- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Affinofile Assay for Identifying Macrophage-Tropic HIV-1
Published: Vol 4, Iss 14, Jul 20, 2014 DOI: 10.21769/BioProtoc.1184 Views: 9305
Reviewed by: Anonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
General Maintenance and Reactivation of iSLK Cell Lines
Ariana C. Calderón-Zavala [...] Ekaterina E. Heldwein
Jun 5, 2025 1875 Views
Inducible HIV-1 Reservoir Reduction Assay (HIVRRA), a Fast and Sensitive Assay to Test Cytotoxicity and Potency of Cure Strategies to Reduce the Replication-Competent HIV-1 Reservoir in Ex Vivo PBMCs
Jade Jansen [...] Neeltje A. Kootstra
Jul 20, 2025 2457 Views
Assembly and Mutagenesis of Human Coronavirus OC43 Genomes in Yeast via Transformation-Associated Recombination
Brett A. Duguay and Craig McCormick
Aug 20, 2025 3033 Views
Abstract
The ability to enter monocyte-derived macrophage (MDM) in vitro is commonly used to define macrophage-tropic HIV-1 despite the fact that viruses vary continuously in their ability to enter MDMs in vitro, and MDMs vary in their ability to support HIV-1 entry (Joseph et al., 2014; Peters et al., 2006). This makes it difficult to distinguish viruses that are adapted to replicating in macrophage from those that are adapted to replicating in T cells. We use the Affinofile cell line ( Johnston et al., 2009) to assay for macrophage tropism by capitalizing on the fact that macrophage-tropic HIV-1 has an enhanced ability to enter cells expressing low levels of CD4 (Joseph et al., 2014; Peters et al., 2006; Duenas-Decamp et al., 2009; Dunfee et al., 2006; Gorry et al., 2002; Martin-Garcia et al., 2006; Peters et al., 2004) and Affinofile cells can be induced to express a wide range of CD4 densities (Johnston et al., 2009). We induce Affinofile cells to express either high or low CD4, infect those cells with pseudotyped reporter virus, and quantify percent infectivity at low CD4 relative to infectivity at high CD4. Macrophage-tropic viruses have an enhanced ability to infect at low CD4. Using this approach we have found that macrophage-tropic strains of HIV-1 are relatively rare and that most HIV-1 variants require high levels of CD4 to enter cells, a phenotype we have referred to as R5 T cell-tropic.
Materials and Reagents
- Affinofile cells (Johnston et al., 2009)
- Luciferase Assay System (Promega Corporation, catalog number: E1501 )
- 5x Reporter Lysis Buffer (Promega Corporation, catalog number: E397A )
- PE-conjugated anti-CD4 antibody (clone RPA-T4) (BD, catalog number: 555347 )
- PE-conjugated anti-CCR5 antibody (clone 2D7) (BD, catalog number: 555993 )
- Aqua live/dead stain (Life Technologies, catalog number: L34957 )
- Env-pseudotyped luciferase reporter virus stock (frozen and tittered)
- Blasticidin (Life Technologies, catalog number: A11139-03 )
- FBS (Atlanta Biologicals, catalog number: S12850 )
- DMEM-H (Cellgro, catalog number: 10-013-CV )
- 10% Buffered Formalin (Thermo Fisher Scientific, catalog number: SF100 )
- Poly-L-Lysine (Sigma-Aldrich, catalog number: P4707 ) (see Recipes)
- Ponasterone A (Life Technologies, catalog number: 45-0478 ) (see Recipes)
- Doxycycline (Sigma-Aldrich, catalog number: D9891 ) (see Recipes)
- DMEM-F10/B (see Recipes)
- 1x staining solution (see Recipes)
- Fixing solution (see Recipes)
Equipment
- 96-well plate (sterile, black) (Corning, Costar®, catalog number: 3916 )
- 24 well plate
- 37 °C, 5% CO2 cell culture incubator (BSL-2)
- Hemocytometer
- Light microscope
- PE-conjugated QuantiBRITE beads (BD, catalog number: 340495 )
- Incubated plate centrifuge (BSL-2)
- Laminar flow biosafety cabinet (BSL-2)
- Plate-reading luminometer
Procedure
- For high versus low CD4 comparison
Day 1- Add 50 µl of diluted Poly-L-Lysine to sterile, black 96 well plates and incubate at 37 °C for 20 min. Remove Poly-L-Lysine.
