- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Detection of the Secreted and Cytoplasmic Fractions of IpaB, IpaC and IpaD by Lysozyme Permeabilization
Published: Vol 4, Iss 20, Oct 20, 2014 DOI: 10.21769/BioProtoc.1271 Views: 9985
Reviewed by: Kanika GeraAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

From Llama to Nanobody: A Streamlined Workflow for the Generation of Functionalised VHHs
Lauren E.-A. Eyssen [...] Raymond J. Owens
Mar 20, 2024 6209 Views
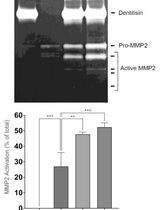
Purification of Native Dentilisin Complex from Treponema denticola by Preparative Continuous Polyacrylamide Gel Electrophoresis and Functional Analysis by Gelatin Zymography
Pachiyappan Kamarajan [...] Yvonne L. Kapila
Apr 5, 2024 2087 Views
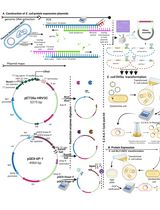
Thermus thermophilus CRISPR Cas6 Heterologous Expression and Purification
Junwei Wei [...] Yingjun Li
Jul 20, 2025 2167 Views
Abstract
Gram negative bacterial pathogens, such as Shigella flexneri, which possess a Type Three Secretion System (T3SS), are able to transfer bacterial proteins, dubbed translocators and effectors, from their cytoplasm into the cytoplasm of their host cells using a syringe like needle complex. For Shigella, it has been shown that during cellular invasion, the intrabacterial pool of translocators and effectors is completely depleted upon activation of the TTS Apparatus and is then progressively replenished while bacteria remain inside host cells. Replenishment of effectors allows for cell-to-cell spreading events, which also necessitate reactivation of the T3SA, and lead to another round of depletion of intrabacterial effector stores. To understand the state of individual intracellular bacteria during infection, it is therefore of interest to be able to locate and evaluate the relative quantity of the intrabacterial and secreted pool of translocators and effectors. We recently adapted a method based on EDTA and lysozyme to permeabilize the cell wall of bacteria present within host cells in order to label the intrabacterial pool of the tip protein IpaD and the translocators IpaB and IpaC. Herein, we describe in detail the protocol to perform the successive labeling of the intrabacterial and secreted pools. This method is theoretically extendable to virulence factors secreted by other secretion systems and other bacterial pathogens.
Materials and Reagents
- Shigella spp. Strain(s) (e.g. M90T)
- Human tissue culture cells such as colonic epithelial TC7 cells (a clone of Caco-2 cells)
Note: Only polarized epithelial cells permit efficient cell-to-cell spread of Shigella spp. We also recommend using human cells because it is the sole natural host of Shigella, although most cell lines of other origins tested are also readily infected and could be used for practical reasons. - Petri dish of Tryptone Casein Soja (TCS) agar (BD Biosciences, catalog number: 236950 ) supplemented with 0.01% Congo red (CR) (SERVA Electrophoresis GmbH, catalog number: 27215.01 ) and the appropriate antibiotic
- TCS broth (BD Biosciences, catalog number: 211825 )
- Polylysine (Sigma-Aldrich, catalog number: P1274 )
- DMEM (Life Technologies, catalog number: 31885 )
- FCS (Biowest, catalog number: S1810-100 )
- Penicillin/streptomycin (Life Technologies, catalog number: 15140 )
- Non-essential amino acids (Life Technologies, catalog number: 11140 )
- Trypsin-EDTA (Life Technologies, catalog number: 25200-056 )
- Fibronectin from human plasma (Sigma-Aldrich, catalog number: F0895 ) (optional)
- HEPES (Life Technologies, catalog number: 15630-056 )
