- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Arabidopsis Metabolome Analysis Using Infusion ESI FT-ICR/MS
Published: Vol 5, Iss 9, May 5, 2015 DOI: 10.21769/BioProtoc.1463 Views: 8971
Reviewed by: Arsalan DaudiSekhar KambakamAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
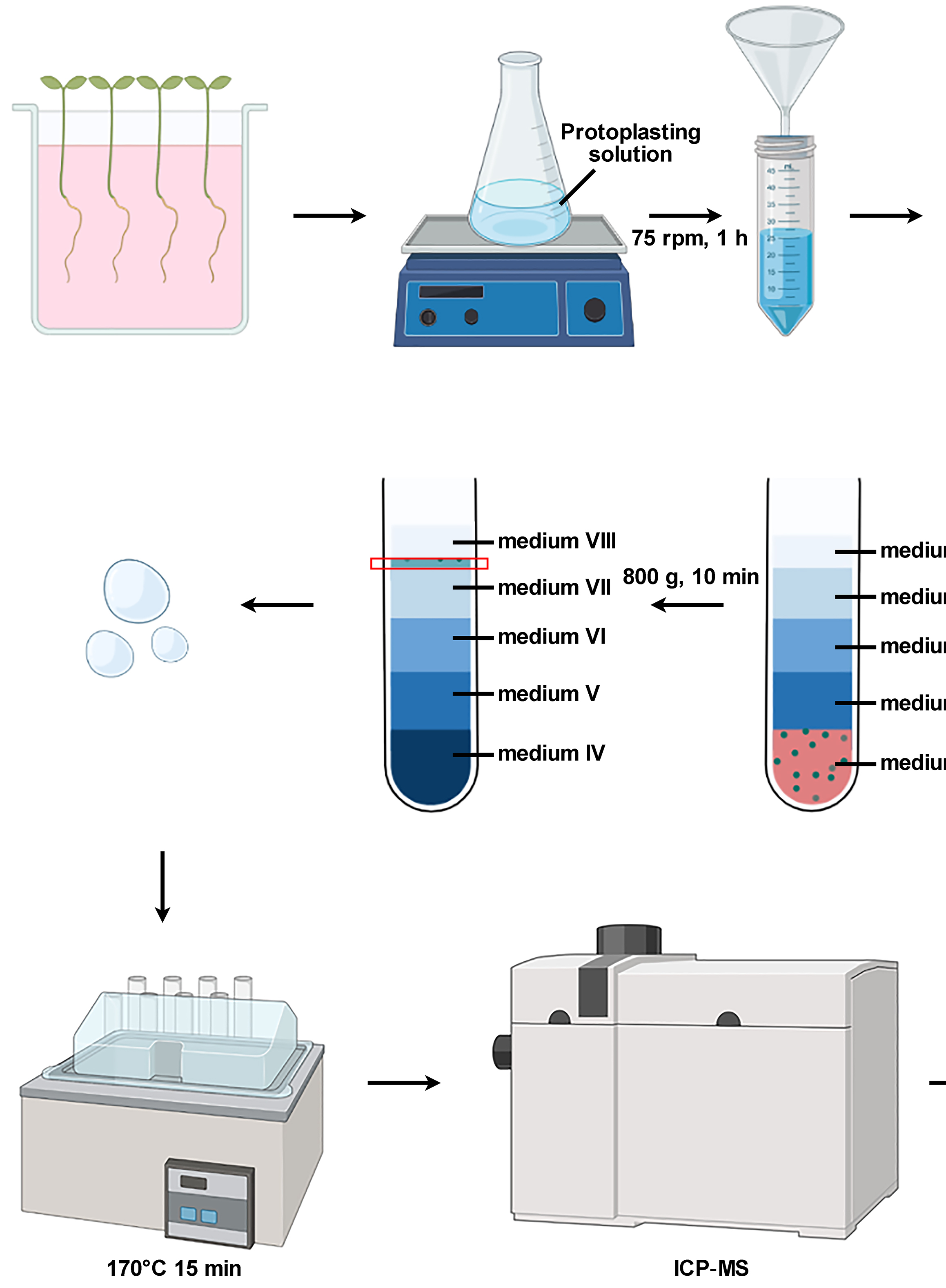
Isolation of Intact Vacuoles from Arabidopsis Root Protoplasts and Elemental Analysis
Chuanfeng Ju [...] Zhenqian Zhang
Mar 5, 2023 2034 Views

Analysis of Modified Plant Metabolites Using Widely Targeted Metabolite Modificomics
Jianing Zhang [...] Jun Yang
Apr 5, 2025 1458 Views
High-Performance Liquid Chromatography Quantification of Glyphosate, Aminomethylphosphonic Acid, and Ascorbate in Culture Medium and Microalgal Cells
Juan Manuel Ostera [...] Gabriela Malanga
Apr 5, 2025 1171 Views
Abstract
We made the method for Arabidopsis metabolome analysis based on direct-infusion Fourier transform ion cyclotron resonance mass spectrometry (FT-ICR/MS) (IonSpec). This method was sufficiently applied to metabolic phenotyping of Arabidopsis. This method is simple in that after homogenizing samples, powdered samples are dissolved in extraction solvents (acetone and methanol) to 20% fresh weight/volume. Extracted sample solutions are dried and dissolved in 50% (v/v) acetonitrile. Mass analysis using FT-ICR/MS (IonSpec) is performed in positive and negative ionization operation modes. Mass spectra are acquired over the 100-1,000 m/z range and accumulated to improve the S/N ratio.
Keywords: MetabolomeMaterials and Reagents
- 3-week-old Arabidopsis plants
- Liquid nitrogen
- Acetone (HPLC grade) (Wako Chemicals USA, catalog number: 014-08681 )
- Methanol (HPLC grade) (Wako Chemicals USA, catalog number: 134-14523 )
- Nitrogen gas
