- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Cell Wall Biomass Preparation and Fourier Transform Mid-infrared (FTIR) Spectroscopy to Study Cell Wall Composition
Published: Vol 5, Iss 11, Jun 5, 2015 DOI: 10.21769/BioProtoc.1494 Views: 10471
Reviewed by: Samik BhattacharyaArsalan DaudiAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Enzymatic Starch Quantification in Developing Flower Primordia of Sweet Cherry
Nestor Santolaria [...] Afif Hedhly
Apr 5, 2025 1886 Views
New Approach to Detect and Isolate Rhamnogalacturonan-II in Arabidopsis thaliana Seed Mucilage
Dayan Sanhueza and Susana Saez-Aguayo
Sep 5, 2025 1240 Views
Detailed Method for the Purification of Rhamnogalacturonan-I (RG-I) in Arabidopsis thaliana
Liang Zhang [...] Breeanna R. Urbanowicz
Feb 5, 2026 99 Views
Abstract
Plant cell wall biomass is an abundant and renewable organic resource. Of the polymers it encloses, cellulose and hemicellulose are regarded as a raw material for the production of fuels and other products (Klemm et al., 2005; Slavov et al., 2013). Nonetheless, current usage of lignocellulosic biomass is still below its full potential due to a series of limiting factors mainly related to the cell wall recalcitrance to saccharification, a severe constraint to maximum biomass usability in downstream processing (Pauly and Keegstra, 2008).
As a strategy to optimise bio-energy and bio-refining applications, an increasing amount of effort is being put into the advancement of our knowledge concerning the cell wall compositional roots of recalcitrance. Fourier transform mid-infrared spectroscopy (FTIR) represents a very useful tool on this enterprise, as it allows for a high-throughput, non-destructive and low unit cost procedure for the examination of cell wall biomass (Allison et al., 2009; Carpita and McCann, 2015). Furthermore, the use of Attenuated Total Reflection (ATR) in conjunction with infrared spectroscopy (IR) enables cell wall biomass samples to be examined in solid state without extensive preparation. Nonetheless, the analysis of purified cell wall preparations instead of the intact plant biomass is highly recommended, as it minimises or even eradicates interference from biomass components which are not part of the cell wall. Further information regarding the fundamentals of FTIR may be found elsewhere (Smith, 2011).
Datasets generated from FTIR spectroscopy can be extensive and complex. In these situations, data-driven modelling techniques are often used as exploratory approaches to identify the most distinctive features of the collected spectra. Here we suggest the use of Principal Component Analysis (PCA), a frequently employed method to transform a large set of variables into a smaller set of new variables (principal components), effectively reducing dataset dimensionality.
When the aim is a complete and detailed biomass characterisation, the FTIR-PCA method here described does not exclude the need for parallel wet gravimetric and analytical procedures. However, it does lead to a rapid identification of the major compositional shifts across large sets of samples; thus contributing to steer research pathways, minimise time-draining analytical procedures and reduce overall research costs.
Materials and Reagents
- Lignocellulosic biomass
Note: Depending on the aims of the researcher, lignocellulosic biomass from different species, organs or tissues may be used, providing it is conveniently prepared as indicated in the Procedure section below. For an example of the application of this protocol, please refer to da Costa et al. (2014). - Deionised H2O
- 70% (v/v) aqueous ethanol (molecular biology grade)
- Chloroform/methanol (1:1 v/v) (molecular biology grade)
- 100% acetone (molecular biology grade)
- Type-I porcine α-amylase (Sigma-Aldrich, catalog number: A6255 ) (saline suspension; 29 mg protein/ml; 1714 units/mg protein)
- 0.1 M sodium acetate buffer (see Recipes)
- 0.1 M ammonium formate buffer (see Recipes)
- 0.001 M sodium azide (see Recipes)
Equipment
- Laboratory safety equipment (gloves, eye protection and safety mask are recommended particularly during sodium azide handling)
- Freezer
- pH meter
- Freeze dryer (Edwards Pirani 501 Super Modulyo, Edwards Ltd.)
