- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Mismatched Primer Extension Assays
Published: Vol 5, Iss 12, Jun 20, 2015 DOI: 10.21769/BioProtoc.1508 Views: 8093
Reviewed by: Yu ChenChang Ho LeeAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
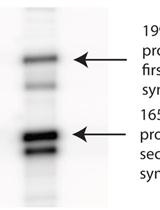
Primer Extension Reactions for the PCR- based α- complementation Assay
Vasudevan Achuthan and Jeffrey J. DeStefano
Jun 20, 2015 11324 Views
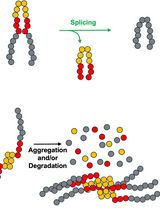
Monitoring Protein Stability In Vivo Using an Intein-Based Biosensor
John S. Smetana [...] Christopher W. Lennon
Apr 20, 2025 1575 Views
Endo-1,4-β-D-xylanase Assay Using Azo-Xylan and Variants Thereof
Luca Bombardi [...] Salvatore Fusco
Apr 20, 2025 1916 Views
Abstract
Steady state kinetic assays have been a reliable way to estimate fidelity of several polymerases (Menendez-Arias, 2009; Rezende and Prasad, 2004; Svarovskaia et al., 2003). The ability to analyze the extension of primers with specific mismatches at the 3ʹ end is a major strength of the mismatched primer extension assays. Recently, we used the mismatched primer extension assays to show that the fidelity of HIV RT increases dramatically when concentration of Mg2+ is reduced to a physiologically relevant concentration (~0.25 mM) (Achuthan et al., 2014). Here, we describe in detail how to perform the mismatched primer extension assay to measure the standard extension efficiency using human immunodeficiency virus reverse transcriptase (HIV RT) at 2 mM Mg2+. The relative fidelity of the polymerase can then be estimated using the standard extension efficiency. The assay described here is based on the method published in Mendelman et al. (1990).
Materials and Reagents
- Deoxynucleoside triphosphate (Roche Diagnostics, catalog number: 11969064001 )
- Gamma [γ-32P] ATP (PerkinElmer, catalog number: Blu502A001MC )
- G-25 Macro spin columns (best suited for volumes of 75-150 μl) (Harvard Apparatus, catalog number: 74-3901 )
- 40% Acrylamide-Bisacrylamide (19:1) solution (VWR International, catalog number: JT4969-0 )
- T4 polynucleotide kinase (PNK) (New England Biolabs, catalog number: M0201L )
- 10X T4 polynucleotide kinase buffer (New England Biolabs, catalog number: B0201S )
- Urea (VWR International, catalog number: 97061-926 )
- Ammonium Persulfate (VWR International, catalog number: 97064-594 )
- HIV Reverse Transcriptase, purified as described in Hou et al. (2004)
- DNA oligonucleotides from Integrated DNA Technologies
- Template:
5'-GGGCGAATTTAG[G/C]TTTTGTTCCCTTTAGTGAGGGTTAATTTCGAGCTTG G-3’. The underlined nucleotides in brackets indicate that templates with either a G or C at this position can be used depending on the type of mismatch examined. - Primer:
5ʹ-TAACCCTCACTAAAGGGAACAAAAX-3ʹ. “X” at the 3ʹ end of the primer denotes A, T, or C depending on the mismatch examined. “X” in the case of a matched primer is G.
- Template:
- 1 M MgCl2
- Extension reaction buffer (see Recipes)
- 2x loading dye (see Recipes)
Equipment
- Eppendorf tubes
- Micropipette
- Table top centrifuge
- Incubator
- Gel apparatus
Software
- Sigmaplot Version 10.0 (Sysstat Software)
Procedure
- Primer labelling
- All the primers should be first radiolabelled in 50 µl of 1x PNK buffer along with 50 pico moles of each primer, 10 μl of [γ-32P] ATP and 5 units of PNK.
Note: The reaction mixture was incubated for 30 min at 37 °C and the PNK was heat inactivated for 15 min at 65 °C. - G-25 spin columns were incubated with 500 µl dH2O for 15 min to equilibrate the column and the water was removed by spinning the columns at a table top centrifuge at 5,000 rpm for 4 min.
- After heat inactivation, the excess [γ-32P] ATP was removed from the reaction mixture by loading it onto an equilibrated column and spinning at 5,000 rpm for 4 min.
- All the primers should be first radiolabelled in 50 µl of 1x PNK buffer along with 50 pico moles of each primer, 10 μl of [γ-32P] ATP and 5 units of PNK.