- Treat Affinofile cells with trypsin-EDTA and harvest in DMEM-F10/B.
- Count cells and dilute to a concentration of 1.8 x 105 cells/ml in DMEM-F10/B.
- Plate 100 µl of cells/well in 96 well plates.
- Plate 0.75 ml of cells/well in 24 well plate for flow cytometry. Each well will be stained for flow cytometry. Plate enough cells to perform the necessary isotype and compensation controls.
- Incubate cells at 37 °C overnight.
- Prepare media for high and low CD4 induction.
- Dilute Doxycycline (Doxy) stock to 4x in DMEM-F10/B. A concentration of 24 ng/ml corresponds to 4x the concentration necessary for maximum CD4 induction.
- Dilute Ponasterone A (Pon A) stock to 4x in DMEM-F10/B. A concentration of 20 µM corresponds to 4x the concentration necessary for maximum CCR5 induction.
- Low CD4 induction medium (Low CD4/High CCR5): Mix equal volumes of 4x Pon A and DMEM-F10/B (=2x Pon A and 0x Doxy).
- High CD4 induction medium (High CD4/High CCR5): Mix equal volumes of 4x Pon A and 4x Doxy (=2x Pon A and 2x Doxy).
- Dilute Doxycycline (Doxy) stock to 4x in DMEM-F10/B. A concentration of 24 ng/ml corresponds to 4x the concentration necessary for maximum CD4 induction.
- Induce cells for infection.
- Remove 50 µl of medium from each well of the 96-well plate.
- For each virus being tested, add 50 µl of low CD4 induction medium to 3 wells (=1x Pon A and 0x Doxy) and 50 µl of High CD4 induction medium to 3 wells (=1x Pon A and 1x Doxy).
- Incubate cells at 37 °C.
- Remove 50 µl of medium from each well of the 96-well plate.
- Induce cells for flow cytometry.
- Remove all medium from the 24 well plate, replace with 750 µl of warm DMEM-F10/B and 250 µl of either 4x Doxy or 4x Pon A (=1x desired amount of Pon A or Doxy).
- Incubate cells at 37 °C.
- Remove all medium from the 24 well plate, replace with 750 µl of warm DMEM-F10/B and 250 µl of either 4x Doxy or 4x Pon A (=1x desired amount of Pon A or Doxy).
- Infect cells 18-24 h after adding drugs (Pon A and Doxy).
- Remove induction medium from wells and immediately replace with 50 µl of warm DMEM-D10/B.
- Warm plate centrifuge to 37 °C.
- Thaw pseudovirus stock and dilute in DMEM-D10/B so that 50 µl of diluted virus produces a desired signal when titered on Affinofile cells expressing maximum CD4 and CCR5. This is determined by pre-titering the virus to ensure the amount of virus added is within the linear dose-response curve for the readout (in this case luciferase) and many fold above the assay background. As an additional control, include a dilution curve of a high titer virus to ensure all signals are within the linear range.
- Add 50 µl of diluted virus to each well of a 96-well plate (3 High CD4 wells and 3 Low CD4 wells per virus).
- Centrifuge plates at 2,000 rpm (849 x g) for 2 h at 37 °C (spinoculation; O'Doherty et al., 2000).
- Incubate cells at 37 °C for 48 h.
- Remove induction medium from wells and immediately replace with 50 µl of warm DMEM-D10/B.
- Prepare cells for flow cytometry.
- Remove medium from a well of a 24 well plate, then use 1 ml of cold PBS to harvest the cells from that well and store the cells in eppendorf tubes on ice until all wells are harvested.
- Centrifuge the harvested cells in a table top microcentrifuge at 4 °C at 2,600 rpm for 10 min.
- While centrifuging, make staining solutions (Recipe 5).
- Resuspend the cells in 50 µl of staining solution and stain at room temperature, in the dark, for 60 min.
- Add, with gentle mixing, 1 ml of fresh fixing solution (Recipe 6).
- Centrifuge in cold room at 2,600 rpm for 10 min, dispose of supernatant.
- Resuspend cells in 1 ml PBS.
- Centrifuge in cold room at 2,600 rpm for 10 min.
- Resuspend cells in 300 µl PBS.
- Store in the dark at 4 °C until ready to analyze.
