- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
RNA Capping by Transcription Initiation with Non-canonical Initiating Nucleotides (NCINs): Determination of Relative Efficiencies of Transcription Initiation with NCINs and NTPs
Published: Vol 7, Iss 12, Jun 20, 2017 DOI: 10.21769/BioProtoc.2336 Views: 9304
Reviewed by: Gal HaimovichMelike ÇağlayanAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Charging State Analysis of Transfer RNA from an α-proteobacterium
Liang Yin and Caroline S. Harwood
Dec 5, 2020 3506 Views

Preparation and Characterization of Internally Modified DNA Templates for Chemical Transcription Roadblocking
Eric J. Strobel
Sep 5, 2021 3597 Views
TGIRT-seq Protocol for the Comprehensive Profiling of Coding and Non-coding RNA Biotypes in Cellular, Extracellular Vesicle, and Plasma RNAs
Hengyi Xu [...] Alan M. Lambowitz
Dec 5, 2021 7288 Views
Abstract
It recently has been established that adenine-containing cofactors, including nicotinamide adenine dinucleotide (NAD+), reduced nicotinamide adenine dinucleotide (NADH), and 3’-desphospho-coenzyme A (dpCoA), can serve as ‘non-canonical initiating nucleotides’ (NCINs) for transcription initiation by bacterial and eukaryotic cellular RNA polymerases (RNAPs) and that the efficiency of the reaction is determined by promoter sequence (Bird et al., 2016). Here we describe a protocol to quantify the relative efficiencies of transcription initiation using an NCIN vs. transcription initiation using a nucleoside triphosphate (NTP) for a given promoter sequence.
Keywords: RNA polymeraseBackground
Transcription in bacteria, archaea, and eukaryotes is carried out by multi-subunit RNA polymerases (RNAPs) conserved in sequence, structure, and mechanism (Ebright, 2000; Lane and Darst, 2010). To initiate transcription, RNAP, together with one or more initiation factors, binds to a specific DNA sequence referred to as a ‘promoter’ and unwinds promoter DNA to form an RNAP-promoter open complex (RPo) containing an unwound ‘transcription bubble’ (Figure 1A; Ruff et al., 2015). RNAP then selects a transcription start site by expanding (‘scrunching’) or contracting (‘antiscrunching’) the transcription bubble to place transcription-start-site nucleotides in the RNAP active-center initiating site (‘i site’) and extending site (‘i+1 site’), binds a complementary initiating nucleotide substrate in the i site and a complementary extending substrate in the ‘i+1’ site, and catalyzes phosphodiester-bond formation to yield an initial RNA product (Winkelman et al., 2016).
In standard de novo transcription initiation, the initiating substrate is a nucleoside triphosphate (NTP), typically ATP or GTP (Nickels and Dove, 2011). However, recently it has been established that adenine-containing cofactors, including nicotinamide adenine dinucleotide (NAD+), reduced nicotinamide adenine dinucleotide (NADH), and 3’-desphospho-coenzyme A (dpCoA), can serve as alternative initiating substrates (‘non-canonical initiating nucleotides’; NCINs), yielding NCIN-capped RNA products that have distinctive 5’-end structures, stabilities, and translation efficiencies (Figures 1B-1C; Bird et al., 2016; Barvik et al., 2016; Jiao et al., 2017; Walters et al., 2017). It further has been established that the relative efficiencies of NCIN-mediated initiation vs. NTP-mediated initiation are determined by promoter sequence (Bird et al., 2016).
Here, we describe a protocol to determine the relative efficiencies of NCIN-mediated transcription initiation versus ATP-mediated transcription initiation, (kcat/KM, NCIN)/(kcat/KM, ATP), for a given promoter sequence. The protocol involves generating radiolabeled initial RNA products in a set of transcription reactions having a constant concentration of NCIN and varying concentrations of ATP, followed by quantifying NCIN-initiated RNA and total RNA, followed by plotting observed ratios of NCIN-initiated RNA to total RNA as a function of ratios of NCIN concentration to ATP concentration. 