- EN1-16 (TAITEC, catalog number: 0076417-000 )
- Formic acid (HPLC grade) (Wako Chemicals USA, catalog number: 063-04192 )
- Lidocaine (anaesthetic) m/z 235.18104 (Wako Chemicals USA, catalog number: 120-02691 )
- Prochloraz (agricultural chemical) m/z 376.03863 (Wako Chemicals USA, catalog number: 164-25131 )
- Reserpine (alkaloid sedative drug) m/z 609.28121 (Wako Chemicals USA, catalog number: 184-00691 )
- Bombesin (peptide) m/z 810.41479 (Wako Chemicals USA, catalog number: 339-40861 )
- 28~30% ammonia solution (Wako Chemicals USA, catalog number: 016-03146 )
- Negative mode internal standards: 2.4-D (plant hormone) m/z 218.96157 (Wako Chemicals USA, catalog number: 040-18532 )
- Ampicillin (antibiotic) m/z 348.10180 (Wako Chemicals USA, catalog number: 017-20531 , CHAPS 8 detergent) m/z 613.388865 (Wako Chemicals USA, catalog number: 341-04721 )
- Tetra-N-acetylchitotetraose [(GluNAc)4] m/z 829.32023 (Tokyo Chemical Industry, catalog number: T2910 )
- 50% (v/v) acetonitrile (HPLC grade) (Wako Chemicals USA, catalog number: 018-19853 ) [use distilled water to dilute acetonitrile to 50%(v/v)]
- Acetic acid (Wako Chemicals USA, catalog number: 014-20063 )
- 28~30% ammonia solution (Wako Chemicals USA, catalog number: 016-03146 )
- Ultrapure water (recommended but not required)
Equipment
- An IonSpec Explorer FT-ICR/MS equipped with a 7-tesla actively shielded superconducting magnet (Agilent, Ionspec)
- Glass vial and teflon cap (GL Biochem, catalog number: 1030-46716 )
- Glass pipette
- 0.45 µm filters (PTFE) (ADVANTEC®, catalog number: 13HP045AN )
- Heat block
Procedure
- Preparation of samples for FT-ICR/MS analysis
- Homogenize samples with liquid nitrogen to create powder samples of whole plants using mortars and pestles.
