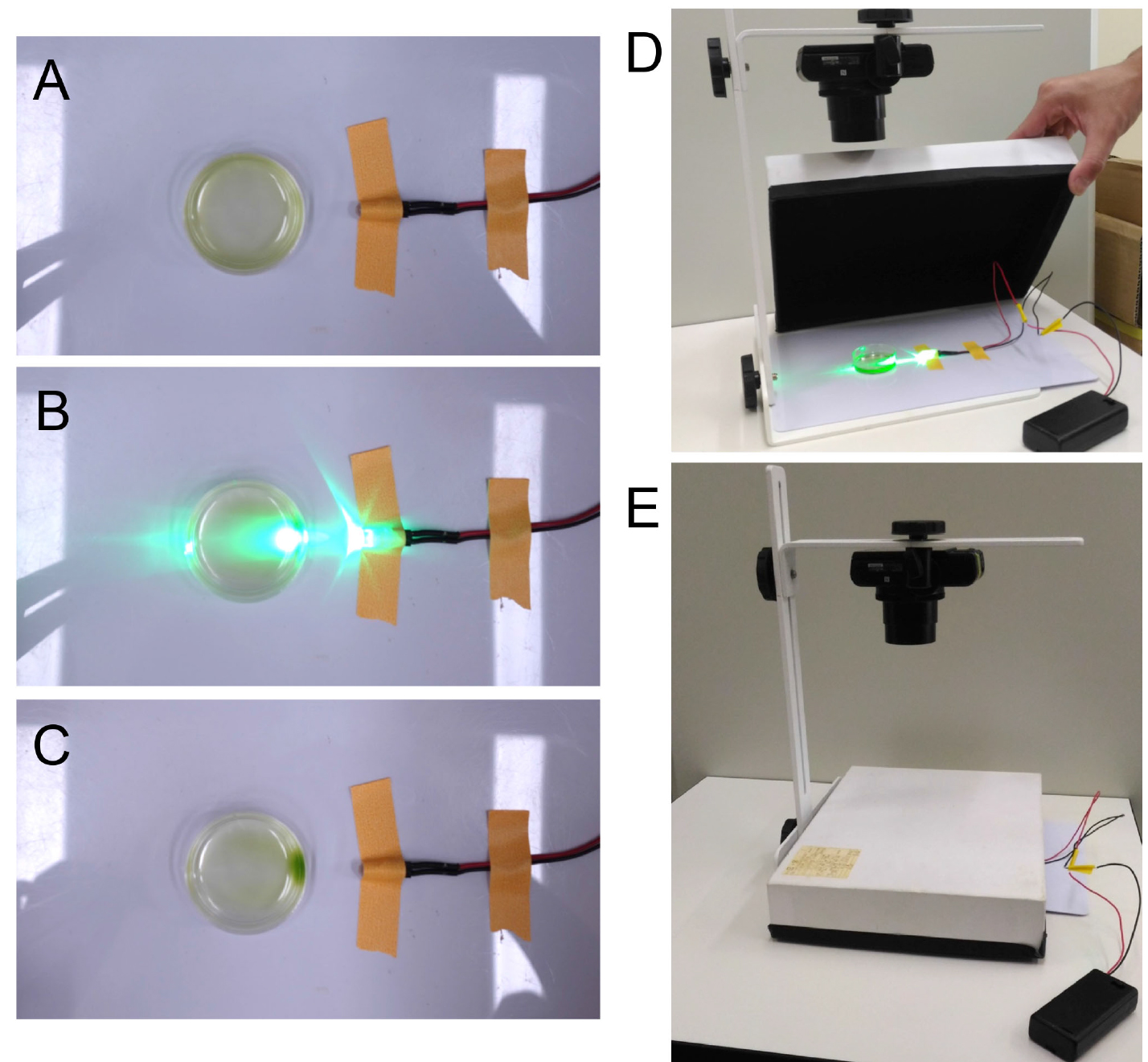
Figure 1. Dish phototaxis assay. A. A 4 cm Petri dish is set on a white plastic sheet. A green LED is set on the side of the dish and fixed. B. During and C. after illumination for 5 min. Positive phototaxis is observed. D. For dish assay, the Petri dish and the LED is covered with a box to block the room light. The inside of the box is masked with pieces of black cloth for antireflection. E. During the dish assay. Before and after 5 min (or longer) illumination, pictures are taken from the top with a camera. - Illuminate the dish from one side with green LED and cover the dish and the LED with a box (Figures 1B, 1D and 1E). Leave them for 5 min (or longer). Check the light intensity by a photometer. Typically, wild-type cells show positive phototaxis at low intensity (< 1 μmol photons m-2 sec-1) and negative phototaxis at higher intensity (> 5 μmol photons m-2 sec-1). To change the light intensity, either change the distance from the LED to the dish and/or set ND filters in front of the LED.