Figure 1. Transcription initiation. A. RNAP-promoter open complex (RPo) with unwound transcription bubble. Gray, RNAP; blue, -10-element nucleotides; i and i+1, RNAP active-center initiating nucleotide binding site and extending nucleotide binding site; boxes, DNA nucleotides (nontemplate-strand nucleotides above template-strand nucleotides). B. Structures of ATP and NAD+, Red, identical atoms in ATP and NAD+; C. Initial RNA products formed in transcription initiation using ATP (top) or transcription initiation using NAD+ (bottom). Left subpanels show initiating ATP or NAD+ bound in i site; right subpanels show initial RNA products formed using CTP as extending nucleotide. Red boxes, adenosine and cytosine moieties of ATP, NAD+, and CTP; green boxes, nicotinamide-riboside moiety of NAD+.
Materials and Reagents
- E. coli RNA polymerase σ70 holoenzyme
Note: Prepared as in Mukhopadhyay et al. (2003) or purchased (New England Biolabs, catalog number: M0551S ) - E. coli RNA polymerase core enzyme
Note: Prepared as in Artsimovitch et al. (2003). - E. coli σ70
Note: Prepared as Marr and Roberts (1997); Perdue and Roberts (2010). - NAD+ (grade I, free acid) (Roche Molecular Systems, catalog number: 10127965001 )
- NADH (grade I, free acid) (Roche Molecular Systems, catalog number: 10107735001 )
- Phusion Flash HF master mix (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: F548L )
Note: For generating transcription templates. - Oligodeoxyribonucleotides (template and primers) (Integrated DNA Technologies, www.IDTdna.com)
- QIAquick PCR purification kit (QIAGEN, catalog number: 28106 )
- TEMED (Avantor Performance Materials, J.T. Baker®, catalog number: 4098-01 )
- Ammonium persulfate (VWR, AMRESCO, catalog number: 97064-594 )
- GeneMate LE Quick-Dissolve agarose (BioExpress, catalog number: E-3119-500 )
- 3’-Desphosphocoenzyme A (Sigma-Aldrich, catalog number: D3385 )
- SequaGel sequencing system (National Diagnostics, catalog number: EC-833 )
- High purity rNTP set (ATP, UTP, GTP, CTP) (100 mM) (GE Healthcare, catalog number: 27-2025-01 )
- [α-32P]-CTP EasyTide (3,000 Ci/mmol) (250 μCi) (Perkin Elmer, catalog number: BLU508H250UC )
- Tris base (VWR, AMRESCO, catalog number: 97061-800 )
- Potassium chloride (KCl) (EMD Millipore, catalog number: PX1405-1 )
- Magnesium chloride hexahydrate (EMD Millipore, catalog number: 5980-500GM )
- EDTA disodium salt dyhydrate (1 kg) (VWR, AMRESCO, catalog number: 97061-018 )
- Dithiothreitol (DTT) (Gold Bio, catalog number: DTT50 )
- Bovine serum albumin (BSA) fraction V (Alfa Aesar, Affymetrix/USB, catalog number: J10857 )
- Sodium dodecylsulfate (SDS) (VWR, AMRESCO, catalog number: 97064-470 )