- Gentamycin (EUROMEDEX, catalog number: EU0540-A )
- PFA (Electron Microscopy Sciences, catalog number: 15714 )
- Glycine (Sigma-Aldrich, catalog number: G7126 )
- Triton X-100 (Sigma-Aldrich, catalog number: T8787 )
- Bovine serum albumin (BSA) (Sigma-Aldrich, catalog number: A 9647 )
- Gelatin (Sigma-Aldrich, catalog number: G1393 )
- Sodium azide (Sigma-Aldrich, catalog number: 0 8591 )
- Goat anti-mouse IgG Alexa568 (highly cross absorbed) (Life Technologies, catalog number: A11036 )
- Goat anti-mouse IgG Alexa647 (highly cross absorbed) (Life Technologies, catalog number: A21236 )
- Lysozyme (Sigma-Aldrich, catalog number: 6876 )
- 5, 000x DAPI stock solution (Sigma-Aldrich, catalog number: D9542 ) (optional) (see Recipes)
- MOWIOL (Sigma-Aldrich, catalog number: 81381 )
- 1,4-diazobicyclo-[2,2,2]-octane d (DABCO) (Sigma-Aldrich, catalog number: D27802 )
- Glycerol (VWR International, catalog number: 24388.295 ) alternatively Prolong mounting medium (Life Technologies, catalog number: P36930 )
- 10x phosphate-buffer saline (PBS) (see Recipes)
- PBS/10 µg/ml polylysine (see Recipes)
- DMEM/20 mM HEPES (see Recipes)
- Chase medium (see Recipes)
- PBS/4% PFA (see Recipes)
- PBS/100 mM glycine (see Recipes)
- PBS/0.1% Triton X-100 (see Recipes)
- PBS/1% BSA/0.2% gelatin (see Recipes)
- Lysozyme reaction buffer (see Recipes)
- 5,000x DAPI stock solution (see Recipes)
- Mounting medium (see Recipes)
- Growth medium for TC7 cells (see Recipes)
Equipment
- 24-well plates
- Coverslips no. 1.5, 12 mm in diameter (Harvard Apparatus, catalog number: 64-0712 )
- Laminar flow hood for cell culture
- Heating water bath
- Tabletop centrifuge for 1.5 ml Eppendorf tubes
- Tabletop centrifuge with plate-holding rotor
- CO2 incubator for cell culture
- Tweezers
Procedure
There are numerous studies in the literature describing protocols to perform immunofluorescence labeling of intrabacterial proteins (Buddelmeijer et al., 1998; Schlumberger et al., 2005). Although the detailed procedure might vary, they have in common the use EDTA and lysozyme, which destabilize the outer membrane [i.e. LPS, reviewed by Nikaido and Vaara (1985)] and degrade the peptidoglycan of the cell wall, respectively, hence allowing antibodies to diffuse into the bacterial cytoplasm to recognize their cognate antigen. Below we present the method that we have used to detect in combination the secreted and intrabacterial fractions of Shigella flexneri IpaB, IpaC and IpaD. See Reference 2 (Campbell-Valois et al., 2014) for application of the procedure.
- 36-48 h before the experiment, confluent TC7 cells monolayer (i.e. 100% coverage of the surface) are detached by incubation with 0.25 % trypsin-EDTA and distributed in their normal growth medium (see Recipes) onto sterile 12 mm coverslips (CS) no. 1.5 within wells of a 24-wells plate. To follow entry events, seed 5 x 104 cells/well (approximately 50-75% confluence the day of the experiment). To favor cell-to-cell spread at longer infection times, seed 1 x 105 cells/well. (Optional) Fibronectin-coated coverslips (CS) can be used to favor attachment of the cells, but in this case, the number of cells seeded must be reduced approximately in half in order to obtain confluence similar to what is described for non-coated coverslips.
- Day before the experiment, a CR–positive S. flexneri colony (i.e. with a red dot at the center of the colony) is picked from a TCS-CR plate and used to inoculate a tube of TCS broth (e.g. 2-8 ml) for overnight growth at 30 °C with shaking.
