- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Extraction and Identification of T Cell Stimulatory Self-lipid Antigens
Published: Vol 5, Iss 11, Jun 5, 2015 DOI: 10.21769/BioProtoc.1491 Views: 10363
Reviewed by: Savita NairRamalingam BethunaickanAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Generation of T cells from Human and Nonhuman Primate Pluripotent Stem Cells
Akhilesh Kumar [...] Igor I. Slukvin
Jul 5, 2020 7275 Views

Expansion and Polarization of Human γδT17 Cells in vitro from Peripheral Blood Mononuclear Cells
Xu Chen [...] Jun Yan
Jan 5, 2024 1949 Views
Proliferation Assay Using Cryopreserved Porcine Peripheral Mononuclear Cells Stimulated With Concanavalin A and Analyzed With FCS ExpressTM 7.18 Software
Marlene Bravo-Parra [...] Luis G. Giménez-Lirola
Jun 5, 2025 2618 Views
Abstract
Autoreactive T cells restricted to CD1 molecules and specific for endogenous lipids are abundant in human blood (de Jong et al., 2010; de Lalla et al., 2011). A few self-lipid molecules recognized by diverse individual T cell clones and accumulated within APCs following stress signals or cell transformation have been identified so far (de Jong et al., 2010; Chang et al., 2008; Lepore et al., 2014). These findings suggested that auto-reactive CD1-restricted T cells display broad lipid specificities and may play critical roles in different types of immune responses including cancer immune surveillance, autoimmunity and antimicrobial immunity. Therefore, the identification of the repertoire of self-lipid molecules recognized by T cells is important to study the physiologic functions of this T cell population and to assess their therapeutic potential (Lepore et al., 2014). Here we describe the protocol we established to isolate and identify endogenous lipids derived from leukemia cells, which stimulate specific autoreactive CD1c-restricted T lymphocytes (Lepore et al., 2014). This protocol can be applied to isolate lipid antigens from any type of target cells and to investigate the self-lipid antigen specificity of autoreactive T cells restricted to all CD1 isoforms (Facciotti et al., 2012).
Keywords: Self-lipid antigensMaterials and Reagents
- Autoreactive CD1-restricted T cell clones (generated as described in de Lalla et al. 2011)
- THP-1 cells (ATCC TIB-202TM, or other target cells able to stimulate the T cell clones in a CD1-dependent manner and in the absence of exogenously provided antigens)
- Antigen presenting cells (APCs) expressing relevant CD1 isoforms and per se poorly stimulating autoreactive T cell
- RPMI 1640 (Amimed, catalog number: 1-41F01-I )
- Stable glutamine (Amimed, catalog number: 5-10K50-H )
- Sodium pyruvate (Amimed, catalog number: 5-60F00-H )
- Non essential amino acids (Amimed, catalog number: 5-13K00-H )
- Kanamycin (Amimed, catalog number: 4-08F00-H )
- Fetal bovine serum (Lonza, catalog number: DE14-802F )
- PBS without Ca2+ and Mg2+ (Amimed, catalog number: 3-05F29-I )
- ELISA
- MAb pairs: Purified anti-human GM-CSF (BioLegend, catalog number: 502202 ), biotin-conjugated anti-human GM-CSF (BioLegend, catalog number: 502304 ); purified anti-human IFN-γ (BioLegend, catalog number: 50750 ), biotin-conjugated anti-IFN-γ (BioLegend, catalog number: 502504 )
- HRP-streptavidin (BioLegend, catalog number: 405210 )
- OPD SIGMAFAST (Sigma-Aldrich, catalog number: P9187-50SET )
- Cytokine standards: Recombinant human GM-CSF (BioLegend, catalog number: 572409 ), recombinant human IFN-γ (BioLegend, catalog number: 570209 )
- MAb pairs: Purified anti-human GM-CSF (BioLegend, catalog number: 502202 ), biotin-conjugated anti-human GM-CSF (BioLegend, catalog number: 502304 ); purified anti-human IFN-γ (BioLegend, catalog number: 50750 ), biotin-conjugated anti-IFN-γ (BioLegend, catalog number: 502504 )