Note: The action spectra of phototaxis peak at 495 to 505 nm (blue-green light) (Foster et al., 1984). We use green light because blue light activates photosynthesis, which changes cellular redox state as well as the phototactic sign (Takahashi and Watanabe, 1993; Wakabayashi et al., 2011). - Take pictures (Figure 1C).
Note: Please note that this ‘dish assay’ is not an accurate method for phototaxis assay, because accumulation of cells caused by light illumination (called photo-accumulation) could occur also by the photoshock response, in that cells either stop or swim backward for a short period upon sensing the sudden change in the light intensity. The purpose of this method is a quick test for phototactic capability. Phototactic movement is defined as the movement along the light beam, which should be examined by the following single-cell analysis. - (For the single-cell analysis) Place a Petri dish on the stage of an inverted microscope and illuminate it with a green LED from the side (Figure 2). Observe the cells in the area near the light source (to estimate the light intensity when necessary) with dim red light (λ > 600 nm, ~5 μmol photons m-2 sec-1) and video-record using a CCD camera.
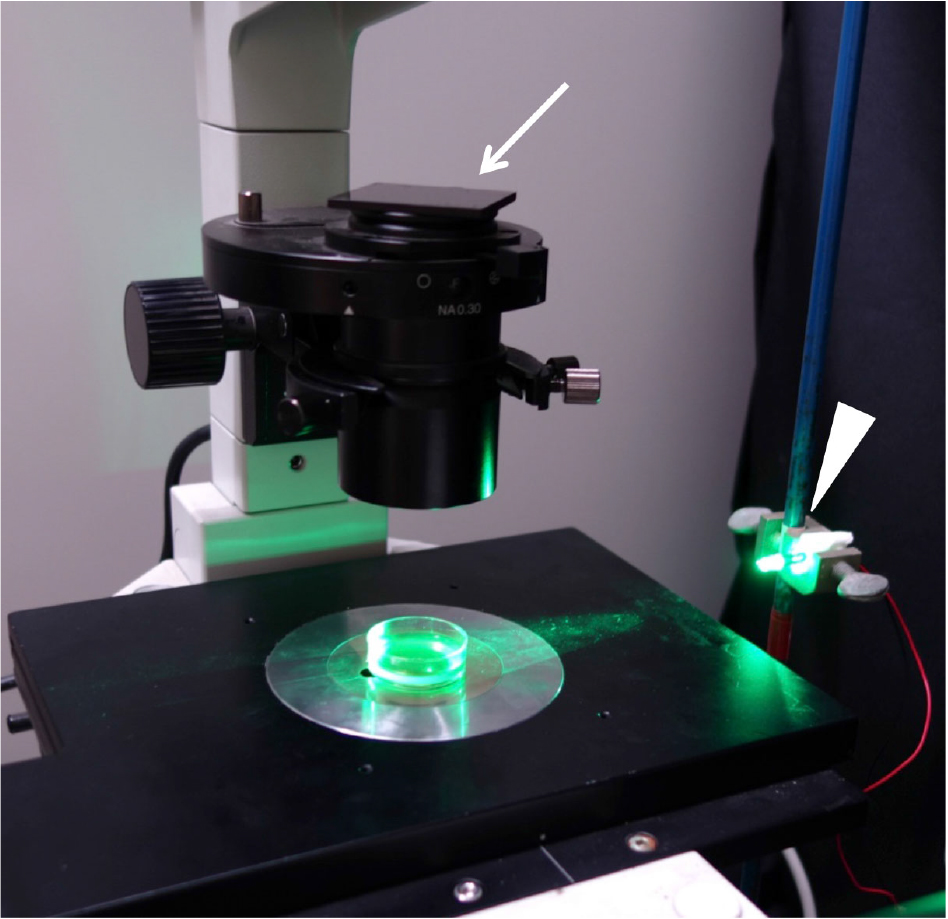

Figure 2. Setting for cell-level phototaxis analysis. A 4 cm Petri dish is set on a stage of the inverted microscope. Arrow indicates the red filter (630 nm long-pass filter). Arrowhead indicates the green LED.