Note: CR induces T3SA activation and effector secretion; colonies with an active T3SA therefore display a red center where CR has accumulated while bacteria that have lost the virulence plasmid show up as larger white colonies (Figure 1).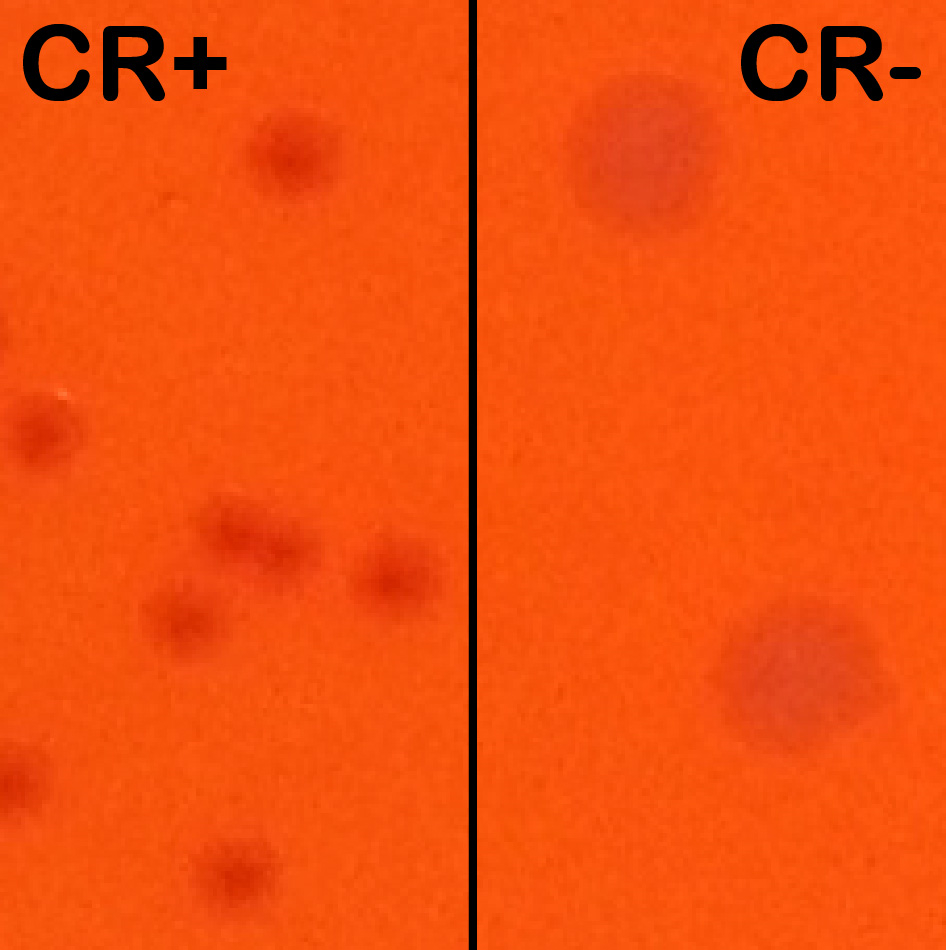
Figure 1. Congo red positive (CR+) versus Congo red negative (CR-) colonies. Wt and plasmid cured M90T derived strains were streaked side-by-side on an agar-containing TCS and congo red plate to show their differing phenotypes. Wt colonies on the left are forming small and CR+ colonies (displaying a red center). Plasmid cured (BS176 strain) are forming large CR- colonies (white). - The next morning, the ON culture is subcultured 1:100 in a tube of 8.0 ml fresh TCS broth without antibiotics and incubated at 37 °C until the OD600 is approximately 1.0 (late exponential phase).
- 2 ml of bacterial culture (sufficient for the infection of 4 wells of a 24-wells plate) is centrifuged at 7,500 x g for 1 min and washed in 1 ml PBS. Repeat two other times the centrifugation and washing steps.
- Resuspend the bacterial pellet in 2 ml DMEM-HEPES at room temperature (RT).
- (Optional) Permeabilization can be directly performed on bacteria grown in tubes. In this case, simply centrifuge 5 min at 450 x g a 1:5 dilution of the washed bacteria onto the required number of polylysine-coated 12 mm coverslips (incubate the sterile coverslips with 50 µg/ml polylysine in PBS for 60 min at RT, wash three times in 1 ml PBS); the use of polylysine-coated coverslips facilitates bacterial adhesion to the glass surface. Place coverslips in a 24-wells plate, wash two times with PBS and proceed with fixation, as described below.
