- EN - English
- CN - 中文
Ribosome Fractionation in Yeast
酵母核糖体的分离
发布: 2012年08月20日第2卷第16期 DOI: 10.21769/BioProtoc.251 浏览次数: 22955
Abstract
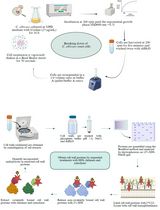
This protocol describes yeast ribosome fractionation in the gradient of sucrose. During the cyclic process of translation, a small (40S) and large (60S) ribosomal subunit associate with mRNA to form an 80S complex (monosome). This ribosome moves along the mRNA during translational elongation, and then dissociates into the 40S and 60S subunits on termination. During elongation by one ribosome, further ribosomes can initiate translation on the same mRNA to form polysomes. The mass of each polysomal complex is determined primarily by the number of ribosomes it contains. Hence, the population of polysomes within the cell can be size-fractionated by sucrose density gradient centrifugation on the basis of the loading of ribosomes on the mRNA. Several compounds help to maintain or to disrupt the polysomes (Figure 1).
Keywords: Ribosome (核糖体)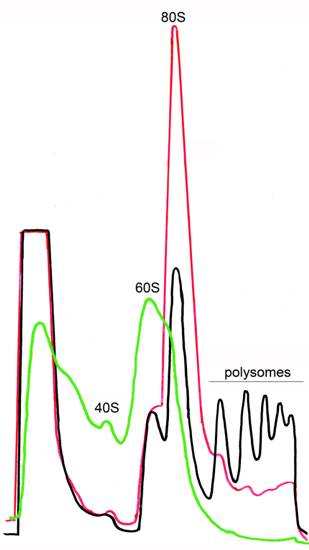
Figure 1. Ribosome profiles after sucrose gradient centrifugation. Ribosome fractionation was performed in the presence of CHX (black), in the absence of CHX (red) or in the presence of EDTA (green). CHX stabilizes polysomes. In the absence of CHX polysomes are destroyed but 80S is still present. 80S can be completely disrupted in cell extracts by treatment with EDTA.
Materials and Reagents
- Glass beads for cells breaking 0.5 mm (Bio Spec Products, catalog number: 110/9105 )
- Cycloheximide (CHX) (Sigma-Aldrich, catalog number: C7698 ) solution 100 mg/ml prepared on ethanol
- Bradford reactive (Bio-Rad Laboratories, catalog number: 500-0006 )
- Protease inhibitor cocktail (F. Hoffmann-La Roche, catalog number: 13560400 )
- Phenylmethylsulfonyl fluoride (PMSF) (Sigma-Aldrich, catalog number: P7625 ) 100 mM solution prepared on isopropanol
- 100% trichloroacetic acid (TCA) (Fluka, catalog number: 91230 )
- Tubes for SW41 rotor Ti (Beckman Coulter, catalog number: 331372 )
- Protease inhibitor cocktail (F. Hoffmann-La Roche, catalog number: 1836170 )
- Lysis buffer
- Ethanol
- Laemmli sample buffer
- Bromophenol blue
- YPD media (see Recipes)
- Buffer A (see Recipes)
- 2x buffer B2 times concentrated (see Recipes)
- Sucrose gradient (see Recipes)
- Foni-inert (see Recipes)
Equipment
- Table Centrifuges
- Glass bead beater Genie Disruptor (Scientific Industries, catalog number: SI-DD38 )
- Ultracentrifuge
- Rotor SW41 Ti (Beckman Coulter, catalog number: 331362 )
- Density Gradient Fractionation System (ISCO, catalog number: 67-9000-177 )
Procedure
文章信息
版权信息
© 2012 The Authors; exclusive licensee Bio-protocol LLC.
如何引用
Panasenko, O. O. (2012). Ribosome Fractionation in Yeast. Bio-protocol 2(16): e251. DOI: 10.21769/BioProtoc.251.
分类
微生物学 > 微生物细胞生物学 > 细胞器分离
生物化学 > 蛋白质 > 分离和纯化
您对这篇实验方法有问题吗?
在此处发布您的问题,我们将邀请本文作者来回答。同时,我们会将您的问题发布到Bio-protocol Exchange,以便寻求社区成员的帮助。
Share
Bluesky
X
Copy link