- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Collecting and Fixing Nuclear GFP/RFP in L1 Larva for Imaging
Published: Vol 2, Iss 5, Mar 5, 2012 DOI: 10.21769/BioProtoc.94 Views: 11054

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
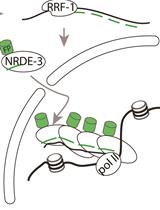
Labelling of Active Transcription Sites with Argonaute NRDE-3—Image Active Transcription Sites in vivo in Caenorhabditis elegans
Antoine Barrière and Vincent Bertrand
Jun 5, 2022 2705 Views
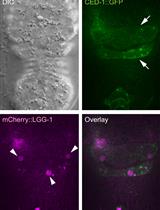
Monitoring the Recruitment and Fusion of Autophagosomes to Phagosomes During the Clearance of Apoptotic Cells in the Nematode Caenorhabditis elegans
Omar Peña-Ramos and Zheng Zhou
Nov 20, 2022 2153 Views
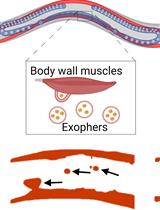
Preparation of Caenorhabditis elegans for Scoring of Muscle-derived Exophers
Katarzyna Banasiak [...] Wojciech Pokrzywa
Jan 5, 2023 1899 Views
Abstract
In this protocol, L1 stage larvae are collected that carry nuclear-localized GFP/wCherry reporters. These can be fixed so that the GFP/wCherry maintains nuclear localization and stain nuclei by DAPI. This protocol therefore achieves the collection and fixation of nuclei in worm L1 larvae.
Materials and Reagents
- Acetone
- Formaldehyde
- DAPI
- Poly-lysine
- Glycerol
- Cytoseal 280 (Richard-Allan Scientific, catalog number: 8311-4 )
- Used Qiaquick Spin Column (QIAGEN)
- 11.58 μm glassbeads (Whitehouse Scientific, catalog number: MS0012 )
- KCl
- NaCl
- Na2EGTA
- Triton X-100
- EDTA
- PIPES
- 2x modified MRWB (see Recipes)
- DAPI staining solution (see Recipes)
- M9 buffer (see Recipes)
- Tris triton buffer (TTB) (see Recipes)
Equipment
- Standard bench top microcentrifuge
- 16-slide glass staining jar (Thermo Fisher Scientific, catalog number: 08-810 )
- Spatula
- Microscope
- Glass container
- Glass coverslip
- Glass slide
- 18 x 18 mm glass cover slip
- 25 x 75 mm glass slide
Procedure
- Preparing larva
- Begin with a plate that contains many eggs (100+)
- Use the spatula to remove any chunks which many retain worms. Be sure to flame between uses so no worms are transferred between plates.
- Using a spatula, carefully displace and remove the agar from the plastic plate. Place the agar in a 16-slide glass-staining jar keeping the surface with worms facing upwards.
- Rinse the agar in the glass container three times with deionized water, taking care that water does not directly hit the agar.
- Using the spatula, place the agar back into the plastic container. Remember which side faces upward!
- Check under the microscope to make sure no worms are left on the plate. If worms are left, repeat rinses until no worms are left. There should be plenty of eggs (100+).
- Leave the plate at room temperature (RT) (25 °C) for 2 h.
- Check that L1 larva has emerged.
- Begin with a plate that contains many eggs (100+)
- Freezing and Fixing the Worms
- Wash the plate with 500 μl M9 buffer and transfer to 1.5 ml centrifuge tube. Repeat twice.
- Spin 3,000 rpm, 2 min. Remove supernatant taking care not to disturb worms at the bottom. Resuspend with 1 ml M9.
- Spin 3,000 rpm, 2 min. Remove supernatant taking care not to disturb worms at the bottom. Resuspend with 500 μl M9.
- Transfer to a used Qiaquick Spin Column. Spin 3,000 rpm, 2 min with lid open.
- Close Qiaquick column cap and place column and collection tube separated into a bucket. Add liquid nitrogen. The next few steps should be performed as quickly as possible after liquid nitrogen is added.