- Anti-CD1 blocking antibodies (anti-CD1c mAb) (Abcam, catalog number: ab18216-100 )
- Anti-human CD19 mAbs (Miltenyi Biotec, catalog number: 130-097-055 )
- Methanol (Applichem, catalog number: A0688, 2500PE )
- Chloroform (Applichem, catalog number: A1585, 1000 )
- Ethyl acetate (Merck KGaA, catalog number: 1.00868.1000 )
- 1-Butanol (Sigma-Aldrich, catalog number: 34867 )
- Diisopropyl ether (Sigma-Aldrich, catalog number: 3827-IL-F )
- Isopropanol (Applichem, catalog number: A1592.2500 )
- Acetic acid (Applichem, catalog number: A2354.0500 )
- Acetone (Applichem, catalog number: A1567.2500 )
- Acetonitrile (Riedel-De Haen, catalog number: 34967 )
- Hexane (Sigma-Aldrich, catalog number: 34994 )
- HCl 37% fuming (Merck KGaA, catalog number: 317.1000 )
- Formic acid (Merck KGaA, catalog number: 1.11670.1000 )
- Ammonium acetate (Sigma-Aldrich, catalog number: A1542 )
- H2O (Sigma-Aldrich, catalog number: 95304 )
- Water-saturated butanol (see Recipes)
- Elution solutions for lipid fractionation on amino-cartridge (see Recipes)
- Complete medium (see Recipes)
Equipment
- Aminopropyl cartridges (SEP-PAK Vac 6 cc, 500 mg NH2 cartridges) (Waters Corporation, catalog number: WAT200606)
- HPLC system (Jasco)
- Nucleodur C18 Pyramid end-capped column (3-μm particle size, 3-mm ID, 125-mm length) (Macherey-Nagel, catalog number: N9040986 )
- Automated fraction collector (Gilson, catalog number: FC203B )
- Glass conical tubes (30 ml and 1 ml volumes, Glass Keller)
- Glass pipettes (Pirex)
- 96 wells flat bottom culture plates (BD Biosciences, Falcon®, catalog number: 353075 )
- 96 wells ELISA immune-plates (Maxisorp, Nunc, catalog number: 439454 )
- Humidified CO2 cell culture incubator (Heraeus, Hera cell 150)
- Spectrophotometer/ELISA Reader (Synergy H1 Hybrid Reader, BioTek Instruments)
- Sonicator (Sonics, Vibra Cell)
- Rotating wheel (Labinco BV, catalog number: 76000 )
- Manometer-regulated N2 gas tank (Carba gas)
Procedure
- Lipid extraction and fractionation
The following lipid extraction and fractionation procedure is adapted from (Facciotti et al., 2012; Folch et al., 1957) and it is optimized for 109 cells. For different numbers of cells, adjust the volumes accordingly. This protocol was used to extract lipids from mouse thymocytes or THP-1 cells (ATCC TIB-202TM). For other types of cells the protocol may require optimization.
Lipid extraction- Pellet the cells by centrifugation at room temperature for 5 min at 300 x g, resuspend them in 10 ml PBS and transfer them in a glass tube.
After this step use glass tubes exclusively. - Wash cells 2x with 10 ml PBS by centrifugation (5 min at 300 x g) and completely remove the PBS by aspiration with a glass Pasteur pipette.
- Resuspend the pellet in 8 ml of a mixture of water/methanol/chloroform (1:1:2 vol/vol/vol).
- Sonicate 2 x 30 sec (5 Hz).
- Incubate 3 h at room temperature in a rotating wheel.
- Centrifuge 5 min at 3,100 x g at room temperature.
- Collect the organic layer (bottom layer) with a glass pipet avoiding contamination with the aqueous phase (upper layer) and store at -20 °C.
- Add 4 ml of methanol/chloroform (1:2 vol/vol) to the remaining aqueous phase.
- Vortex 2 min and incubate 1 h at room temperature in rotation.
- Centrifuge 5 min at 3,100 x g.
- Collect the organic (bottom) layer with a glass pipet avoiding contamination with the aqueous phase (upper layer) and store at -20 °C until use.
- Repeat steps A8-11 one additional time.
- Pool the collected organic phases and dry for ~2 h under nitrogen flow delivered as a gas stream (~3 bars) through a Pasteur pipette connected to a N2 source. Dissolve in 2 ml methanol/chloroform (1:2 vol/vol) and store at -20 °C. We refer to this extraction as “apolar” because it contains most of the cellular lipids with exception of the highly polar ones.