- Analytical mill (Ika, A11 basic)
- Sieves (pore sizes: 0.18 and 0.85 mm)
- Plastic centrifuge tubes (50 ml, with screw cap) (Greiner Bio-One GmbH)
- Vortex mixer
- Shaking incubator
- Centrifuge
- Fume hood
- Block heater
- FTIR spectrometer (Equinox 55, Bruker Optik) equipped with a Golden Gate ATR accessory (Specac)
Software
- Bruker OPUS IR spectroscopy software (version 5.0; Bruker Optik)
Procedure
- Biomass preparation-organic solvent wash
- Pre-freeze and freeze-dry lignocellulosic biomass (time may vary depending on sample source (for whole Miscanthus spp. tillers the samples were freeze-dried for an excess of 7 days to ensure complete dryness).
- Grind tissues to a particle size in the range 0.18-0.85 mm (suggested equipment: IKA A11 Handheld Analytical Mill; sieves with mesh sizes of 80 and 20 µm).
- Weigh approximately 1 g of the ground plant biomass into a 50 ml plastic centrifuge tube.
- Add 30 ml 70% (v/v) aqueous ethanol, mix thoroughly using a vortex mixer and leave in a shaking incubator set at 40 °C/150 rpm for 12 h.
- Centrifuge at 900 x g for 10 min and discard the supernatant by decantation or aspiration.
- Add 30 ml 70% (v/v) aqueous ethanol, mix using a vortex mixer, but this time incubate the samples for 30 min at 40 °C/150 rpm.
- Centrifuge at 900 x g for 10 min and discard the supernatant.
- Repeat steps A6-7.
- Add 20 ml of the chloroform/methanol (1:1 v/v) solution, mix to re-suspend the pellet and leave in a shaking incubator for 30 min at 25 °C and 150 rpm.
- Centrifuge at 900 x g for 10 min and discard the supernatant after the chloroform/methanol wash.
- Repeat steps A9-10 twice.
- Add 15 ml of acetone, mix to re-suspend the pellet and leave in a shaking incubator for 30 min at 25 °C and 150 rpm.
- Centrifuge at 900 x g for 10 min and discard the supernatant after the acetone wash.
- Repeat steps A12-13 twice.
- Let the organic-washed biomass samples dry overnight in a fume hood (alternatively they can be left in an oven set at 35 °C for approximately 16 h).
- Pre-freeze and freeze-dry lignocellulosic biomass (time may vary depending on sample source (for whole Miscanthus spp. tillers the samples were freeze-dried for an excess of 7 days to ensure complete dryness).
- Biomass preparation-starch removal
- Re-suspend the dry, organic solvent-extracted biomass in 15 ml of 0.1 M sodium acetate buffer (pH 5.0).
- Heat for 20 min at 80 °C in a heating block to induce starch gelatinisation.
- Cool suspensions on ice for 15 min.
- Centrifuge at 900 x g for 10 min and discard the supernatant.
- Wash the pellet twice with 30 ml H2O, with centrifugation (900 x g for 10 min) and supernatant removal after each wash.
- Add to the pellet the following reagents: 10 ml 0.1 M ammonium formate buffer (pH 6.0), 10 μl type-I porcine α-amylase (47 units per 100 mg cell wall) and 500 µl 0.001 M sodium azide solution.
- Leave in a shaking incubator for 48 h at 25 °C/110 rpm.
- Terminate the digestion by heating to 95 °C/15 min.
- Cool samples on ice for 15 min.
- Centrifuge at 900 x g for 10 min and discard the supernatant.
- Wash the pellet three times with 30 ml H2O and twice with 20 ml acetone, with centrifugation (900 x g for 10 min) and supernatant removal after each wash.
- Let the prepared cell wall material samples dry overnight in a fume hood (alternatively they can be left in an oven set at 35 °C for approximately 16 h; the samples are stable for months at room temperature if kept in a sealed container and protected from direct light).
- Re-suspend the dry, organic solvent-extracted biomass in 15 ml of 0.1 M sodium acetate buffer (pH 5.0).
- Biomass examination-Fourier transform mid-infrared spectroscopy
- Without further preparation, place approximately 10 mg of the dry cell wall powder onto the Golden Gate ATR crystal.
- Press the sample into optimal contact with the ATR crystal using the anvil of the Golden Gate ATR accessory.
- Collect spectra in duplicate for each biomass sample
Note: Each spectrum is collected by ATR in a mid-infrared range of 4,000-600 cm-1 and consists of the average over 32 scans at a resolution of 4 cm-1. Spectra were corrected for background absorbance by subtraction of the spectrum of the empty ATR crystal. - Recover the sample using a spatula or a razor blade (Video 1).
- Clean the contact areas with a laboratory tissue wipe and ethanol between samples.