- Add 200 μl acetone (-20 °C) to the column and immediately spin 2,000 rpm, 30 sec.
- Add 200 μl acetone (-20 °C) to the column and place in -20 °C freezer for 1 min. Then spin 2,000 rpm 30 sec.
- Add 200 μl fresh MWRB/formaldehyde solution (50% 2x modified MRWB, 5% formaldehyde = 100 μl 2x modified MRWB, 100 μl 10% formaldehyde) and let sit at RT for 1 h. Then spin at 2,000 rpm, 30 sec.
- Add 200 μl TTB and spin 2,000 rpm, 30 sec. Repeat to remove all formaldehyde.
- Add 200 μl DAPI staining solution. Let sit at RT for 1 h. Spin 2,000 rpm, 30 sec.
- Add 200 μl TTB and pipette up and down to resuspend the worms in the solution. Transfer to 1.5 ml centrifuge tube to collect worms.
- Wash the plate with 500 μl M9 buffer and transfer to 1.5 ml centrifuge tube. Repeat twice.
- Preparing the slides
- Add 75 μl of .5% poly-lysine (in H2O) to a 18 x 18 mm glass cover slip. Cover using plastic dish lid and let sit at RT for at least 30 min.
- Recollect excess poly-lysine.
- Wash cover slip using distilled H2O and air dry.
- Add a drop of well suspended 11.58 μm glass beads (in acetone) onto the treated surface of a 25 x 75 mm glass slide. Air dry.
- Add 75 μl of .5% poly-lysine (in H2O) to a 18 x 18 mm glass cover slip. Cover using plastic dish lid and let sit at RT for at least 30 min.
- Mounting the Worms
- Add 20 μl of worms in TTB to the poly-lysine treated side of the coverslip. Leave at RT for 30 min to allow worms to stick to the coverslip.
- Remove as much TTB as possible but observe this removal step under the microscope to make sure most worms are stuck to the poly-lysine.
- Add 75 μl 50% glycerol to the coverslip. Carefully remove glycerol from the sides. Add 50 μl 50% glycerol to the middle of the coverslip. This helps to disperse an excess TTB, making the drop closer to 50% glycerol. Remove excess liquid from the sides.
- Place slide over the cover slip and seal with Cytoseal 280.
- Place in a slide holder and refrigerate until use.
- Add 20 μl of worms in TTB to the poly-lysine treated side of the coverslip. Leave at RT for 30 min to allow worms to stick to the coverslip.
Recipes
- 2x modified MRWB
160 mM KCl
40 mM NaCl
20 mM Na2EGTA
10 mM Spermidine HCl
30 mM PIPES (pH 7.4) - TTB (Tris triton buffer)
100 mM Tris-HCl (pH 7.4)
1% Triton X-100
1 mM EDTA - M9 buffer
Refer to: common worm media and buffers - DAPI staining solution
100 μl M9
100 μl deionized H2O
0.1 μl 1 mg/ml DAPI
Acknowledgments
This protocol has been adapted from Rigaut et al. (1999) and Puig et al. (2001).
References
- Puig, O., Caspary, F., Rigaut, G., Rutz, B., Bouveret, E., Bragado-Nilsson, E., Wilm, M. and Seraphin, B. (2001). The tandem affinity purification (TAP) method: a general procedure of protein complex purification. Methods 24(3): 218-229.
- Rigaut, G., Shevchenko, A., Rutz, B., Wilm, M., Mann, M. and Seraphin, B. (1999). A generic protein purification method for protein complex characterization and proteome exploration. Nat Biotechnol 17(10): 1030-1032.
Article Information
Copyright
© 2012 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Liu, X. (2012). Collecting and Fixing Nuclear GFP/RFP in L1 Larva for Imaging. Bio-protocol 2(5): e94. DOI: 10.21769/BioProtoc.94.
Category
Cell Biology > Cell imaging > Fluorescence
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