- Analyze cells and QuantiBRITE beads using flow cytometer equipped with a 405 nm laser and a filter that measures emissions at approximately 526 nm and a 488 nm laser with a filter that measures emissions at approximately 578 nm.
- Use Quantibrite beads, with their four known levels of PE, to determine the relationship between mean fluorescence and number of PE molecules (Figure 1). This relationship can be used to translate the PE fluorescence of stained cells into mean number of CD4 or CCR5 antibody binding sites (ABS) per cell. This method requires that cells be stained with saturating levels of antibody and assumes that each antibody is conjugated to a single PE molecule.
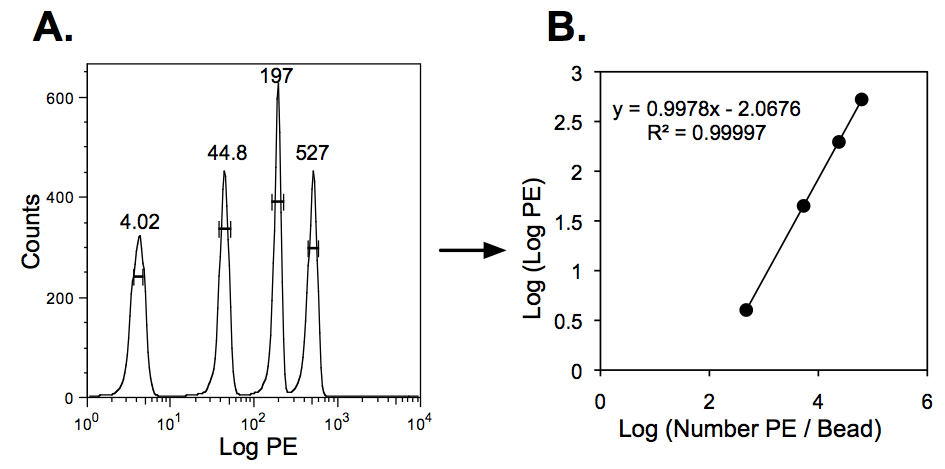
Figure 1. QuantiBRITE beads are used to quantify the relationship between PE fluorescence and number of PE molecules attached to a bead/cell. A) QuantiBRITE beads are analyzed by flow cytometry to calculate the mean, log PE fluorescence of the four bead populations, each conjugated to a different, known number of PE molecules. B) The relationship between log (PE molecules / bead) and log (log PE) is then determined. This relationship can then be used to translate the mean PE fluorescence of stained cells into the mean number of CD4 or CCR5 antibody binding sites (ABS) per cell.
- Remove medium from a well of a 24 well plate, then use 1 ml of cold PBS to harvest the cells from that well and store the cells in eppendorf tubes on ice until all wells are harvested.
Day 5- Harvest infected cells 48 h after infection.
- Remove medium from wells.
- Gently rinse 2x with warm PBS, discard PBS after each wash.
- Add 50 µl of 1x Reporter Lysis Buffer and incubate at room temperature for 15 min.
- Seal plates with aluminum foil covers and freeze at -80 °C for at least 2 h.
- Thaw plates and Luciferase Assay System reagent at room temperature.
- Prepare Luciferase Assay reagents following the manufacturer’s protocol, and read plate using a plate-reading luminometer.
- Calculate percent infectivity at low CD4 relative to infectivity at high CD4 for each individual virus.
- Remove medium from wells.
- Add 50 µl of diluted Poly-L-Lysine to sterile, black 96 well plates and incubate at 37 °C for 20 min. Remove Poly-L-Lysine.
- For continuous CD4 comparison
Macrophage-tropism can also be assessed by infecting Affinofile cells expressing a wide range of CD4 densities (Joseph et al., 2014). CD4 usage curves reveal that, relative to T cell-tropic viruses, macrophage-tropic viruses are significantly better at infecting cells expressing low levels of CD4, reach 50% infectivity at lower CD4 levels (CD4 ED50) and often approach a plateau level of infectivity at a lower CD4 density (Figure 2; Joseph et al., 2014). This procedure is identical to the or ‘Procedure for High Versus Low CD4 Comparison’ (part A above) except that cells are induced to express 10 levels of CD4 and 30 wells of a 96-well plate are needed to test each virus (10 CD4 levels x 3 replicates per level).