- Dissolve in extraction solvents to 20% fresh weight/volume. Two extraction solvents, 100% acetone and 100% methanol, are used to elute various polar compounds.
- Filter extracted sample solutions through 0.45 µm filters (PTFE).
- Transfer filtered sample solutions to vials.
- Transfer 1 ml of sample solution to a separate vial. Place the sample solution into a draft chamber and apply nitrogen gas using EN1-16. At 40 °C solvents will evaporate using a heat block.
- Dissolve samples in 50% acetonitrile (0.1~1 ml depending on sample varieties) and store at -80 °C.
- Dilute samples 1:14 prior to ESI analysis.
- Homogenize samples with liquid nitrogen to create powder samples of whole plants using mortars and pestles.
- FT-ICR/MS analysis
- Perform Mass analysis using an IonSpec Explorer FT-ICR/MS in positive and negative ionization operation modes. Ions are generated from an ESI source with a fused silica needle of 0.005-inch i. d. (Oikawa et al., 2006). Samples are infused using a Harvard syringe pump mode 22 at a flow rate of 0.5 to 1.0 µl/min through a 100 µl Hamilton syringe. Set the potentials on the electrospray emitters to 3.0 kV and -0.3 kV for the positive and the negative electrosprays, respectively.
- For positive mode, add 99.5% formic acid to extracted sample solutions (step A7) at a final concentration of 0.1% (v/v). Positive mode internal standards: Lidocaine (anaesthetic) m/z 235.18104, Prochloraz (agricultural chemical) m/z 376.03863, Reserpine (alkaloid sedative drug) m/z 609.28121, Bombesin (peptide) m/z 810.41479.
- For negative mode, add 28~30% ammonia solution to extracted sample solutions at a final concentration of 0.1% (v/v).
Negative mode internal standards: 2.4-D (plant hormone) m/z 218.96157, Ampicillin (antibiotic) m/z 348.10180, CHAPS 8 detergent) m/z 613.388865, (GluNAc)4 m/z 829.32023.
- Mass spectra are acquired over the 100-1,000 m/z range and accumulate to improve the S/N ratio. The time period for accumulation depends on the total ion concentration. Analyze peaks using the IonSpec Omega ver.8 software. Proofread m/z of each peak referencing the internal standards. Measure the product ion mass spectra of each sample three times.
- When ion peaks are detected at least twice out of three successive spectral scans, they are subjected to further data processing as ion signals from actual analytes.
- A total of four mass spectral peaks from two different extraction solvents (methanol and acetone) and two ionization operation modes (positive and negative) are aligned using our in-house Java program. (If you are interested in this in-house Java program, please contact Prof. Ohta.)
- Perform Mass analysis using an IonSpec Explorer FT-ICR/MS in positive and negative ionization operation modes. Ions are generated from an ESI source with a fused silica needle of 0.005-inch i. d. (Oikawa et al., 2006). Samples are infused using a Harvard syringe pump mode 22 at a flow rate of 0.5 to 1.0 µl/min through a 100 µl Hamilton syringe. Set the potentials on the electrospray emitters to 3.0 kV and -0.3 kV for the positive and the negative electrosprays, respectively.
- Normalization of data
Apply global normalization to data. To avoid zero division, missing values are filled with 105 as a background signal of FT-ICR/MS. Peak intensities are transformed using a logarithmic scale with a factor of 10. Four data matrices are used to apply global normalization.
Global normalization calculation methods are as follows:
- Average intensity is calculated by dividing the total signal by the number of detected peaks in each spectrum.
- Average signal is calculated for all spectra in each elution and charge pair.
- Normalization factor is calculated for each spectrum by dividing the average intensity for each spectrum by total average intensity.