Data analysis
- Track the swimming cells using a particle-tracking software. The Image Hyper software can be used in a semi-automatic manner. For tracking cells using Image J software, save the video in the uncompressed Audio Video Interleave (AVI) format. Playback the AVI file using ImageJ. Binarize the images (cells appear in black and background in white). Run MTrack2 plugin.
- Export the data of the trajectories (i.e., positions of cells at each frame) as the Comma-Separated Values (CSV) or any format for spreadsheet softwares.
- Measure the angle (θ) between the light direction and the swimming direction from the trajectories during 1.5 sec following illumination with a green LED for 15 sec (Figure 3A). The angle can be calculated by a spreadsheet software. (If Microsoft Excel is used, it can be calculated as following: ‘= degrees(atan2((x2 - x1),(y2 - y1)))’, where (x1, y1) represents the columns for the primary position of a cell and (x2, y2) represents those for the position after 1.5 sec.)
Note: For a few seconds after illumination, the photoshock response could occur and the phototactic behavior should be recorded after that period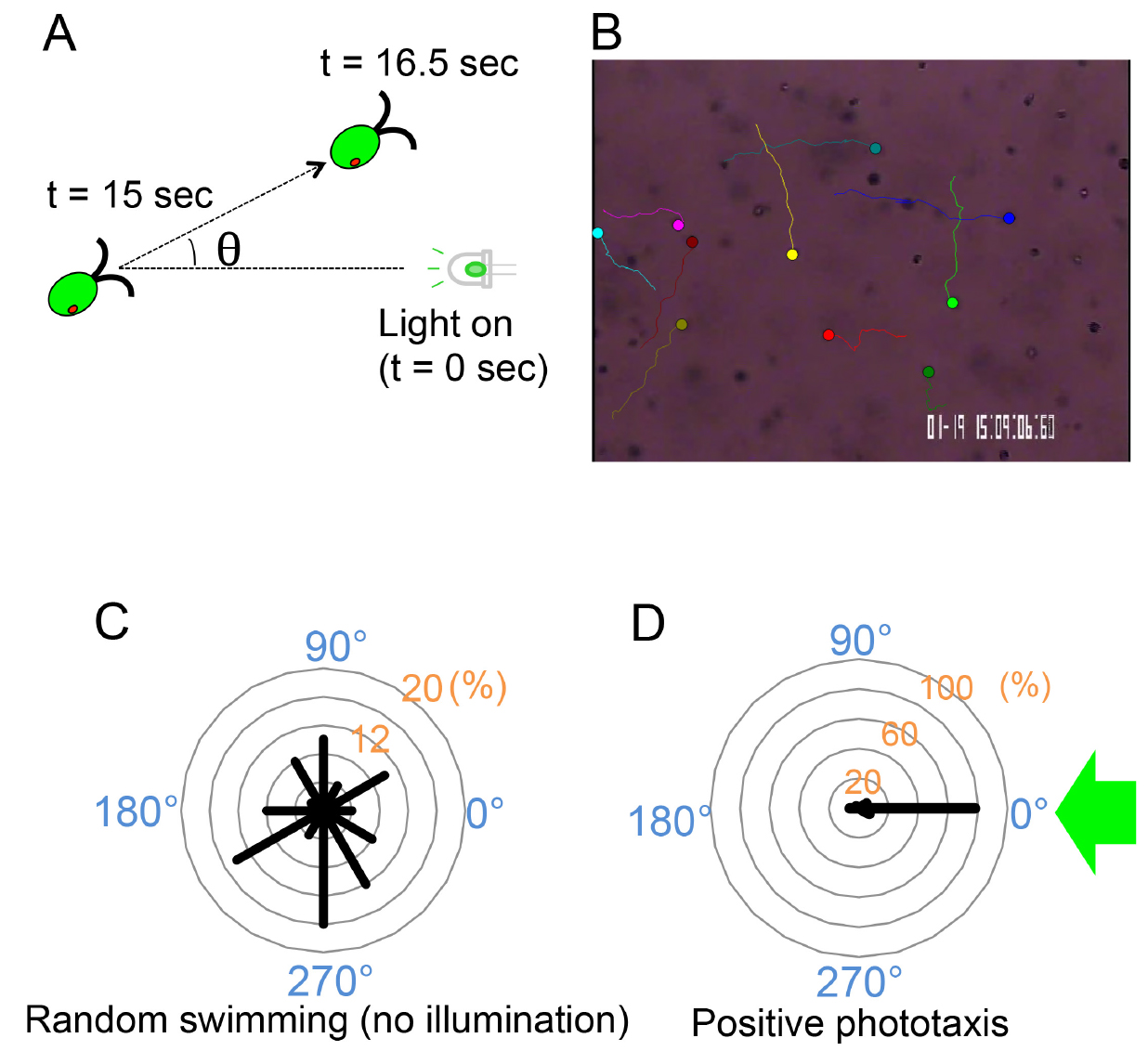
Figure 3. Cell-level phototaxis analysis. A. Swimming direction of each cell is measured for 1.5 sec following 15 sec illumination with a green LED to avoid the effects of photoshock response. B. An example of cell-level analysis (wild-type cells, random swimming). Well-focused images of swimming cells were auto-tracked