- Deionized formamide (EMD Millipore, catalog number: 4610-100ML )
- Xylene cyanol (Sigma-Aldrich, catalog number: X4126-10G )
- Bromophenol blue (EMD Millipore, catalog number: BX1410-7 )
- Amaranth red (Acros Organics, catalog number: AC15303-0250 )
- Boric acid (ACS grade) (VWR, AMRESCO, catalog number: 97061-980 )
- Sodium acetate, trihydrate (Avantor Performance Materials, MACRON, catalog number: 7364-06 )
- Hydrochloric acid (ACS plus) (Fisher Scientific, catalog number: A144-212 )
- Glycerol (ACS grade) (EMD Millipore, catalog number: GX0185-5 )
- Transcription buffer (1x) (see Recipes)
- Transcription buffer (5x) (see Recipes)
- Transcription stop buffer (see Recipes)
- Tris-borate EDTA buffer (TBE) (see Recipes)
- TBE + 0.3 M sodium acetate (see Recipes)
Equipment
- NanoDrop 2000c spectrophotometer ND2000C (Thermo Fisher Scientific, Thermo ScientificTM, model: NanoDropTM 2000/2000c , catalog number: ND-2000C)
- Glass plate
- Block digital heater w/20 tapered hole blocks (VWR, catalog numbers: 12621-088 ; 13259-002 )
- 5424 table top centrifuge with w/FA-45-24-11 rotor (Eppendorf, mdoel: 5424/5424 R , catalog number: 5424000410)
- Powerpack HV powersupply (Bio-Rad Laboratories, model: PowerPac HV Power Supply, catalog number: 1645056 )
- Sequi-gen GT sequencing gel system (38 x 30 cm gel) (Bio-Rad Laboratories, catalog number: 1653862 )
Note: This product has been discontinued. - Hydrotech vacuum pump (Bio-Rad Laboratories, catalog number: 1651781 )
- Model 583 gel dryer (Bio-Rad Laboratories, model: Model 583, catalog number: 1651745 )
- DNA Engine Dyad PCR Machine 4 x 48 well blocks (Bio-Rad Laboratories)
- Unmounted phosphor exposure screen (35 x 43 cm) (GE Healthcare, model: General Purpose Screens, catalog number: 63-0034-79 )
- Storm 840 scanner (Molecular Dynamics, model: Storm 840 )
- Windows computer (HP, model: Compaq dc7700 )
Software
- Excel (Microsoft)
- ImageQuant (GE)
- SigmaPlot (Systat)
Procedure
Notes:
- The presented procedure is for analysis of Escherichia coli RNAP and the transcription initiation factor σ70. The procedure can be adapted to analysis of any other RNAP by replacing E. coli RNAP with the RNAP of interest, replacing the transcription factor σ70 by the transcription initiation factor(s) used by the RNAP of interest, replacing the promoter by a promoter used by the RNAP of interest, and replacing the transcription buffer with a transcription buffer suitable for the RNAP of interest.