- Repeat steps C1-5 for all samples.
- Convert each collected spectrum into an individual text file containing the spectral two dimensional Cartesian coordinates (x, y) (Figure 1) in two separate columns (we used the Bruker OPUS IR spectroscopy software). Subsequently, matrices containing the raw data may be created and the underlying relationships between the spectra may be investigated using numerical or statistical computing software (such as R or MATLAB). As an example, we used PCA to identify the features most distinctive of spectra collected from Miscanthus spp. leaf and stem cell wall biomass (Figure 2).
Video 1. Collection of FTIR spectra from cell wall biomass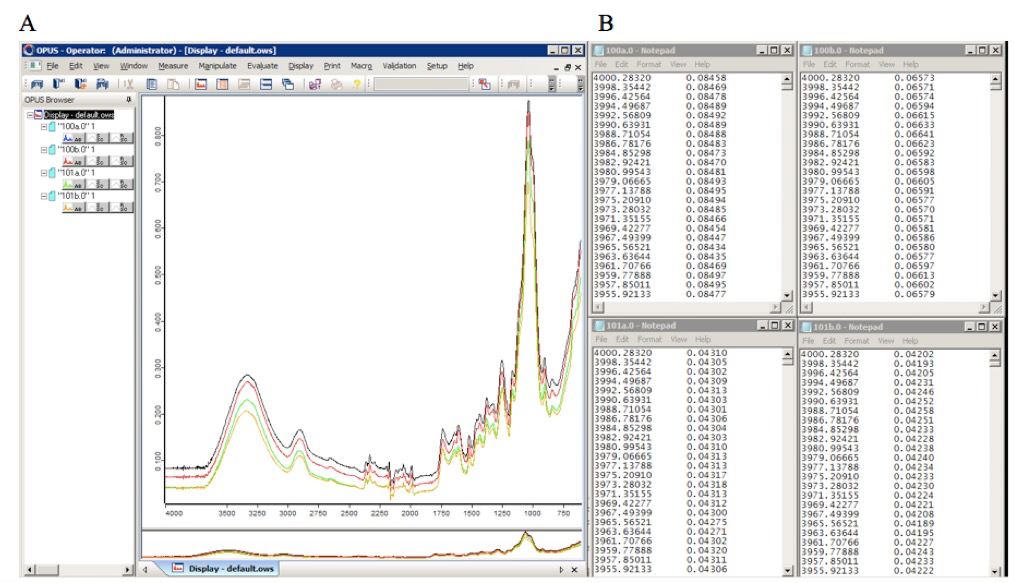
Figure 1. FTIR spectra of cell wall biomass (Miscanthus spp.), plotted using Opus IR spectroscopy software (A), and after being converted into text files containing their Cartesian coordinates in two separate columns (B)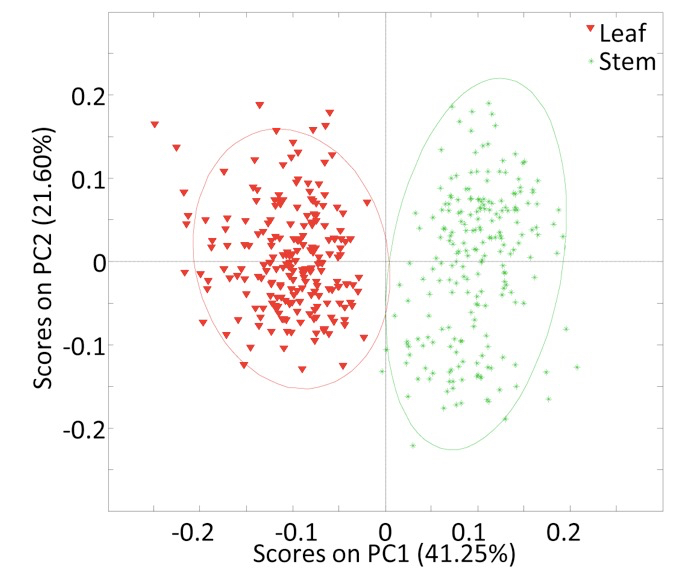
Figure 2. Plot of principal component one (PC1) and principal component two (PC2) scores for Miscanthus spp. leaf and stem cell wall spectra generated by FTIR (da Costa et al., 2014)
- Without further preparation, place approximately 10 mg of the dry cell wall powder onto the Golden Gate ATR crystal.