- TC7 cells are washed two times with DMEM-HEPES (RT); after the washes place DMEM-HEPES at 37 °C for step 10.
- 500 µl of the DMEM-HEPES bacterial suspension is added to each well of the 24-wells plate containing the TC7 cells.
- Bacteria are brought into contact with the cells by centrifugation for 5 min at 450 x g at RT. Cells are maintained in contact with the bacterial suspension for approximately 10 min at RT in total. (Optional) If desirable, polylysine-coated bacteria can be used instead of centrifugation of non-coated bacteria. Polylysine-coated bacteria can be rapidly prepared by incubating bacteria washed once with 1 ml PBS (see step 4), in PBS/polylysine (10 µg/ml) for 10 min with gentle agitation and washing of the bacteria three times prior to infection. Entry efficiency is greatly enhanced using polylysine treatment, so a bacterial suspension with OD600 equal to 0.02-0.2 should be incubated with the cells at RT for 10 min without centrifugation.
- The bacteria suspension is aspirated and replaced by 500 µl of DMEM-HEPES preheated to 37 °C.
- Incubate between 15 to 30 min in a cell culture incubator with 5% CO2 at 37 °C.
- Aspirate and add the chase medium for the desired time. To study cell-to-cell spread in TC7 cells, a chase of 210 min is recommended.
- At the desired time after infection, wash the cell monolayer once with PBS.
- Fix for 10 min with 200 µl PBS/4% PFA at room temperature (RT).
- Aspirate the fixative and incubate 5 min in 200 µl PBS/100 mM glycine to quench the PFA.
- Wash with 500 µl PBS 3 times.
- Incubate 4 min in 200 µl PBS/0.1% Triton X-100.
- Wash with 500 µl PBS 4 times.
- Incubate CS for 15 min in 200 µl blocking solution PBS/1% BSA/0.2% gelatin.
- Incubate CS ON at 4 °C with the primary antibody, directed either against IpaB, IpaC, IpaD or alternatively against any other secreted protein of interest, diluted in 200 µl blocking solution/coverslip. The dilution of the antibody is variable and must be determined empirically for each antibody.
- Wash 3 times with 500 µl PBS.
- Incubate for 120 min with the appropriate secondary antibody conjugated to Alexa Fluor 568 and diluted in blocking solution.
- Wash 3 times in 500 µl PBS.
- Incubate 5 min in 200 µl PBS/4% PFA to crosslink secondary and primary antibodies to their antigen and thus, avoid cross-reaction with the intrabacterial pool following lysozyme permeabilization.
- Incubate 5 min in 200 µl PBS/glycine.
- Wash 3 times in PBS.
- Incubate CS 20 min with the lysozyme reaction buffer at 37 °C.
- Wash 3 times in 500 µl PBS.
- Incubate overnight (ON) at 4 °C or 2-4 h at RT with the primary antibody directed against IpaB, IpaC, IpaD or alternatively against any other secreted proteins of interest diluted 200 µl blocking solution/coverslip. Again, the dilution of the antibody is variable and must be determined empirically by the user.
- Wash 3 times in 500 µl PBS.
- Incubate for 120 min with the appropriate secondary antibody conjugated to Alexa Fluor 647 or equivalent fluorophore diluted 1/500-1/1,000 or according to supplier in 200 µl blocking solution/coverslip. At this step, DAPI can be added at 0.2 µg/ml during the incubation.
- Wash 3 times in PBS and heat mounting medium to 37 °C.
- Take out the CS from the plate with a fine pair of tweezers, briefly immerse in distilled water and remove excess water using a sheet of absorbing paper.
- Mount on a slide using 10 µl of pre-warmed (37 °C) mounting medium.
- Let dry without disturbing the CS for at least one hour at RT.
- Store at 4 °C until ready to observe.