- DNA templates are designed to facilitate generation and analysis of defined initial RNA products formed using NAD+, NADH, dpCoA, or ATP as the initiating substrate (Figure 2A). Initiation using NAD+, NADH, dpCoA, or ATP requires A:T (i.e., template-strand T) at the transcription start site (position +1; Bird et al., 2016). Use of CTP as the extending nucleotide requires C:G (i.e., template-strand G) at the first position downstream of the transcription start site (position +2). Generation of defined initial RNA products representing one, and only one, nucleotide-addition reaction requires a T:A or C:G (i.e., template-strand A or G, which are not complementary to NTPs present in reactions).
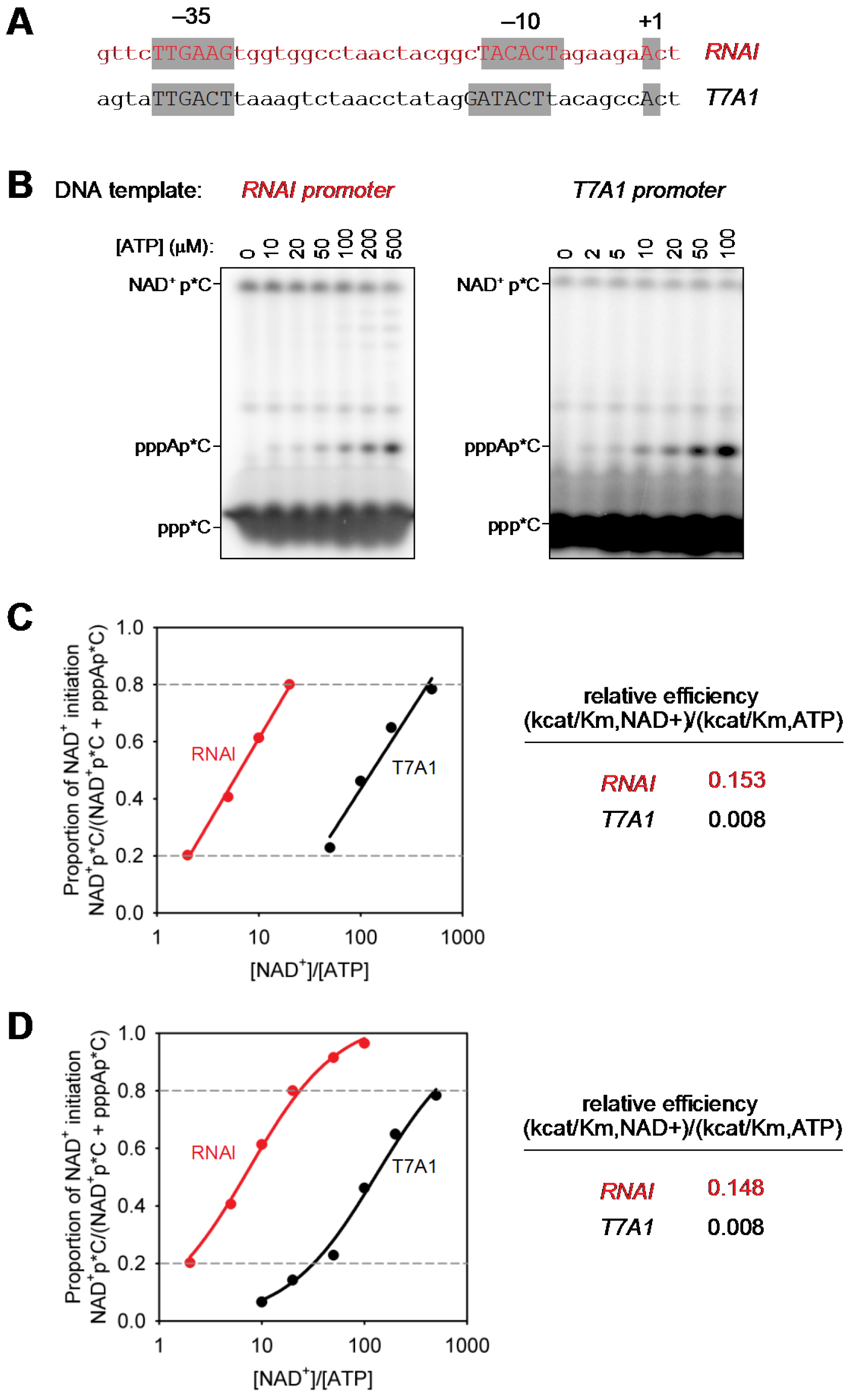
Figure 2. Determination of relative efficiencies of transcription initiation with NAD+ vs. transcription initiation with ATP. A. dsDNA transcription templates containing RNAI and T7A1 promoters (positions -40 to +3; promoter elements and transcription start sites in gray boxes); B. Representative raw data (initial RNA products of transcription reactions performed in the presence of 1 mM NAD+, 0.01-0.5 mM ATP, and [α32P]-CTP as extending nucleotide. C and D. Relative efficiencies of transcription initiation with NAD+ vs. transcription initiation with ATP [(kcat/KM, NCIN)/(kcat/KM, ATP)]. Calculation using logarithmic regression (C; best-fit line for data points with NAD+pC/(pppApC + NAD+pC) values between 0.2 and 0.8 in (C) or non-linear regression (D); best-fit curve for data points with NAD+pC/(pppApC + NAD+pC) values between 0 and 1.
- Generation of transcription templates
- Prepare template and primers as follows:
- 100 nt oligodeoxyribonucleotide corresponding to promoter template-strand positions -65 to +35 (‘template’; IDT, Inc.; dissolved in water to 100 μM).
- 20 nt oligodeoxyribonucleotide complementary to promoter template-strand positions -65 to -46 (‘forward primer’; IDT, Inc.; dissolved in water to 100 μM).
- 20 nt oligodeoxyribonucleotide identical to promoter template-strand positions +16 to +35 (‘reverse primer’; IDT, Inc.; dissolved in water to 100 μM).
- 100 nt oligodeoxyribonucleotide corresponding to promoter template-strand positions -65 to +35 (‘template’; IDT, Inc.; dissolved in water to 100 μM).
- Run PCR as follows (35 cycles):

PCR cycles:
- Purify PCR products using QIAquick PCR purification kit (two columns per PCR reaction, each eluted in 40-50 μl water). (Two columns are needed because the DNA quantity exceeds the binding capacity of a single column.)
- Analyze aliquots by 2% agarose gel electrophoresis to confirm production of 100 bp double-stranded DNA fragment (‘dsDNA transcription template’).
- Quantify dsDNA transcription template by UV-Vis spectrophotometry, and adjust concentration to 0.5-1 μM in water.