- Measure the volume of the remaining aqueous phase and add 1 vol of water-saturated butanol (see Recipes).
- Vortex 2 min and incubate 1 h at room temperature in a rotating wheel.
- Centrifuge for 5 min at 3,100 x g.
- Collect the upper layer (butanol phase), dry under nitrogen flow, dissolve in 2 ml methanol and store at -20 °C. As this extraction contains highly polar lipids poorly soluble in methanol/chloroform we call it “polar”.
Lipid fractionation
The following fractionation procedure allows the separation of both apolar and polar lipid extractions in 10 individual fractions of different polarity using aminopropyl cartridges. Apply the same procedure described here to both the lipid extractions separately using two different cartridges.- Equilibrate the cartridge with 6 ml hexane.
- Load the cartridge with the total volume of extracted lipids (2 ml, Figure 1 A).
- Sequentially apply the indicated volumes of the eluents (see Recipes) and collect the fractions individually eluted by gravity in glass conical tubes (Figure 1B; each letter represents an eluent and it also identifies individual fractions, e.g. the fraction eluted with solution “a” is called “a”, etc.):
- 4 ml
- 4 ml
- 3 ml
- 11 ml
- 9 ml
- 3 ml
- 4.5 ml
- 6 ml
- 4 ml
- 4 ml
- 4 ml
- Dry eluted fractions under nitrogen flow and dissolve each fraction in the solutions and volumes indicated below:
- 280 µl methanol/chloroform (1:2 vol/vol)
- 280 µl methanol/chloroform (1:1 vol/vol)
- 280 µl methanol/chloroform (1:1 vol/vol)
- 280 µl methanol/chloroform (1:1 vol/vol)
- 280 µl methanol/chloroform (1:1 vol/vol)
- 280 µl methanol/chloroform (9:1 vol/vol)
- 280 µl methanol/chloroform (9:1 vol/vol)
- 280 µl methanol/chloroform (9:1 vol/vol)
- 280 µl methanol/chloroform (9:1 vol/vol)
- 500 µl methanol/chloroform (9:1 vol/vol)
- 280 µl methanol/chloroform (1:2 vol/vol)
- Store each resuspended fraction at -20 °C.
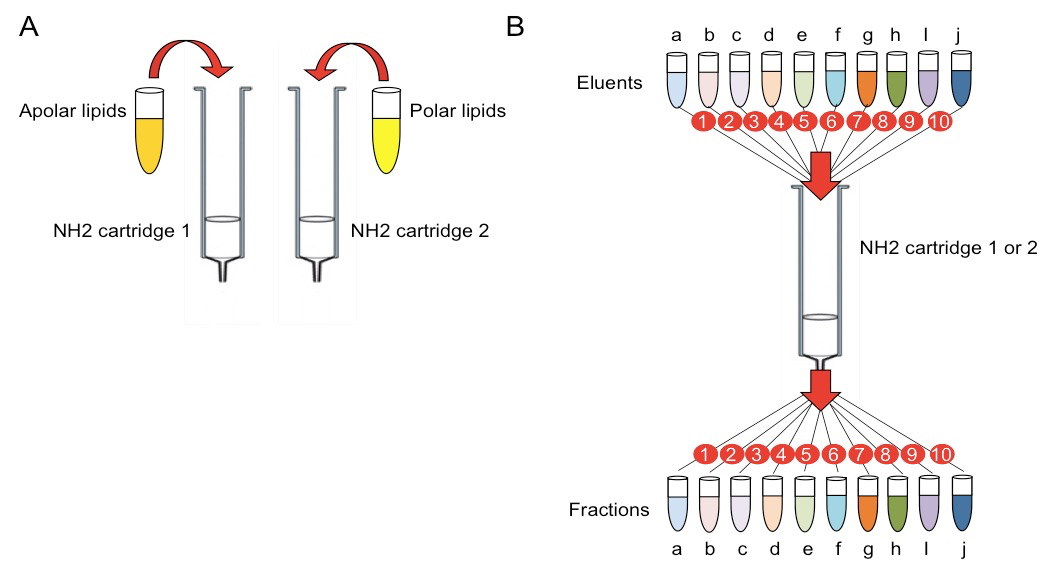
Figure 1. Scheme illustrating the lipid fractionation procedure. A. Loading of apolar and polar extracted lipids on two separate amino (NH2) cartridges. B. Sequential elution of lipid fractions from the amino cartridges. Letters indicate eluents and corresponding fractions. Numbers indicate the temporal order of elution and fraction collection.