Recipes
- 0.1 M sodium acetate buffer (pH 5.0)
Prepare 0.2 M acetic acid (A) by mixing 11.55 ml glacial acetic acid in 500 ml H2O and adjusting to 1 L with H2O
Prepare 0.2 M sodium acetate solution (B) by dissolving 27.21 g sodium acetate trihydrate in 800 ml H2O and adjust to 1 L with H2O
Mix 14.8 ml of A, 35.2 ml of B and 50 ml of H2O
Confirm pH=5.0 with a pH meter (if needed, the pH may be adjusted with 10 M sodium acetate or with glacial acetic acid) - 0.1 M ammonium formate buffer (pH 6.0)
Dissolve 6,306 mg ammonium formate in 500 ml H2O and adjust pH to 6.0 with formic acid
Make up to 1 L with H2O
Confirm pH=6.0 with a pH meter (if needed, the pH may be adjusted with formic acid) - 0.001 M sodium azide
Dissolve 10 mg sodium azide in 145 ml H2O
Acknowledgments
The methods here described were employed in da Costa et al. (2014) for the analysis of Miscanthus spp. cell wall samples. Portions of this procedure were modified from various sources: organic solvent washing and starch gelatinization from Foster et al. (2010); starch removal from Persson et al. (2007) and Kong et al. (2011); and Fourier transform mid-infrared spectroscopy from Allison et al. (2009). The original work was supported by the European Regional Development Funding through the Welsh Government for BEACON Grant number 8056 to R. M. F. da Costa, G. G. Allison and M. Bosch.
References
- Allison, G. G., Thain, S. C., Morris, P., Morris, C., Hawkins, S., Hauck, B., Barraclough, T., Yates, N., Shield, I., Bridgwater, A. V. and Donnison, I. S. (2009). Quantification of hydroxycinnamic acids and lignin in perennial forage and energy grasses by Fourier-transform infrared spectroscopy and partial least squares regression. Bioresour Technol 100(3): 1252-1261.
- Carpita, N. C. and McCann, M. C. (2015). Characterizing visible and invisible cell wall mutant phenotypes. J Exp Bot. (Epub ahead of print)
- da Costa, R. M., Lee, S. J., Allison, G. G., Hazen, S. P., Winters, A. and Bosch, M. (2014). Genotype, development and tissue-derived variation of cell-wall properties in the lignocellulosic energy crop Miscanthus. Ann Bot 114(6): 1265-1277.
- Foster, C. E., Martin, T. M. and Pauly, M. (2010). Comprehensive compositional analysis of plant cell walls (Lignocellulosic biomass) part I: lignin. J Vis Exp (37): e1745.
- Klemm, D., Heublein, B., Fink, H. P. and Bohn, A. (2005). Cellulose: fascinating biopolymer and sustainable raw material. Angew Chem Int Ed Engl 44(22): 3358-3393.
- Kong, Y., Zhou, G., Yin, Y., Xu, Y., Pattathil, S. and Hahn, M. G. (2011). Molecular analysis of a family of Arabidopsis genes related to galacturonosyltransferases. Plant Physiol 155(4): 1791-1805.
- Pauly, M. and Keegstra, K. (2008). Cell‐wall carbohydrates and their modification as a resource for biofuels. Plant J 54(4): 559-568.
- Persson, S., Caffall, K. H., Freshour, G., Hilley, M. T., Bauer, S., Poindexter, P., Hahn, M. G., Mohnen, D. and Somerville, C. (2007). The Arabidopsis irregular xylem8 mutant is deficient in glucuronoxylan and homogalacturonan, which are essential for secondary cell wall integrity. Plant Cell 19(1): 237-255.
- Slavov, G., Allison, G. and Bosch, M. (2013). Advances in the genetic dissection of plant cell walls: tools and resources available in Miscanthus. Front Plant Sci 4: 217.
- Smith, B. (2011). Fundamentals of Fourier Transform Infrared Spectroscopy, Second Edition. CRC Press.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
da Costa, R. M. F., Allison, G. G. and Bosch, M. (2015). Cell Wall Biomass Preparation and Fourier Transform Mid-infrared (FTIR) Spectroscopy to Study Cell Wall Composition. Bio-protocol 5(11): e1494. DOI: 10.21769/BioProtoc.1494.
Category
Plant Science > Plant biochemistry > Carbohydrate
Biochemistry > Carbohydrate > Polysaccharide
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