- Normalized intensity is calculated by multiplying the raw intensity in each spectrum by the previously calculated normalization factor. Empirical formulas are inferred by the accuracy of the FT-ICR/MS. Because the sample ions become adduct ions to attach protons and sodium ions, etc. on the ESI source, we assumed the following were involved in the detected peaks [M+H]+, [M+Na]+, [M+K]+, [M+H+methanol]+, [M+ammonium]+ in positive ionization mode and [M-H]- in negative ionization mode.
- Search for candidate compounds using KEGG (http://www.genome.jp/kegg/), NIST (http://webbook.nist.gov/chemistry/) and KNApSAck (Shinbo et al., 2006).
- Average intensity is calculated by dividing the total signal by the number of detected peaks in each spectrum.
Notes
- In order to obtain reliably reproducible data, it is imperative that the plant growth environment be as uniform/consistent as possible. For example depending on the light environment the value of a plant chlorophyll fluorescence will fluctuate. The amount of light a plant receives when next to the side light on the growth incubator is completely different from the light it receives when on the center of the shelf. For this reason it is important to shuffle the location of growth plates, etc.
- It is important that FT-ICRMS maintains a steady vacuum. We must also be careful about the size of the sample cone on the end of the ESI spray nozzle. It is also important to make minor adjustments where necessary to the direction of ESI ionspray, in an effort to capture as many ions into ICR cells as possible.
- FT-ICRMS can have problems when the concentration of plant extraction samples is too high, or when other unwanted substances are included in samples. Regular baking and removal of ions from the chamber inner wall as well as cleaning of the ion source is required.
Acknowledgments
This work was supported by the Ministry of Education, Culture, Sports, Science and Technology (Japan) [Grants-in-Aid for Scientific Research (No.17681022 to R.M.)].
FT-ICR/MS (IonSpec) machine belongs to Osaka Prefecture University, and for this project it was used according to directions as indicated by Professors Ohta and Oikawa of the same university. Data normalization techniques were established by Mr. Satou.
This protocol is modified and appended referencing the original, as featured in "Integrated analysis of transcriptome and metabolome of Arabidopsis albino or pale green mutants with disrupted nuclear-encoded chloroplast proteins" (Satou et al., 2014).
References
- Oikawa, A., Nakamura, Y., Ogura, T., Kimura, A., Suzuki, H., Sakurai, N., Shinbo, Y., Shibata, D., Kanaya, S. and Ohta, D. (2006). Clarification of pathway-specific inhibition by Fourier transform ion cyclotron resonance/mass spectrometry-based metabolic phenotyping studies. Plant Physiol 142(2): 398-413.
- Satou, M., Enoki, H., Oikawa, A., Ohta, D., Saito, K., Hachiya, T., Sakakibara, H., Kusano, M., Fukushima, A., Saito, K., Kobayashi, M., Nagata, N., Myouga, F., Shinozaki, K. and Motohashi, R. (2014). Integrated analysis of transcriptome and metabolome of Arabidopsis albino or pale green mutants with disrupted nuclear-encoded chloroplast proteins. Plant Mol Biol 85(4-5): 411-428.
- Shinbo, Y., Nakamura, Y., Altaf-Ul-Amin, M., Asahi, H., Kurokawa, K., Arita, M., Saito, K., Ohta, D., Shibata, D. and Kanaya, S. (2006). KNApSAcK: a comprehensive species-metabolite relationship database. Biotech Agric Forest 57: 165-181.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Motohashi, R., Satou, M., Myouga, F., Oikawa, A. and Ohta, D. (2015). Arabidopsis Metabolome Analysis Using Infusion ESI FT-ICR/MS. Bio-protocol 5(9): e1463. DOI: 10.21769/BioProtoc.1463.
Category
Plant Science > Plant biochemistry > Other compound
Plant Science > Plant metabolism > Metabolite profiling
Systems Biology > Metabolomics > Whole organism
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