Figure 2. Dose-response curves for infectivity of a macrophage-tropic virus (Ba-L) and a T cell-tropic viruses (JRCSF) to Affinofile cells expressing various densities of CD4. The sensitivity of pseudotyped viruses to CD4 levels was assessed by measuring their ability to infect Affinofile cells expressing 10 CD4 levels (1,425, 4,590, 4,981, 9,374, 22,667, 33,842, 46,204, 58,153, 69,897, and 81,649 CD4 ABS per cell), with CCR5 fully induced. In this assay, the vast majority of blood-derived viruses have an infectivity profile similar to that of JRCSF ( Ping et al., 2013).
Day 2- Prepare media for continuous CD4 comparison.
- Dilute Doxycycline stock to 24 ng/ml in DMEM-F10/B.
- Serially dilute Doxycycline to the following concentrations: 6, 4, 2.8, 2, 1.2, 0.8, 0.4, 0.28 ng/ml.
- Dilute Ponasterone A stock to 20 µM in DMEM-F10/B.
- Induction medium (10 levels of Doxycycline and 1 high level of Pon A): Mix equal volumes of 4x Doxycycline and 4x Ponasterone A (Table 1).
Table 1. Dilutions of Doxycycline and Ponasterone A to induce ten levels of CD4 expression and a single, high level of CCR5 expression4x Doxycycline (ng/ml) 4x Ponasterone A (µM) 1x Final Doxycycline concentration 1x Final Ponasterone A concentration 24 20 6 5 6 20 1.5 5 4 20 1 5 2.8 20 0.7 5 2 20 0.5 5 1.2 20 0.3 5 0.8 20 0.2 5 0.4 20 0.1 5 0.28 20 0.07 5 0 20 0 5
- Dilute Doxycycline stock to 24 ng/ml in DMEM-F10/B.
- Induce cells for infection.
- Remove 50 µl of media from each well of the 96-well plate.
- For each virus being tested, at each Doxycycline level, add 50 µl of each induction medium to 3 wells (10 induction levels x 3 replicates = 30 wells / virus).
- Incubate cells at 37 °C.
- Remove 50 µl of media from each well of the 96-well plate.
- Induce cells for flow cytometry
- Remove all medium from the 24 well plate, replace with 750 µl of warm DMEM-F10/B and 250 µl of either 4x Doxy or 4x Pon A (=1x desired amount of Pon A or Doxy).
- Incubate cells at 37 °C.
Days 3 and 5 are the same as described in ‘Procedure for High Versus Low CD4 Comparison’ (part A above).
- Remove all medium from the 24 well plate, replace with 750 µl of warm DMEM-F10/B and 250 µl of either 4x Doxy or 4x Pon A (=1x desired amount of Pon A or Doxy).
- Prepare media for continuous CD4 comparison.
Recipes
- Poly-L-Lysine (0.1 mg/ml)
Diluted 10-1 in sterile PBS (10 µg/ml) and stored at 4 °C - Ponasterone A
Diluted in 50% ethanol to 1 mM and stored in 100 µl aliquots at -20 °C - Doxycycline
Diluted in sterile water to 10 mg/ml and frozen in 20 µl aliquots at -20 °C (do not freeze-thaw) - DMEM-F10/B
450 ml Cellgro DMEM-H
2.5 ml of blasticidin (final conc = 50 mg/ml, do not freeze/thaw blasticidin)
50 ml of dialyzed FBS (final = 10%)
Stored at 4 °C - 1x staining solution
3 µl of antibody (anti-CD4, anti-CCR5 or isotype controls)
2 µl of aqua live/dead stain
45 µl of PBS
1% FBS - Fixing solution
200 µl of 10% formalin
800 µl H2O
Acknowledgments
We would like to thank Kathryn T. Arrildt for helpful comments that improved this protocol. This work was funded by the National Institutes of Health grant R37 AI44667 to RS.
References
- Dunfee, R. L., Thomas, E. R., Gorry, P. R., Wang, J., Taylor, J., Kunstman, K., Wolinsky, S. M. and Gabuzda, D. (2006). The HIV Env variant N283 enhances macrophage tropism and is associated with brain infection and dementia. Proc Natl Acad Sci U S A 103(41): 15160-15165.
- Duenas-Decamp, M. J., Peters, P. J., Burton, D. and Clapham, P. R. (2009). Determinants flanking the CD4 binding loop modulate macrophage tropism of human immunodeficiency virus type 1 R5 envelopes. J Virol 83(6): 2575-2583.