- Prepare template and primers as follows:
- Transcription reactions
Template + RNAP mix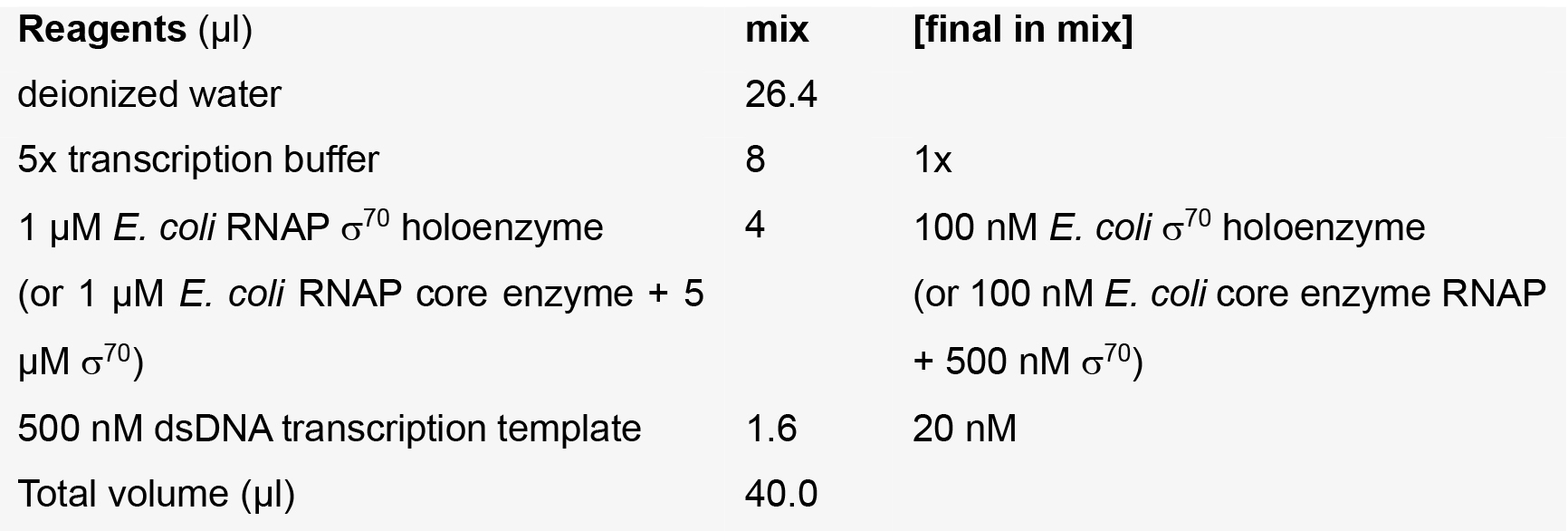
NCIN + NTP mixes
Note: The water, 5x transcription buffer, [α32P]-CTP, CTP, and NCIN are pre-mixed to yield a ‘master mix’, of which 4 μl is added to 1 μl of the appropriate ATP dilution.- Incubate Template + RNAP mix at 37 °C for 15 min to enable formation of a catalytically competent RNAP-promoter open complex.
- Add 5 μl Template + RNAP mix to pre-warmed NCIN + NTP mixes, and incubate 10 min at 37 °C.
- Add 10 μl ice-cold stop buffer. (Keep samples on ice if gel is to be loaded immediately, or at -20 °C if gel is to be loaded later.)
Note: Do not boil samples that contain NCIN-capped RNA before loading gels. (NCINs can be heat-labile.) - Analyze products by electrophoresis on 7.5 M urea 20% polyacrylamide nucleotide sequencing gel (National Diagnostics, Inc.; prepared per instructions of the vendor, but using one-half of specified ammonium persulfate and Temed concentrations)
- Pour gel.
- Pre-run gel at 50 W for 20-60 min before loading samples (top reservoir buffer, TBE; bottom reservoir buffer, TBE containing 0.3 M sodium acetate).
- Load samples (5 μl per lane).
Optional: In a lane adjacent to sample lanes, load 1 μl transcription stop buffer with added 0.025% amaranth red as a marker. Amaranth red migrates similarly to free NTPs and allows a visual estimation of migration of dinucleotide products. - Run gel at 50 W (buffers as above) for 120 min or until desired separation is achieved.