- Pellet the cells by centrifugation at room temperature for 5 min at 300 x g, resuspend them in 10 ml PBS and transfer them in a glass tube.
- T cell activation assay
In this section we describe how to evaluate the T cell stimulatory capacity of lipid preparations. To maximize assay sensitivity, a series of essential points have to be considered.- Autoreactive T cell clones recognizing the cells from which the lipids are extracted, should be highly responsive.
- APCs should express high levels of the restriction molecules for optimal antigen presentation.
- APCs should be chosen for their poorly stimulatory capacity in the absence of exogenously added antigen, in order to minimize the levels of background stimulation of autoreactive T cells in the absence of tumor-derived lipids.
- Plate 1-5 x 105 APCs/well in 96 wells flat bottom plates in 45 µl of complete medium.
- Prepare at least 3 dilutions of each lipid fraction. Tested dilutions should be in the range 1:10-1:1,000 of the original preparation. As the fractions are dissolved in organic solvents, which are toxic for cells (volumes and type of solvents are indicated in the step B4 of the paragraph “Lipid fractionation”), they need to be dried (under nitrogen, to avoid oxygen-induced lipid alterations) and offered to the cells in a solution, which is compatible with cell viability. Generally, 20 µl of each fraction are transferred into a new glass conical tube, dried, dissolved in 20 µl of PBS 20% methanol (vol/vol, vehicle) and sonicated. Dissolved fractions are then used to pulse APCs at various dilutions.
- Add fractions to plated APCs. As example, 5 µl of each dissolved fraction (undiluted or previously diluted 1:10 or 1:100 using PBS 20% methanol) are added to the wells containing APCs in 45 µl of complete medium. In this way each lipid fraction will be finally diluted 1:10, 1:100 or 1:1000, respectively in the culture wells. Importantly, the methanol contained in the vehicle in which the fractions are dissolved will also be diluted to a final concentration of 2%, which is compatible with APC viability and T cell activation. Include control wells in which only the vehicle (5 µl of PBS 20% methanol) is added to the 45 µl of APCs. Perform triplicate replicas of each experimental condition.
- Incubate 4 h at 37 °C in humidified CO2 incubator.
- Add 5 x 105 T cells/well in 50 µl of complete medium.
- Incubate 24-48 h at 37 °C in humidified CO2 incubator.
- Collect supernatants and measure cytokine release by standard sandwich ELISA (according to the manufacturing protocols; see Materials and Reagents). We measured release of GM-CSF (after 24 h) and IFN-γ (after 48 h) as read out of T cell activation. However, it is important to note that the choice of the cytokine needs to be done according to the cytokines more abundantly released by tested T cells and excluding those released by the APCs, which should be determined in preliminary experiments. In general, GM-CSF release represents a very sensitive and relatively fast read out for in vitro T cell clone activation. Measuring at least two different cytokines is recommendable to avoid false positive results. Lipid fractions are considered positive if they induce dose-dependent T cell cytokine release. Results are expressed as fold change over background (cytokine release in the presence of APCs incubated with lipid/cytokine release with APCs incubated with vehicle). Fold change ≥ 2 is considered relevant.
- The activity of stimulatory fraction(s) has to be confirmed in a second set of experiments, in which blocking antibodies specific for the CD1 isoform that restricts the response of the T cells used are included. Blocking antibodies and appropriate isotype control antibodies are used at a final concentration of 20 µg/ml and are added to APCs incubated with active lipid fractions at least 30 min before T cells.
- Autoreactive T cell clones recognizing the cells from which the lipids are extracted, should be highly responsive.
- Sub-fractionation of active lipid fractions
Once one or more T cell stimulatory lipid fractions are identified a second fractionation step is made to further purify the antigenic lipid molecules.