- Gorry, P. R., Taylor, J., Holm, G. H., Mehle, A., Morgan, T., Cayabyab, M., Farzan, M., Wang, H., Bell, J. E., Kunstman, K., Moore, J. P., Wolinsky, S. M. and Gabuzda, D. (2002). Increased CCR5 affinity and reduced CCR5/CD4 dependence of a neurovirulent primary human immunodeficiency virus type 1 isolate. J Virol 76(12): 6277-6292.
- Johnston, S. H., Lobritz, M. A., Nguyen, S., Lassen, K., Delair, S., Posta, F., Bryson, Y. J., Arts, E. J., Chou, T. and Lee, B. (2009). A quantitative affinity-profiling system that reveals distinct CD4/CCR5 usage patterns among human immunodeficiency virus type 1 and simian immunodeficiency virus strains. J Virol 83(21): 11016-11026.
- *Joseph, S. B., Arrildt, K. T., Swanstrom, A. E., Schnell, G., Lee, B., Hoxie, J. A. and Swanstrom, R. (2014). Quantification of Entry Phenotypes of Macrophage-Tropic HIV-1 across a Wide Range of CD4 Densities. J Virol 88(4): 1858-1869.
- O'Doherty, U., Swiggard, W. J. and Malim, M. H. (2000). Human immunodeficiency virus type 1 spinoculation enhances infection through virus binding. J Virol 74(21): 10074-10080.
- Peters, P. J., Bhattacharya, J., Hibbitts, S., Dittmar, M. T., Simmons, G., Bell, J., Simmonds, P. and Clapham, P. R. (2004). Biological analysis of human immunodeficiency virus type 1 R5 envelopes amplified from brain and lymph node tissues of AIDS patients with neuropathology reveals two distinct tropism phenotypes and identifies envelopes in the brain that confer an enhanced tropism and fusigenicity for macrophages. J Virol 78(13): 6915-6926.
- Peters, P. J., Sullivan, W. M., Duenas-Decamp, M. J., Bhattacharya, J., Ankghuambom, C., Brown, R., Luzuriaga, K., Bell, J., Simmonds, P., Ball, J. and Clapham, P. R. (2006). Non-macrophage-tropic human immunodeficiency virus type 1 R5 envelopes predominate in blood, lymph nodes, and semen: implications for transmission and pathogenesis. J Virol 80(13): 6324-6332.
- Ping, L. H., Joseph, S. B., Anderson, J. A., Abrahams, M. R., Salazar-Gonzalez, J. F., Kincer, L. P., Treurnicht, F. K., Arney, L., Ojeda, S., Zhang, M., Keys, J., Potter, E. L., Chu, H., Moore, P., Salazar, M. G., Iyer, S., Jabara, C., Kirchherr, J., Mapanje, C., Ngandu, N., Seoighe, C., Hoffman, I., Gao, F., Tang, Y., Labranche, C., Lee, B., Saville, A., Vermeulen, M., Fiscus, S., Morris, L., Karim, S. A., Haynes, B. F., Shaw, G. M., Korber, B. T., Hahn, B. H., Cohen, M. S., Montefiori, D., Williamson, C., Swanstrom, R., Study, C. A. I. and the Center for, H. I. V. A. V. I. C. (2013). Comparison of viral Env proteins from acute and chronic infections with subtype C human immunodeficiency virus type 1 identifies differences in glycosylation and CCR5 utilization and suggests a new strategy for immunogen design. J Virol 87(13): 7218-7233.
- Martin-Garcia, J., Cao, W., Varela-Rohena, A., Plassmeyer, M. L. and Gonzalez-Scarano, F. (2006). HIV-1 tropism for the central nervous system: Brain-derived envelope glycoproteins with lower CD4 dependence and reduced sensitivity to a fusion inhibitor. Virology 346(1): 169-179.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Joseph, S. B., Lee, B. and Swanstrom, R. (2014). Affinofile Assay for Identifying Macrophage-Tropic HIV-1. Bio-protocol 4(14): e1184. DOI: 10.21769/BioProtoc.1184.
Category
Microbiology > Microbe-host interactions > Virus
Microbiology > Microbial cell biology > Cell viability
Immunology > Immune cell function > Macrophage
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