- Expected results: Secreted fraction will appear in both the Alexa568 and Alexa647 channels, while the intrabacterial fraction will only appear in the Alexa647 channel. Localization of the secreted fraction will appear different than those of the intrabacterial fraction. Due to the sequential permeabilization procedure and cross-linking with PFA between labeling of the secreted and intrabacterial fractions, it is possible to use sequentially the same primary antibodies directed against the translocators or effectors to perform this protocol without compromising interpretation of the results [see Figure 4C in Campbell-Valois et al. (2014) and Figure 2 below]. In most cases, bacteria with secreted translocators or tip proteins in their vicinity have depleted intrabacterial store. Sometime bacteria with no detectable intrabacterial or secreted store are observed. They most likely represent bacteria that have recently depleted their intrabacterial store and have not replenished their stock yet.
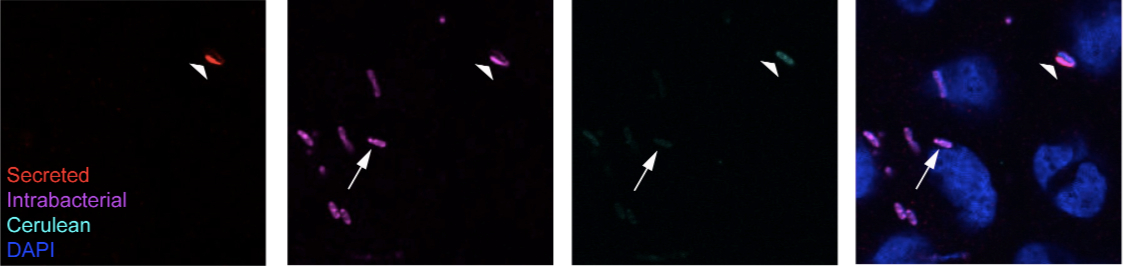
Figure 2. Example of labeling obtained following the protocol. Here we show the final result obtained with the labeling protocol described above using a single monoclonal antibody (A8.1) directed against the tip protein paD expressed by Wt Shigella flexneri. The arrow points at a bacterium, which has only intrabacterial IpaD protein (+lysozyme) and no detectable secreted fraction (-lysozyme). The large arrowhead points at a bacterium with secreted IpaD in its vicinity and demonstrate the cross-reaction described in step 37 that is systematically observed for bacteria with secreted IpaB, IpaC and IpaD in their vicinity, following lysozyme treatment. Contrary to bacteria similar to the one indicated by the arrow, the bacteria indicated by the large arrowhead do not have IpaD in its cytoplasm, as indicated by the hollow shape of the labeling.
Note: Cerulean is a cyan variant of the green fluorescent protein from Aequoria victoria, here expressed from a constitutively active promoter by the bacteria.
Notes
- Results are usually highly reproducible. However to ensure that the ratio of secreting bacteria to not secreting bacteria be constant at a given time post-challenge under scrutiny, it is recommended to ensure that the infected cells be studied at a similar level of confluence because when bacteria are unable to perform cell-to-cell spread efficiently, the proportion of immobile, hence not secreting bacteria increases. Similarly, bacteria subculturing prior to infection must be performed in highly controlled conditions, for example, by taking care of stopping incubation at similar OD600 each time an experiment is repeated. This is particularly important if the labelings are performed directly on bacteria grown in broth or at early time points post-challenge as the amount of translocators found in the bacterial cytoplasm is known to increase throughout the exponential growth phase.