The procedure described in this section uses reverse-phase HPLC performed with a Nucleodur C18 Pyramid end-capped column (3-μm particle size, 3-mm ID) and two mobile phases:
Mobile phase A: methanol/water/formic acid (74:25:1, vol/vol/vol) (pH 4.0)
Mobile phase B: methanol/formic acid (99:1, vol/vol) (pH 4.0)
The following gradient was used to isolate polar lipids:- B 20% from time 0 to min 1
- B from 20% to 50% from min 1 to min 2
- B 50% from min 2 to min 4
- B from 50% to 100% to min 4 to min 34
- B 100% from min 34 to min 54
- B from 100% to 20% from min 54 to min 55
- B 20% from min 55 to min 60
- Inject the active lipid fraction in the HPLC system.
- Collect individual sub-fractions every 30 sec using an automated collector connected to the HPLC system.
- Dry the sub-fractions under nitrogen flow and resuspend fractionated lipids in 10 µl methanol.
- Perform T cell activation assay as described above by testing all individual sub-fractions at various dilutions.
- Store the rest of the sub-fractions at -20 °C for further analyses.
- B 20% from time 0 to min 1
Representative data
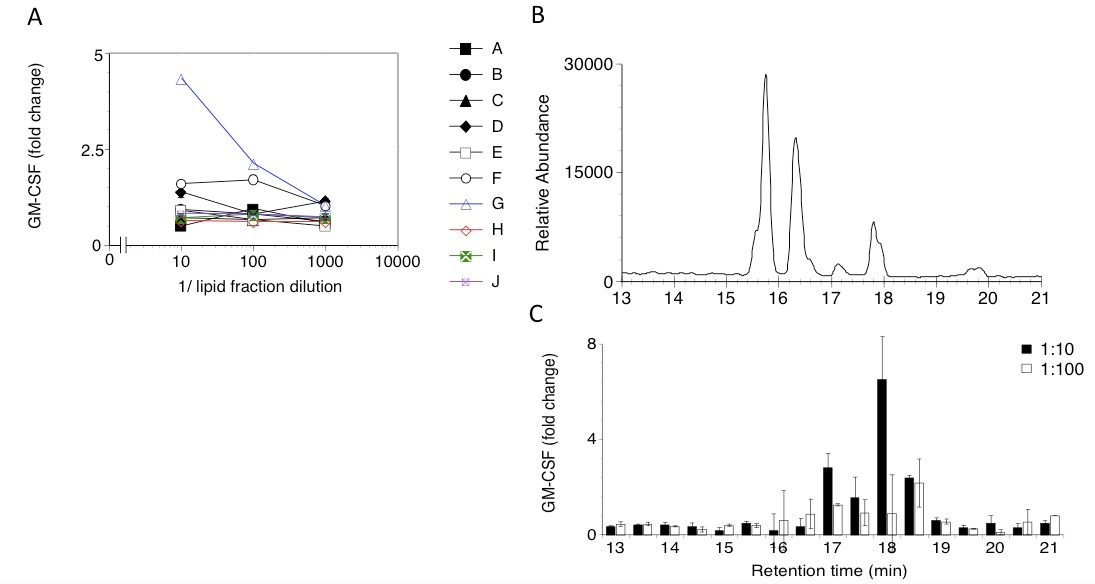
Figure 2. A. Response of a CD1-restricted human T cell clone to lipid fractions (A-J) extracted from THP-1 cells. Three dilutions of each fraction were tested. GM-CSF released in the supernatant was measured by ELISA and expressed as fold change over background. B. HPLC profile corresponding to the sub-fractionation of the T cell stimulatory lipid fraction G in panel A. Sub-fractions were collected every 30 sec. C. GM-CSF release by the CD1-restricted T cell clone to HPLC sub-fractions of fraction G. Each sub-fraction was tested at two dilutions (1:10, filled columns, and 1:100, open columns). Bars indicate sd.
For more representative data also refer to the following: T cell activation assay, Figure 3 (Lepore et al., 2014); Lipid sub-fractionation, Figure 1 (Facciotti et al., 2012) and Figure 2 (Lepore et al., 2014).