using the Image Hyper software. Circles represent the starting points of cells, and lines indicate their swimming trajectories for 1.5 sec. (see Video 1). C and D. Polar histograms depicting the percentage of cells moving in a particular direction relative to light illuminated from the right (0°) (12 bins of 30°; n = 50 cells per condition). C. Representative data for random swimming when not illuminated; D. Representative data for positive phototaxis. ~80% of cells swim to 0°. Modified from (Wakabayashi et al., 2011).
Video 1. Tracking cells using Image Hyper software. Cells showing phototactic swimming were auto-tracked under a dark-field microscope. Four cells were tracked in this movie, and the former three cells show positive, and the last cell show negative phototaxis. (This software is a Japanese product and some letters in the screen are written in Japanese.) - For drawing polar histograms, typically, set 12 bins of 30° and draw a histogram (Figures 3C and 3D). (If necessary, more bins can be set [such as 24 bins of 15°].) If Microsoft Excel is used, draw graphs with ‘radar’ (Figure 4).
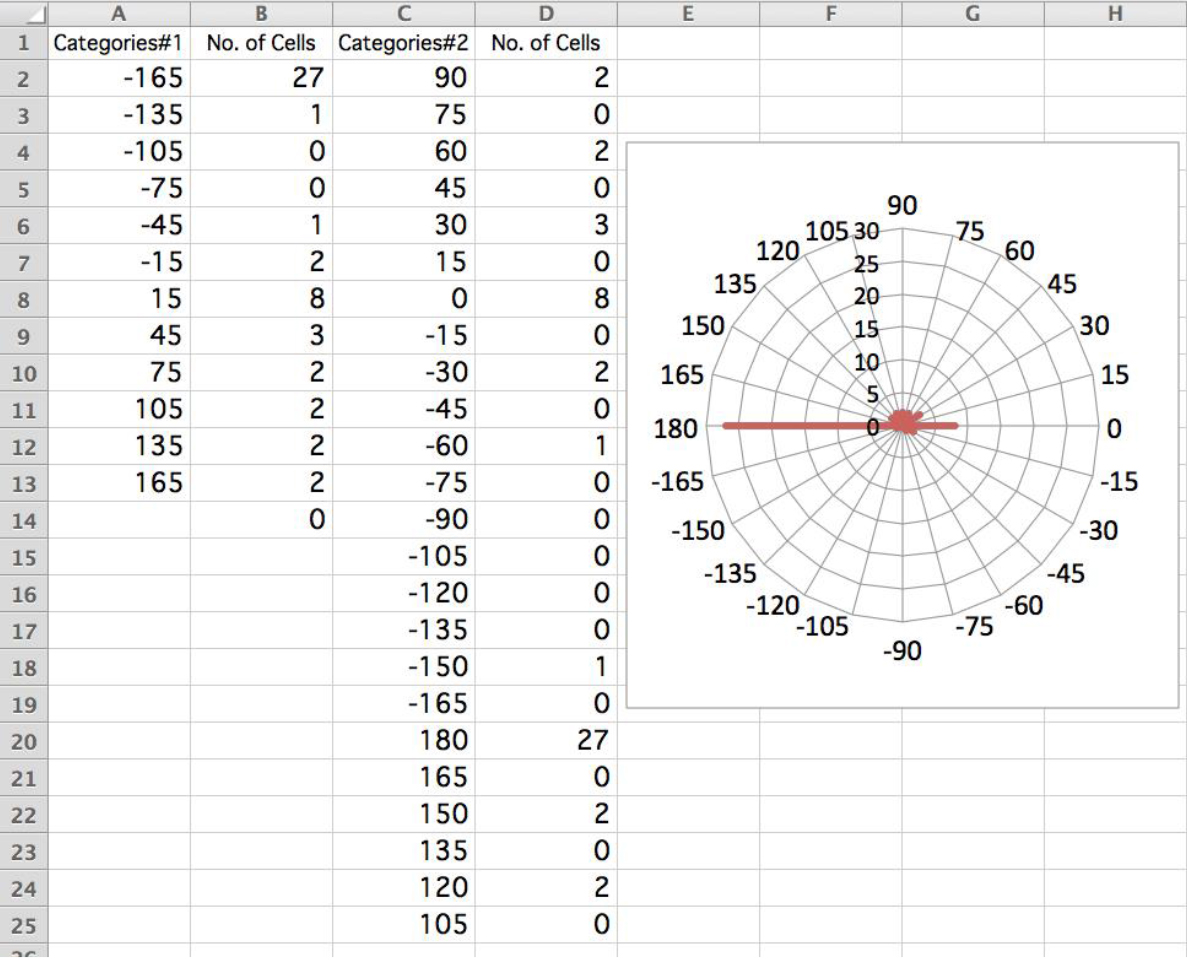
Figure 4. Drawing polar histogram. A screen shot of Microsoft Excel is shown. Calculate the first histogram (columns A and B) by the swimming angle against the light illumination axis. The numbers of cells are then pooled to different angular categories (columns C and D) as followings: D2 = B11, D4 = B10, D6 = B9, D8 = B8, D10 = B7, D12 = B6, D14 = B5, D16 = B4, D18 = B3, D20 = B2 + B14, D22 = B13, and D24 = B12. Insert an empty bin between each two categories (e.g., 75 between categories 90 and 60) so that the values are drawn as bars, not as the points of a polygon in the radar graph. In this example, most cells show negative phototaxis (the light is illuminated from 0°). - For estimation of phototactic index, calculate cosθ for each cell. When the sign of phototaxis is examined, average the value of cosθ. When cells are not illuminated and swim in random directions, the phototactic index should be ~0. When 100% of cells show clear positive or negative phototaxis, the phototactic index is 1 or -1, respectively. When the phototactic capability (i.e., swimming parallel to the light beam) is examined and the phototactic sign can be disregarded, average the value of |cosθ|. When 100% of cells swim parallel to the light beam, this index is 1. When cells swim in random directions, index is ~2/π (Okita et al., 2005).
Recipes
- Phototaxis assay solution (Okita et al., 2005)
5 mM HEPES (pH 7.4)
0.2 mM EGTA
1 mM KCl
0.3 mM CaCl2
Acknowledgments
This work was supported by JSPS KAKENHI Grant Numbers 15H01206, 15H01314, and 16K14752 to KW. This protocol was used in Wakabayashi et al., 2011 and Ueki et al., 2016.
References
- Berthold, P., Tsunoda, S. P., Ernst, O. P., Mages, W., Gradmann, D. and Hegemann, P. (2008). Channelrhodopsin-1 initiates phototaxis and photophobic responses in chlamydomonas by immediate light-induced depolarization. Plant Cell 20(6): 1665-1677.