Note: The addition of 0.3 M sodium acetate to the bottom reservoir buffer creates a salt gradient that compresses and improves resolution for short RNA products (Vo et al., 2003).
- Pour gel.
- Disassemble gel apparatus. Wrap gel on glass plate with polyethylene wrap, and expose to storage-phosphor screen for 3-18 h at 4 °C.
- Incubate Template + RNAP mix at 37 °C for 15 min to enable formation of a catalytically competent RNAP-promoter open complex.
Data analysis
- Scan storage phosphor screen using storage phosphor imager (Storm, Typhoon, or equivalent; GE, Inc.).
- Open gel images in ImageQuant (GE, Inc.). and quantify band intensities using one of the following methods:
- Box method: Using the ImageQuant ‘Rectangle’ tool, draw uniform-size boxes around bands of interest (defining box for first band, and then copying and pasting box to each other band of interest). Perform background correction (either by using ImageQuant ‘Background Correction’ tool to define and subtract average background, or by manually subtracting background for an identically sized box placed in a region without bands). Quantify band intensities using the ImageQuant ‘Volume Report’ tool.
- Line method: Using the ImageQuant ‘Line’ tool, draw uniform-width lines vertically through each lane (defining line and adjusting line-width to be slightly narrower than bands for first lane, and then copying and pasting line to each other lane). Using ImageQuant ‘Create Graph’ tool, create graph reporting area under the line for each lane, define bands, and correct for background using ImageQuant ‘Peak Finder’ tool or ImageQuant ‘Define Peak’, ‘Split Peak’, and base-line-adjustment tools. Quantify band intensities using ImageQuant ‘Area Report’ tool.
- Calculate relative efficiencies of NCIN-mediated initiation vs. ATP-mediated initiation [(kcat/KM, NCIN)/(kcat, ATP/KM, ATP)] using one of the following methods:
- Logarithmic regression (using Excel or SigmaPlot): Plot observed values of NCINpC/(pppApC + NCINpC) vs. [NCIN]/[ATP] on a semi-log plot, using only observed values of NCINpC/(pppApC + NCINpC) between 0.2 and 0.8 [i.e., using only values of NCINpC/(pppApC + NCINpC) for the part of the curve that can be approximated as a line]. Perform logarithmic regression, fitting data to:
y = y0 + a[ln(x)]
where, y is NCINpC/(pppApC + NCINpC), x is [NCIN]/[ATP], and y0 and a are regression parameters. The resulting fit yields the value of x for which y = 0.5. The relative efficiency (kcat/KM, NCIN)/(kcat/KM, ATP) is equal to 1/x. - Non-linear regression (using SigmaPlot): Plot observed values of NCINpC/(pppApC + NCINpC) vs. [NCIN]/[ATP] on semi-log plot, using all observed values of NCINpC/(pppApC + NCINpC) [i.e., using values of NCINpC/(pppApC + NCINpC) not only for the part of the curve that can be approximated as a line but also for the parts of the curve that cannot be approximated as a line]. Perform non-linear regression, fitting data to:
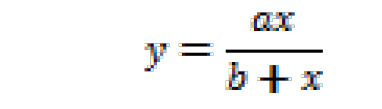
where, y is NCINpC/(pppApC + NCINpC), x is [NCIN]/[ATP], and a and b are regression parameters. The resulting fit yields the value of x for which y = 0.5. The relative efficiency (kcat/KM, NCIN)/(kcat/KM, ATP) is equal to 1/x.
Recipes
- Transcription buffer (1x)
10 mM Tris HCl pH 8.0
40 mM KCl
10 mM MgCl2
0.1 mM EDTA
1 mM DTT
0.1 mg/ml BSA - Transcription buffer (5x)
50 mM Tris HCl pH 8.0
200 mM KCl
50 mM MgCl2
0.5 mM EDTA
5 mM DTT
0.5 mg/ml BSA - Transcription stop buffer
100 mM Tris HCl pH 8.0
18 mM EDTA
1.25% SDS
90% formamide
0.025% xylene cyanol
0.025% bromophenol blue
0.025% amaranth red - Tris-borate EDTA buffer (TBE)
90 mM Tris base
90 mM boric acid
2 mM EDTA disodium salt - TBE + 0.3 M sodium acetate
90 mM Tris base
90 mM boric acid
2 mM EDTA disodium salt
300 mM sodium acetate
Acknowledgments
This work was supported by National Institutes of Health grants GM118059 (B.E.N.) and GM041376 (R.H.E.). The protocol was adapted from methods reported in Bird et al. (2016).