Recipes
- 10x phosphate-buffer saline (PBS)
Mix 80 g of NaCl with 2 g of KCl, 14.4 g of Na2HPO4, 2.4 g of KH2PO4
Adjust pH to 7.4 with NaOH
Add distilled H2O to 1,000 ml
Sterilize by autoclaving
Stored at RT
Diluted to 1x concentration for all uses - PBS/10 µg/ml polylysine
1x PBS
1:1,000 of the polylysine stock solution (10 mg/ml in water, filter sterilized and stored at -20 °C in 1 ml aliquots) - DMEM/20 mM HEPES
DMEM
1:50 dilution of 1 M HEPES - Chase medium
DMEM
5% FBS
Gentamycin 50 µg/ml - PBS/4% PFA
10 ml PFA (32%)
8 ml 10x PBS
Add distilled H2O to 80 ml
Aliquot and store at -20 °C
Discard aliquot after one week at 4 °C - PBS/100 mM glycine
0.375 g glycine
5 ml 10x PBS
Add H2O to 50 ml
Filter sterilize (0.2 µm)
Stored at 4 °C - PBS/0.1% Triton X-100
500 µl PBS/10% Triton X-100 stock solution
5 ml 10x PBS stock solution
Filter sterilize (0.2 µm)
Add distilled H2O to 50 ml
Stored at 4 °C - PBS/1% BSA/0.2% gelatin
0.5 g BSA
0.1 g gelatin
50 ml 1x PBS
Add sodium azide to final 2 mM
Filter sterilize (0.2 µm)
Stored at 4 °C - Lysozyme reaction buffer
PBS 0.8x from 10x PBS stock solution
50 mM glucose
5 mM EDTA
Supplement immediately before use with 5 mg/ml lysozyme using a 100 mg/ml stock solution Aliquoted and stored at -20 °C - 5,000x DAPI stock solution
1 ml H2O
1 mg DAPI - Mounting medium
Note: Recipe from the University of Rochester Medical Center, Michael Mastrangelo, Eric Yehling; for details see http://www.urmc.rochester.edu/confocal-conventional-microscopy/documents/Mowiol52810.pdf.
24 g analytical grade glycerol
9.6 g Mowiol 4-88
24 ml distilled H2O
48 ml 0.2 M Tris buffer (pH 8.5)
To dissolve, completely assemble the solution in a 500 ml flask and stir on a heating plate at 60-70 °C for 4-5 h
Aliquot in 20-40 ml per 50 ml conical tube
Centrifuge at 5,000 x g for 15 min to remove undissolved residues and carefully recover the supernatant
Add 2.5% DABCO as anti-bleaching agent
Aliquot in 15 or 50 ml conical tubes and stored at -20 °C for long-term storage - Growth medium for TC7 cells
DMEM
20% FCS
1x penicillin/streptomycin
1x non-essential amino acids
Acknowledgments
We are grateful to Claude Parsot for his comments on the manuscript. FXCV was a CIHR, EMBO, Marie-Curie-IRG and FRM fellow. PS was an EMBO and Marie Curie-IEF fellow while PJS is a HHMI senior scholar. This work was supported by the ERC (PJS Advanced Grant, no. 232798). The procedure described above was originally described in Campbell-Valois et al. (2014).
References
- Buddelmeijer, N., Aarsman, M. E., Kolk, A. H., Vicente, M. and Nanninga, N. (1998). Localization of cell division protein FtsQ by immunofluorescence microscopy in dividing and nondividing cells of Escherichia coli. J Bacteriol 180(23): 6107-6116.
- Campbell-Valois, F. X., Schnupf, P., Nigro, G., Sachse, M., Sansonetti, P. J. and Parsot, C. (2014). A fluorescent reporter reveals on/off regulation of the Shigella type III secretion apparatus during entry and cell-to-cell spread. Cell Host Microbe 15(2): 177-189.
- Nikaido, H. and Vaara, M. (1985). Molecular basis of bacterial outer membrane permeability. Microbiol Rev 49(1): 1-32.
- Schlumberger, M. C., Müller, A. J., Ehrbar, K., Winnen, B., Duss, I., Stecher, B. and Hardt, W.-D. (2005). Real-time imaging of type III secretion: Salmonella SipA injection into host cells. Proc Natl Acad Sci U S A 102(35): 12548-12553.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Campbell-Valois, F. X., Schnupf, P. and Sansonetti, P. J. (2014). Detection of the Secreted and Cytoplasmic Fractions of IpaB, IpaC and IpaD by Lysozyme Permeabilization. Bio-protocol 4(20): e1271. DOI: 10.21769/BioProtoc.1271.
Category
Microbiology > Microbial biochemistry > Protein > Isolation and purification
Microbiology > Microbe-host interactions > Bacterium
Biochemistry > Protein > Labeling
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link