- Matched primer extension reactions
To obtain information about the standard extension efficiency, extension of matched as well as mismatched primers should be performed. The standard extension efficiency can then be calculated as the ratio of efficiency of extending mismatched primers to efficiency of extending matched primers.- Eight matched primer extension reactions were set up. For each reaction, 14 nM of the radiolabeled primer was hybridized to 14 nM of the template (1:1 ratio of primer:template) in 7 μl of the extension reaction buffer. The mixture was heated at 65 °C for 5 min and then slowly cooled to room temperature.
- The hybrid was then incubated for 3 min at 37 °C in the reaction buffer along with 2 μl of 10 mM MgCl2 (final concentration of 2 mM MgCl2) and 2 μl of the nucleotide substrate (concentration varies for each reaction, see below) for each reactions. The nucleotide substrate is the next correct nucleotide to be added and it depends on the template used in the reactions. For this particular template, dCTP was the substrate (Figure 1). For matched primer extension reactions, the eight reactions had a final concentration of dCTP in the order of 0, 0.02, 0.04, 0.1, 0.2, 0.3, 0.6 and 1 μM respectively.

Figure 1. Constructs used in mismatched primer extension assays. The sequence of the DNA templates is shown at the bottom and the sequence of the primer at the top. The underlined nucleotides show the only differences between the two templates. - The extension was then initiated by addition of 2 μl of 13 nM HIV RT (2 nM final concentration). The total reaction volume was 13 μl.
- After 2 min, reactions were terminated by addition of 13 μl of 2x loading dye.
Note: Reactions were run only for 2 min to ensure the primer is extended by only one nucleotide. - The reaction products were then electrophoresed on 16% denaturing 7 M urea-polyacrylamide gels, dried, and imaged using a Fujifilm FLA5100 phosphorimager.
Note: The samples were run far enough to separate the extended band from the primer band (Figure 2).
- Mismatched primer extension reactions
- For mismatched primer extension reactions, a different radiolabeled primer, depending on the mismatch analyzed (Figure 1), is used. Primer-template hybrids were made as described above.
- Eight individual reactions were set up. 7 μl of primer-template hybrids was incubated at 37 °C in the reaction buffer for 3 min along with 2 μl of 10 mM MgCl2 (final concentration of 2 mM MgCl2) and 2 μl of the nucleotide substrate. The total reaction volume was 13 μl.
Note: Mismatched primer-template sequences require more substrate for extension than matched primer-template sequences. So, the eight reactions had a final concentration of dCTP in the order of 0, 50, 100, 200, 400, 630, 1,200 and 1,870 μM respectively (Figure 2). - Extension was initiated by addition of 2 μl of 13 nM HIV RT.
- After 5 min of extension, the reactions were terminated by addition of 13 μl of 2x loading dye and the extension products were processed on a 16% denaturing polyacrylamide gel as described above. The gel was run at 75 Watts for 90 min.
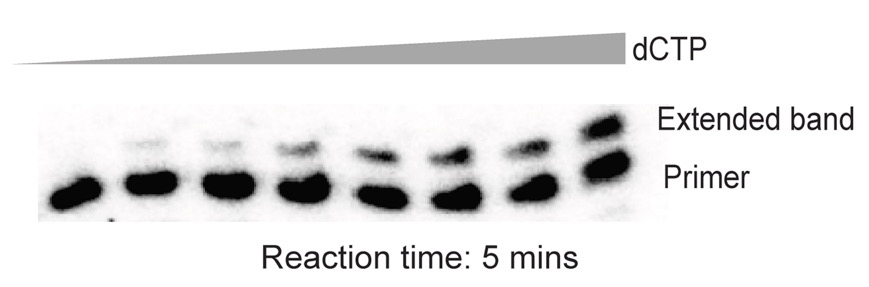
Figure 2. Representative data for the mismatched primer extension assay. Primer: template hybrid used here had a C.A mismatch at the 3ʹ terminus. Concentration of dCTP used in each of the lanes (left to right): 0, 50, 100, 200, 400, 630, 1,200 and 1,870 μM.
- Calculation of standard extension efficiency
- Velocity measurements were performed according to Mendelman et al. (1990). Velocity (
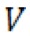 rel) of extending each primer by one nucleotide was calculated according to the formula:
rel) of extending each primer by one nucleotide was calculated according to the formula: 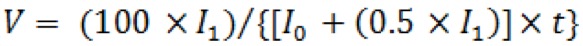 , where
, where 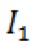 is the intensity of the extended band, is the intensity of the primer band, and t is the time of extension.
is the intensity of the extended band, is the intensity of the primer band, and t is the time of extension. 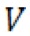 rel (percentage of primer extended per minute) was then plotted against [S], where [S] is the concentration of the substrate (dCTP) used in the reaction (Figure 3).
rel (percentage of primer extended per minute) was then plotted against [S], where [S] is the concentration of the substrate (dCTP) used in the reaction (Figure 3).