References
- Artsimovitch, I., Svetlov, V., Murakami, K. S. and Landick, R. (2003). Co-overexpression of Escherichia coli RNA polymerase subunits allows isolation and analysis of mutant enzymes lacking lineage-specific sequence insertions. J Biol Chem 278(14): 12344-12355.
- Barvik, I., Rejman, D., Panova, N., Sanderova, H. and Krasny, L. (2016). Non-canonical transcription initiation: the expanding universe of transcription initiating substrates. FEMS Microbiol Rev.
- Bird, J. G., Zhang, Y., Tian, Y., Panova, N., Barvik, I., Greene, L., Liu, M., Buckley, B., Krasny, L., Lee, J. K., Kaplan, C. D., Ebright, R. H. and Nickels, B. E. (2016). The mechanism of RNA 5’ capping with NAD+, NADH and desphospho-CoA. Nature 535(7612): 444-447.
- Ebright, R. H. (2000). RNA polymerase: structural similarities between bacterial RNA polymerase and eukaryotic RNA polymerase II. J Mol Biol 304(5): 687-698.
- Jiao, X., Doamekpor, S. K., Bird, J. G., Nickels, B. E., Tong, L., Hart, R. P. and Kiledjian, M. (2017). 5' end nicotinamide adenine dinucleotide cap in human cells promotes RNA decay through DXO-mediated deNADding. Cell 168(6): 1015-1027 e1010.
- Lane, W. J. and Darst, S. A. (2010). Molecular evolution of multisubunit RNA polymerases: sequence analysis. J Mol Biol 395(4): 671-685.
- Marr, M. T. and Roberts, J. W. (1997). Promoter recognition as measured by binding of polymerase to nontemplate strand oligonucleotide. Science 276(5316): 1258-1260.
- Mukhopadhyay, J., Mekler, V., Kortkhonjia, E., Kapanidis, A. N., Ebright, Y. W. and Ebright, R. H. (2003). Fluorescence resonance energy transfer (FRET) in analysis of transcription-complex structure and function. Methods Enzymol 371: 144-159.
- Nickels, B. E. and Dove, S. L. (2011). NanoRNAs: a class of small RNAs that can prime transcription initiation in bacteria. J Mol Biol 412(5): 772-781.
- Perdue, S. A. and Roberts, J. W. (2010). A backtrack-inducing sequence is an essential component of Escherichia coli sigma(70)-dependent promoter-proximal pausing. Mol Microbiol 78(3): 636-650.
- Ruff, E. F., Record, M. T., Jr. and Artsimovitch, I. (2015). Initial events in bacterial transcription initiation. Biomolecules 5(2): 1035-1062.
- Vo, N. V., Hsu, L. M., Kane, C. M. and Chamberlin, M. J. (2003). In vitro studies of transcript initiation by Escherichia coli RNA polymerase. 3. Influences of individual DNA elements within the promoter recognition region on abortive initiation and promoter escape. Biochemistry 42(13): 3798-3811.
- Walters, R. W., Matheny, T., Mizoue, L. S., Rao, B. S., Muhlrad, D. and Parker, R. (2017). Identification of NAD+ capped mRNAs in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A 114(3): 480-485.
- Winkelman, J. T., Vvedenskaya, I. O., Zhang, Y., Zhang, Y., Bird, J. G., Taylor, D. M., Gourse, R. L., Ebright, R. H. and Nickels, B. E. (2016). Multiplexed protein-DNA cross-linking: Scrunching in transcription start site selection. Science 351(6277): 1090-1093.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Bird, J. G., Nickels, B. E. and Ebright, R. H. (2017). RNA Capping by Transcription Initiation with Non-canonical Initiating Nucleotides (NCINs): Determination of Relative Efficiencies of Transcription Initiation with NCINs and NTPs. Bio-protocol 7(12): e2336. DOI: 10.21769/BioProtoc.2336.
Category
Microbiology > Microbial biochemistry > RNA
Molecular Biology > RNA > Transcription
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









