- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Probing Yeast Protein Microarrays for Protein-protein Interactions Using V5-epitope Tagged Fusion Protein Probes
Published: Vol 2, Iss 5, Mar 5, 2012 DOI: 10.21769/BioProtoc.123 Views: 13535

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
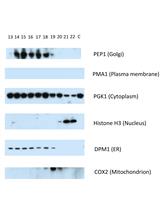
A Protocol to Map the Spatial Proteome Using HyperLOPIT in Saccharomyces cerevisiae
Daniel J. H. Nightingale [...] Stephen G. Oliver
Jul 20, 2019 7186 Views
A Yeast Chromatin-enriched Fractions Purification Approach, yChEFs, from Saccharomyces cerevisiae
Abel Cuevas-Bermúdez [...] Francisco Navarro
Jan 5, 2020 4717 Views
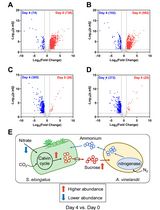
Applying LFQRatio Normalization in Quantitative Proteomic Analysis of Microbial Co-culture Systems
Mengxun Shi [...] Jagroop Pandhal
May 5, 2025 2294 Views
Abstract
Protein microarray is probably the only technique currently available for systematic investigation of protein-protein interactions. This protocol describes an optimized method to probe yeast protein microarrays for protein-protein interactions using purified V5-epitope tagged fusion protein. It should also apply to any other proteins with appropriate modifications.
Keywords: KinaseMaterials and Reagents
- Zirconia beads (Biospec Products, catalog number: 11079110zx 1.0mm dia)
- His-tag affinity resin (His Spintrap, GE Life sciences, catalog number: 28-4013-53 )
- Protein microarray (Life Technologies, catalog number: PAH0525101 )
- V5-AlexaFluor 647 antibody (Life Technologies, catalog number: 451098 )
- Glycerol (Sigma-Aldrich, catalog number: G5516 )
- Potassium chloride
- Sodium chloride
- Triton X-100
- Dithiothreitol (DTT) (Promega, catalog number: P1170 )
- Phenylmethylsulfonyl fluoride (PMSF) (Sigma-Aldrich, catalog number: P7626-250MG )
- Imidazole (Sigma-Aldrich, catalog number: I5513-5G )
- ATP (for kinases)
- BSA
- Tween-20
- Yeast extract
- Peptone
- Galactose
- Yeast nitrogen base
- (NH4)2SO4
- Raffinose (Sigma-Aldrich, catalog number: R0514-100G )
- Agar
- Lysis buffer (see Recipes)
- Wash buffer 1 (see Recipes)
- Wash buffer 2 (see Recipes)
- Wash buffer 3 (see Recipes)
- Elution buffer (see Recipes)
- Probe buffer (see Recipes)
- Blocking buffer (see Recipes)
- 3x YEP-GAL (see Recipes)
- Synthetic complete minus uracil media (Sc-ura) (see Recipes)
Equipment
- FastPrep Cell Lyser (MP Bio, catalog number: 116004500 - 1 each )
- High speed centrifuges
- JA-10 (or comparable) rotor
- Axon 4200 AL microarray reader
- 50 ml conical tube
- Wheel
- Humidified chamber
Procedure
- Probe protein purification
- Construct a V5-His6-protein fusion construct (i.e. pYES-DEST52, Life Technologies, Invitrogen™) of the yeast protein you wish to profile by using the methods outlined by Fasolo and Snyder (2009).
- Yeast (Y258 strain optimized for protein purification (Zhu et al., 2001)) that have been transformed with a pYES-DEST52 fusion construct under control of the GAL1 inducible promoter are grown in Sc-Ura/Dex overnight in 5 ml starter cultures.
- The next day OD600 are determined for the starter cultures and used to inoculate a 400 ml. Sc-Ura/Raffinose culture to an OD600 = 0.1. The culture is grown at 30 °C shaking to an OD600 = 0.55-0.6 and induced with 200 ml 3XYEP-GAL for 6 h at 30 °C shaking.