Note: These curves are typical of Michaelis-Menten kinetics curves.- The values of Vmax and Km were obtained by curve-fitting. The data were fitted to a “Single rectangular 2-parameter hyperbola equation” using the Sigmaplot software.
- The constant “a” obtained after curve fitting corresponds to the Vmax and “b” corresponds to the Km for this extension. Vmax and Km values were individually determined for both matched and mismatched primer extension reactions.
- The standard extension efficiency,
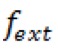 , was calculated as the ratio of
, was calculated as the ratio of 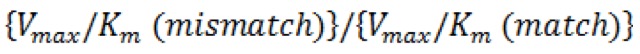 .
.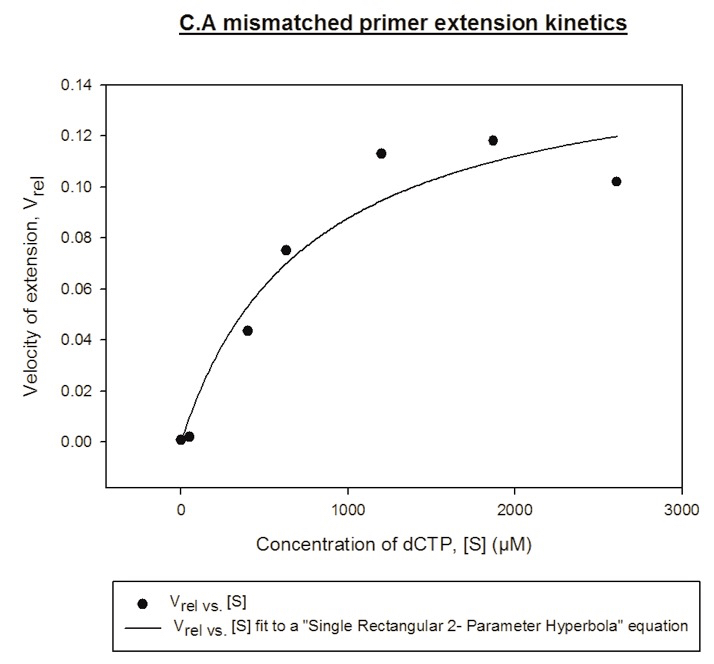
Figure 3. Representative graph for the mismatched primer extension assay. Primer:template hybrid used here had a C.A mismatch at the 3ʹ terminus. Vmax and Km were determined by fitting the points to a “Single rectangular 2-parameter hyperbola equation” using the Sigmaplot software.
- Velocity measurements were performed according to Mendelman et al. (1990). Velocity (
Recipes
- Extension reaction buffer (50 ml)
1 M Tris HCl (pH 8) 25 ml 3 M KCl 13.3 ml 1 M DTT 1 ml RNase free water 10.7 ml - 2x loading dye (10 ml)
50 mM Tris HCl (pH 6.8) 500 μl 100 mM DTT 1 ml 2% SDS 2 ml 0.05% Bromophenol blue 500 μl 10% glycerol 1 ml RNase free water 5 ml
References
- Achuthan, V., Keith, B. J., Connolly, B. A. and DeStefano, J. J. (2014). Human immunodeficiency virus reverse transcriptase displays dramatically higher fidelity under physiological magnesium conditions in vitro. J Virol 88(15): 8514-8527.
- Hou, E. W., Prasad, R., Beard, W. A. and Wilson, S. H. (2004). High-level expression and purification of untagged and histidine-tagged HIV-1 reverse transcriptase. Protein Expr Purif 34(1): 75-86.
- Mendelman, L. V., Petruska, J. and Goodman, M. F. (1990). Base mispair extension kinetics. Comparison of DNA polymerase alpha and reverse transcriptase. J Biol Chem 265(4): 2338-2346.
- Menendez-Arias, L. (2009). Mutation rates and intrinsic fidelity of retroviral reverse transcriptases. Viruses 1(3): 1137-1165.
- Rezende, L. F. and Prasad, V. R. (2004). Nucleoside-analog resistance mutations in HIV-1 reverse transcriptase and their influence on polymerase fidelity and viral mutation rates. Int J Biochem Cell Biol 36(9): 1716-1734.
- Svarovskaia, E. S., Cheslock, S. R., Zhang, W. H., Hu, W. S. and Pathak, V. K. (2003). Retroviral mutation rates and reverse transcriptase fidelity. Front Biosci 8: d117-134.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Achuthan, V. and DeStefano, J. J. (2015). Mismatched Primer Extension Assays. Bio-protocol 5(12): e1508. DOI: 10.21769/BioProtoc.1508.
Category
Microbiology > Microbial biochemistry > Protein > Activity
Biochemistry > Protein > Activity
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