- Foster, K. W., Saranak, J., Patel, N., Zarilli, G., Okabe, M., Kline, T. and Nakanishi, K. (1984). A rhodopsin is the functional photoreceptor for phototaxis in the unicellular eukaryote Chlamydomonas. Nature 311(5988):756-759.
- Foster, K. W. and Smyth, R. D. (1980). Light Antennas in phototactic algae. Microbiol Rev 44(4): 572-630.
- Gorman, D. S. and Levine R. P. (1965). Cytochrome f and plastocyanin: their sequence in the photosynthetic electron transport chain of Chlamydomonas reinhardi. Proc Natl Acad Sci U S A 54: 1665-1669.
- Hegemann, P. and Berthold, P. (2009). Sensory photoreceptors and light control of flagellar activity. In: George, B. W. (Ed). The Chlamydomonas Sourcebook Second Edition Volume 3. Academic Press pp: 395-430.
- Ide, T., Mochiji, S., Ueki, N., Yamaguchi, K., Shigenobu, S., Hirono, M. and Wakabayashi, K. (2016). Identification of the agg1 mutation responsible for negative phototaxis in a “wild-type” strain of Chlamydomonas reinhardtii. Biochem Biophys Reports 7: 379-385.
- Kamiya, R. and Witman, G. B. (1984). Submicromolar levels of calcium control the balance of beating between the two flagella in demembranated models of Chlamydomonas. J Cell Biol 98(1): 97-107.
- Kim, J. Y., Kwak, H. S., Sung, Y. J., Choi, H. I., Hong, M. E., Lim, H. S., Lee, J. H., Lee, S. Y. and Sim, S. J. (2016). Microfluidic high-throughput selection of microalgal strains with superior photosynthetic productivity using competitive phototaxis. Sci Rep 6: 21155.
- Mergenhagen, D. (1984). Circadian clock: genetic characterization of a short period mutant of Chlamydomonas reinhardii. Eur J Cell Biol 33(1): 13-18.
- Okita, N., Isogai, N., Hirono, M., Kamiya, R. and Yoshimura, K. (2005). Phototactic activity in Chlamydomonas 'non-phototactic' mutants deficient in Ca2+-dependent control of flagellar dominance or in inner-arm dynein. J Cell Sci 118(Pt 3): 529-537.
- Sineshchekov, O., Lebert, M. and Hader, D. P. (2000). Effects of light on gravitaxis and velocity in Chlamydomonas reinhardtii. J Plant Physiol 157(3): 247-254.
- Takahashi, T. and Watanabe, M. (1993). Photosynthesis modulates the sign of phototaxis of wild-type Chlamydomonas reinhardtii. Effects of red background illumination and 3-(3',4'-dichlorophenyl)-1,1-dimethylurea. FEBS Lett 336(3): 516-520.
- Takahashi, T., Yoshihara, K., Watanabe, M., Kubota, M., Johnson, R., Derguini, F. and Nakanishi, K. (1991). Photoisomerization of retinal at 13-ene is important for phototaxis of Chlamydomonas reinhardtii: simultaneous measurements of phototactic and photophobic responses. Biochem Biophys Res Commun 178(3): 1273-1279.
- Ueki, N., Ide, T., Mochiji, S., Kobayashi, Y., Tokutsu, R., Ohnishi, N., Yamaguchi, K., Shigenobu, S., Tanaka, K., Minagawa, J., Hisabori, T., Hirono, M. and Wakabayashi, K. (2016). Eyespot-dependent determination of the phototactic sign in Chlamydomonas reinhardtii. Proc Natl Acad Sci U S A 113(19): 5299-5304.
- Wakabayashi, K., Misawa, Y., Mochiji, S. and Kamiya, R. (2011). Reduction-oxidation poise regulates the sign of phototaxis in Chlamydomonas reinhardtii. Proc Natl Acad Sci U S A 108(27): 11280-11284.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Ueki, N. and Wakabayashi, K. (2017). Phototaxis Assay for Chlamydomonas reinhardtii. Bio-protocol 7(12): e2356. DOI: 10.21769/BioProtoc.2356.
Category
Plant Science > Plant physiology > Phenotyping
Plant Science > Plant cell biology > Cell imaging
Cell Biology > Cell imaging > Live-cell imaging
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link