- Cell pellet is harvested by centrifugation using a JA-10 (or comparable) rotor by spinning 400 ml volumes of cell suspension at 4,000 rpm, 4 °C, 5 min.
- Wash cell pellet in ice cold PBS buffer by transferring the cell pellet to 50 ml conical tube and centrifuging in a tabletop centrifuge at 3,000 rpm, 4 °C, 5 min.
- Re-suspend cell pellet in ice cold PBS once more and aliquot evenly into 4 screw cap 2.0 ml. FASTPREP compatible tubes (pellet weight is ~0.5 g/tube).
- Spin cell suspension at 14,000 rpm, 4 °C, 1 min in a tabletop microfuge and aspirate supernatant. Snap freeze cell pellet immediately in liquid nitrogen and store at -80 °C or lyse immediately (you can stop here and proceed on the following day if not prepared for following steps).
- Add 0.5 mm zirconia beads, and lysis buffer at a 1:1:1 ratio to cell pellet and store on ice.
- Place tubes on FastPrep machine for 6 cycles of lysis at maximum setting for 1 min each, with 1 min on ice in between each interval.
- Spin suspension down in tabletop microcentrifuge at 14,000 rpm, 4 °C, 10 min, 10 min and collect supernatant.
- Repeat steps 9 and 10 then discard extracted pellet.
- Combine supernatant from both extractions into a single tube and incubate with Ni2+, or Co2+ affinity resin for 3 h/4 °C/rotating on a wheel.
- Centrifuge resin and save small aliquot of cleared supernatant for quality control.
- Wash resin 2x 10 min in wash buffer 1 /4 °C/rotating on a wheel.
- Wash resin 2x 10 min in wash buffer 2 /4 °C/rotating on a wheel.
- Wash resin 1x 10 min in wash buffer 3 /4 °C/rotating on a wheel.
- Collect washed resin in G25 or comparable column and elute with elution buffer and store at -80 °C until ready to use in probing assay (stop here is not ready to proceed).
- Construct a V5-His6-protein fusion construct (i.e. pYES-DEST52, Life Technologies, Invitrogen™) of the yeast protein you wish to profile by using the methods outlined by Fasolo and Snyder (2009).
- Protein microarray probing
- Dilute the purified protein probe over a concentration range of 5-500 μg/ml in the probe buffer (concentration of probe is empirically determined for each protein-protein interaction assay).
- Block the arrays in blocking buffer for 1 h by shaking at 50 rpm on a stage at 4 °C.
- After blocking, transfer the arrays to a humidified chamber, and add 90 μl of diluted probe directly to the array surface. Overlay the arrays with a raised lifter slip and incubate static (no shaking) in the humidor for 1.5 h.
- Wash the arrays 3 times for 1 min each in probe buffer in three 50-ml conical tubes.
- To detect interactions, dilute the V5-AlexaFluor 647 antibody to 260 ng/ml in probe buffer and mix thoroughly by shaking on a wheel for 30 min at 4 °C.
- After washing the arrays several times as indicated, add antibody solution directly to array and overlay with a raised lifter slip as before. Incubate the arrays for 30 min/static/4 °C.
- Finally, wash the arrays for 1 min (3x) in probe buffer, and spin in a 50-ml conical tube at 800 x g in a tabletop centrifuge for 5 min at RT. Air-dry the arrays in a slide holder in the dark for 30 min prior to scanning the array at 647 nm on an Axon 4200 AL microarray reader (or comparable).
- Dilute the purified protein probe over a concentration range of 5-500 μg/ml in the probe buffer (concentration of probe is empirically determined for each protein-protein interaction assay).
Notes
- Protein microarrays used in this protocol are produced using full-length GST-fusion fusion proteins that were purified using glutathione sepharose beads. Protein arrays produced using proteins purified with antibody-conjugated matrices may result in high background when probed with anti-V5 antibody for detection due to the presence of residual IgG contamination. It is therefore important to test the background of each array with a negative control consisting of the detection antibody alone.