Recipes
- Water-saturated butanol
1-Butanol/H2O 1:1, vol/vol
Vigorously shake 5 min, let stand for at least 30 min to allow phase separation. Use the saturated butanol (upper phase). - Eluents for lipid fractionation on amino-cartridges
- Ethyl acetate/hexane 15:85, vol/vol
- Chloroform/methanol 23:1 vol/vol
- Diisopropyl ether/ acetic acid 98:5, vol/vol
- Acetone/methanol 9:1.35, vol/vol
- Chloroform/methanol 2:1, vol/vol
- Methanol
- Isopropanol/3 N HCl in methanol 4:1, vol/vol
- Methanol/3 N HCl in methanol 9:1, vol/vol
- Chloroform/methanol 2:1, vol/vol
- Chloroform/methanol/3.6 M ammonium acetate in water 30/60/8, vol/vol/vol
- Ethyl acetate/hexane 15:85, vol/vol
- Complete medium
RPMI 1640
2 mM stable glutamine
1 mM sodium pyruvate
1 mM non-essential amino acids
50 µg/ml kanamycin
10% heat-inactivated fetal bovine serum
Acknowledgments
This work was supported by Grants of the Swiss National Science Foundation (NMS1813), A*STAR/Australian NHMRC (1201826277) and University of Basel (DMS2306). The protocols described here were used in (Facciotti et al., 2012) and (Lepore et al., 2014).
References
- Bodennec, J., Koul, O., Aguado, I., Brichon, G., Zwingelstein, G. and Portoukalian, J. (2000). A procedure for fractionation of sphingolipid classes by solid-phase extraction on aminopropyl cartridges. J Lipid Res 41(9): 1524-1531.
- Chang, D. H., Deng, H., Matthews, P., Krasovsky, J., Ragupathi, G., Spisek, R., Mazumder, A., Vesole, D. H., Jagannath, S. and Dhodapkar, M. V. (2008). Inflammation-associated lysophospholipids as ligands for CD1d-restricted T cells in human cancer. Blood 112(4): 1308-1316.
- de Jong, A., Pena-Cruz, V., Cheng, T. Y., Clark, R. A., Van Rhijn, I. and Moody, D. B. (2010). CD1a-autoreactive T cells are a normal component of the human alphabeta T cell repertoire. Nat Immunol 11(12): 1102-1109.
- de Lalla, C., Lepore, M., Piccolo, F. M., Rinaldi, A., Scelfo, A., Garavaglia, C., Mori, L., De Libero, G., Dellabona, P. and Casorati, G. (2011). High-frequency and adaptive-like dynamics of human CD1 self-reactive T cells. Eur J Immunol 41(3): 602-610.
- De Libero, G., Moran, A. P., Gober, H. J., Rossy, E., Shamshiev, A., Chelnokova, O., Mazorra, Z., Vendetti, S., Sacchi, A., Prendergast, M. M., Sansano, S., Tonevitsky, A., Landmann, R. and Mori, L. (2005). Bacterial infections promote T cell recognition of self-glycolipids. Immunity 22(6): 763-772.
- Facciotti, F., Ramanjaneyulu, G. S., Lepore, M., Sansano, S., Cavallari, M., Kistowska, M., Forss-Petter, S., Ni, G., Colone, A., Singhal, A., Berger, J., Xia, C., Mori, L. and De Libero, G. (2012). Peroxisome-derived lipids are self antigens that stimulate invariant natural killer T cells in the thymus. Nat Immunol 13(5): 474-480.
- Folch, J., Lees, M. and Sloane Stanley, G. H. (1957). A simple method for the isolation and purification of total lipides from animal tissues. J Biol Chem 226(1): 497-509.
- Lepore, M., de Lalla, C., Gundimeda, S. R., Gsellinger, H., Consonni, M., Garavaglia, C., Sansano, S., Piccolo, F., Scelfo, A., Haussinger, D., Montagna, D., Locatelli, F., Bonini, C., Bondanza, A., Forcina, A., Li, Z., Ni, G., Ciceri, F., Jeno, P., Xia, C., Mori, L., Dellabona, P., Casorati, G. and De Libero, G. (2014). A novel self-lipid antigen targets human T cells against CD1c(+) leukemias. J Exp Med 211(7): 1363-1377.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Lepore, M., Sansano, S., Lalla, C. D., Dellabona, P., Casorati, G., Libero, G. D. and Mori, L. (2015). Extraction and Identification of T Cell Stimulatory Self-lipid Antigens. Bio-protocol 5(11): e1491. DOI: 10.21769/BioProtoc.1491.
Category
Immunology > Immune cell isolation > Lymphocyte
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link