- Protein probe concentration must be high enough to dilute to working concentration in probe buffer without effecting solute concentration of probe buffer. This can be accomplished by scaling up culture to >10 L if necessary. Downstream applications like molecular weight cut-off columns may be used to exchange solutions and remove excess imidazole from final probe solution.
- Imidazole gradient in wash steps may be altered to optimize protein purification which can vary from protein to protein; protein probes must be at ~80-90% pure in order to which can be assessed by mass spectrometry and SDS-PAGE.
Recipes
- Lysis buffer
PBS (0.01 M phosphate buffer, 0.0027 M potassium chloride, 0.137 M sodium chloride) (pH 7.4)
150 mM NaCl
10% glycerol
0.1% Triton X-100
0.5 mM DTT
1 mM PMSF
1x complete protease inhibitor tablet (F. Hoffmann-La Roche) - Wash buffer 1
PBS (0.01 M phosphate buffer, 0.0027 M potassium chloride, 0.137 M sodium chloride) (pH 7.4)
150 mM NaCl
50 mM Imidazole
10% glycerol
0.1% Triton X-100 - Wash buffer 2
PBS (0.01 M phosphate buffer, 0.0027 M potassium chloride, 0.137 M sodium chloride) (pH 7.4)
150 mM NaCl
100 mM Imidazole
10% glycerol
0.1% Triton X-100 - Wash buffer 3
PBS (0.01 M phosphate buffer, 0.0027 M potassium chloride, 0.137 M sodium chloride) (pH 7.4)
150 mM NaCl
150 mM Imidazole
10% glycerol
0.1% Triton X-100 - Elution buffer
PBS (0.01 M phosphate buffer, 0.0027 M potassium chloride, 0.137 M sodium chloride) (pH 7.4)
0.1% Triton X-100
500 mM NaCl
0.5 mM DTT
500 mM imidazole
2 mM MgCl2
25% glycerol (if freezing at -80 °C) - Probe buffer
PBS (0.01 M phosphate buffer, 0.0027 M potassium chloride, 0.137 M sodium chloride) (pH 7.4)
1% glycerol
2 mM MgCl2
0.5 mM DTT
0.05% Triton X-100
50 mM NaCl
500 μM ATP (for kinases)
1% BSA - Blocking buffer
PBS (0.01 M phosphate buffer, 0.0027 M potassium chloride, 0.137 M sodium chloride) (pH 7.4)
1% BSA
0.1% Tween-20 - 3x YEP-GAL (yeast extract–peptone–galactose)
30 g yeast extract
60 g peptone
Make up to 700 ml with dH2O
Add 300 ml of sterile filtered 20% galactose to media after autoclaving. - Synthetic complete minus uracil media (Sc-ura)
1.5 g yeast nitrogen base
5 g (NH4)2SO4
2 g Sc-ura drop out mix (commercially available)
20 g raffinose (or dextrose for starter culture media and SD-ura plates)
20 g agar (for plates)
Acknowledgments
This protocol was developed in the Snyder Lab, Department of Genetics, Stanford University, Stanford, CA, USA. It was adapted from Fasolo et al. (2011), and Fasolo and Snyder, (2009). This work was supported by the NIH.
References
- Fasolo, J., Sboner, A., Sun, M. G., Yu, H., Chen, R., Sharon, D., Kim, P. M., Gerstein, M. and Snyder, M. (2011). Diverse protein kinase interactions identified by protein microarrays reveal novel connections between cellular processes. Genes Dev 25(7): 767-778.
- Fasolo, J. and Snyder, M. (2009). Protein microarrays. Methods Mol Biol 548: 209-222.
Article Information
Copyright
© 2012 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Fasolo, J. and Snyder, M. (2012). Probing Yeast Protein Microarrays for Protein-protein Interactions Using V5-epitope Tagged Fusion Protein Probes. Bio-protocol 2(5): e123. DOI: 10.21769/BioProtoc.123.
Category
Systems Biology > Proteomics > Protein microarray
Microbiology > Microbial proteomics > Whole organism
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link